-
Notifications
You must be signed in to change notification settings - Fork 2.6k
Commit
This commit does not belong to any branch on this repository, and may belong to a fork outside of the repository.
[Project] Medical semantic seg dataset: Chest x ray images with pneum…
…othorax masks (#2687)
- Loading branch information
1 parent
c923f4d
commit d3f2922
Showing
9 changed files
with
305 additions
and
6 deletions.
There are no files selected for viewing
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
119 changes: 119 additions & 0 deletions
119
...cts/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/README.md
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,119 @@ | ||
| # Chest X-ray Images with Pneumothorax Masks | ||
|
|
||
| ## Description | ||
|
|
||
| This project support **`Chest X-ray Images with Pneumothorax Masks `**, and the dataset used in this project can be downloaded from [here](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks). | ||
|
|
||
| ### Dataset Overview | ||
|
|
||
| A pneumothorax (noo-moe-THOR-aks) is a collapsed lung. A pneumothorax occurs when air leaks into the space between your lung and chest wall. This air pushes on the outside of your lung and makes it collapse. Pneumothorax can be a complete lung collapse or a collapse of only a portion of the lung. | ||
|
|
||
| A pneumothorax can be caused by a blunt or penetrating chest injury, certain medical procedures, or damage from underlying lung disease. Or it may occur for no obvious reason. Symptoms usually include sudden chest pain and shortness of breath. On some occasions, a collapsed lung can be a life-threatening event. | ||
|
|
||
| Treatment for a pneumothorax usually involves inserting a needle or chest tube between the ribs to remove the excess air. However, a small pneumothorax may heal on its own. | ||
|
|
||
| ### Statistic Information | ||
|
|
||
| | Dataset Name | Anatomical Region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release date | License | | ||
| | --------------------------------------------------------------------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- | | ||
| | [Chest-x-ray-images-with-pneumothorax-masks](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks) | throax | segmentation | x_ray | 2 | 10675/-/1372 | yes/-/yes | 2020 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) | | ||
|
|
||
| | Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test | | ||
| | :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: | | ||
| | background | 10675 | 99.7 | - | - | 1372 | 99.71 | | ||
| | pneumothroax | 2379 | 0.3 | - | - | 290 | 0.29 | | ||
|
|
||
| ### Visualization | ||
|
|
||
| 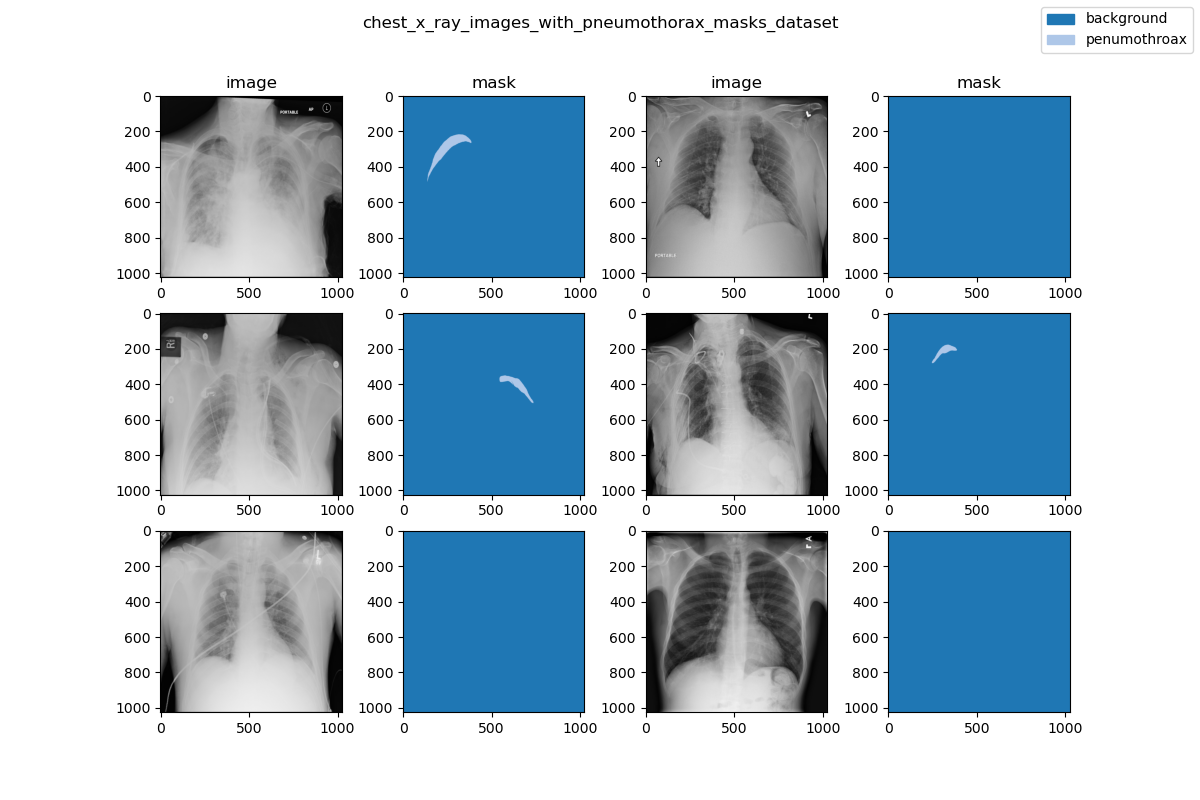 | ||
|
|
||
| ### Prerequisites | ||
|
|
||
| - Python 3.8 | ||
| - PyTorch 1.10.0 | ||
| - pillow(PIL) 9.3.0 | ||
| - scikit-learn(sklearn) 1.2.0 | ||
| - [MIM](https://github.com/open-mmlab/mim) v0.3.4 | ||
| - [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4 | ||
| - [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher | ||
| - [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5 | ||
|
|
||
| All the commands below rely on the correct configuration of PYTHONPATH, which should point to the project's directory so that Python can locate the module files. In chest_x_ray_images_with_pneumothorax_masks/ root directory, run the following line to add the current directory to PYTHONPATH: | ||
|
|
||
| ```shell | ||
| export PYTHONPATH=`pwd`:$PYTHONPATH | ||
| ``` | ||
|
|
||
| ### Dataset preparing | ||
|
|
||
| - download dataset from [here](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks) and decompression data to path 'data/'. | ||
| - run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below. | ||
| - run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly. | ||
|
|
||
| ```none | ||
| mmsegmentation | ||
| ├── mmseg | ||
| ├── projects | ||
| │ ├── medical | ||
| │ │ ├── 2d_image | ||
| │ │ │ ├── x_ray | ||
| │ │ │ │ ├── chest_x_ray_images_with_pneumothorax_masks | ||
| │ │ │ │ │ ├── configs | ||
| │ │ │ │ │ ├── datasets | ||
| │ │ │ │ │ ├── tools | ||
| │ │ │ │ │ ├── data | ||
| │ │ │ │ │ │ ├── train.txt | ||
| │ │ │ │ │ │ ├── val.txt | ||
| │ │ │ │ │ │ ├── images | ||
| │ │ │ │ │ │ │ ├── train | ||
| │ │ │ │ | │ │ │ ├── xxx.png | ||
| │ │ │ │ | │ │ │ ├── ... | ||
| │ │ │ │ | │ │ │ └── xxx.png | ||
| │ │ │ │ │ │ ├── masks | ||
| │ │ │ │ │ │ │ ├── train | ||
| │ │ │ │ | │ │ │ ├── xxx.png | ||
| │ │ │ │ | │ │ │ ├── ... | ||
| │ │ │ │ | │ │ │ └── xxx.png | ||
| ``` | ||
|
|
||
| ### Training commands | ||
|
|
||
| ```shell | ||
| mim train mmseg ./configs/${CONFIG_PATH} | ||
| ``` | ||
|
|
||
| To train on multiple GPUs, e.g. 8 GPUs, run the following command: | ||
|
|
||
| ```shell | ||
| mim train mmseg ./configs/${CONFIG_PATH} --launcher pytorch --gpus 8 | ||
| ``` | ||
|
|
||
| ### Testing commands | ||
|
|
||
| ```shell | ||
| mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH} | ||
| ``` | ||
|
|
||
| ## Checklist | ||
|
|
||
| - [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`. | ||
|
|
||
| - [x] Finish the code | ||
| - [x] Basic docstrings & proper citation | ||
| - [x] Test-time correctness | ||
| - [x] A full README | ||
|
|
||
| - [x] Milestone 2: Indicates a successful model implementation. | ||
|
|
||
| - [x] Training-time correctness | ||
|
|
||
| - [ ] Milestone 3: Good to be a part of our core package! | ||
|
|
||
| - [ ] Type hints and docstrings | ||
| - [ ] Unit tests | ||
| - [ ] Code polishing | ||
| - [ ] Metafile.yml | ||
|
|
||
| - [ ] Move your modules into the core package following the codebase's file hierarchy structure. | ||
|
|
||
| - [ ] Refactor your modules into the core package following the codebase's file hierarchy structure. |
42 changes: 42 additions & 0 deletions
42
...ges_with_pneumothorax_masks/configs/chest-x-ray-images-with-pneumothorax-masks_512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,42 @@ | ||
| dataset_type = 'ChestPenumoMaskDataset' | ||
| data_root = 'data/' | ||
| img_scale = (512, 512) | ||
| train_pipeline = [ | ||
| dict(type='LoadImageFromFile'), | ||
| dict(type='LoadAnnotations'), | ||
| dict(type='Resize', scale=img_scale, keep_ratio=False), | ||
| dict(type='RandomFlip', prob=0.5), | ||
| dict(type='PhotoMetricDistortion'), | ||
| dict(type='PackSegInputs') | ||
| ] | ||
| test_pipeline = [ | ||
| dict(type='LoadImageFromFile'), | ||
| dict(type='Resize', scale=img_scale, keep_ratio=False), | ||
| dict(type='LoadAnnotations'), | ||
| dict(type='PackSegInputs') | ||
| ] | ||
| train_dataloader = dict( | ||
| batch_size=16, | ||
| num_workers=4, | ||
| persistent_workers=True, | ||
| sampler=dict(type='InfiniteSampler', shuffle=True), | ||
| dataset=dict( | ||
| type=dataset_type, | ||
| data_root=data_root, | ||
| ann_file='train.txt', | ||
| data_prefix=dict(img_path='images/', seg_map_path='masks/'), | ||
| pipeline=train_pipeline)) | ||
| val_dataloader = dict( | ||
| batch_size=1, | ||
| num_workers=4, | ||
| persistent_workers=True, | ||
| sampler=dict(type='DefaultSampler', shuffle=False), | ||
| dataset=dict( | ||
| type=dataset_type, | ||
| data_root=data_root, | ||
| ann_file='val.txt', | ||
| data_prefix=dict(img_path='images/', seg_map_path='masks/'), | ||
| pipeline=test_pipeline)) | ||
| test_dataloader = val_dataloader | ||
| val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice']) | ||
| test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice']) |
20 changes: 20 additions & 0 deletions
20
...6_unet-{use-sigmoid}_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,20 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', | ||
| './chest-x-ray-images-with-pneumothorax-masks_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict( | ||
| imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.01) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict( | ||
| num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
19 changes: 19 additions & 0 deletions
19
...n-unet-s5-d16_unet_1xb16-0.0001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,19 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', | ||
| './chest-x-ray-images-with-pneumothorax-masks_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict( | ||
| imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.0001) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict(num_classes=2), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
19 changes: 19 additions & 0 deletions
19
...cn-unet-s5-d16_unet_1xb16-0.001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,19 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', | ||
| './chest-x-ray-images-with-pneumothorax-masks_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict( | ||
| imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.001) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict(num_classes=2), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
19 changes: 19 additions & 0 deletions
19
...fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,19 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', | ||
| './chest-x-ray-images-with-pneumothorax-masks_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict( | ||
| imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.01) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict(num_classes=2), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
31 changes: 31 additions & 0 deletions
31
...es_with_pneumothorax_masks/datasets/chest-x-ray-images-with-pneumothorax-masks_dataset.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,31 @@ | ||
| from mmseg.datasets import BaseSegDataset | ||
| from mmseg.registry import DATASETS | ||
|
|
||
|
|
||
| @DATASETS.register_module() | ||
| class ChestPenumoMaskDataset(BaseSegDataset): | ||
| """ChestPenumoMaskDataset dataset. | ||
| In segmentation map annotation for ChestPenumoMaskDataset, | ||
| 0 stands for background, which is included in 2 categories. | ||
| ``reduce_zero_label`` is fixed to False. The ``img_suffix`` | ||
| is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'. | ||
| Args: | ||
| img_suffix (str): Suffix of images. Default: '.png' | ||
| seg_map_suffix (str): Suffix of segmentation maps. Default: '.png' | ||
| reduce_zero_label (bool): Whether to mark label zero as ignored. | ||
| Default to False. | ||
| """ | ||
| METAINFO = dict(classes=('background', 'penumothroax')) | ||
|
|
||
| def __init__(self, | ||
| img_suffix='.png', | ||
| seg_map_suffix='.png', | ||
| reduce_zero_label=False, | ||
| **kwargs) -> None: | ||
| super().__init__( | ||
| img_suffix=img_suffix, | ||
| seg_map_suffix=seg_map_suffix, | ||
| reduce_zero_label=reduce_zero_label, | ||
| **kwargs) |
36 changes: 36 additions & 0 deletions
36
...edical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/tools/prepare_dataset.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,36 @@ | ||
| import glob | ||
| import os | ||
| import shutil | ||
|
|
||
| from PIL import Image | ||
| from sklearn.model_selection import train_test_split | ||
|
|
||
| root_path = 'data/' | ||
| img_suffix = '.png' | ||
| seg_map_suffix = '.png' | ||
| save_img_suffix = '.png' | ||
| save_seg_map_suffix = '.png' | ||
|
|
||
| all_imgs = glob.glob('data/siim-acr-pneumothorax/png_images/*' + img_suffix) | ||
| x_train, x_test = train_test_split(all_imgs, test_size=0.2, random_state=0) | ||
|
|
||
| print(len(x_train), len(x_test)) | ||
| os.system('mkdir -p ' + root_path + 'images/train/') | ||
| os.system('mkdir -p ' + root_path + 'images/val/') | ||
| os.system('mkdir -p ' + root_path + 'masks/train/') | ||
| os.system('mkdir -p ' + root_path + 'masks/val/') | ||
|
|
||
| part_dir_dict = {0: 'train/', 1: 'val/'} | ||
| for ith, part in enumerate([x_train, x_test]): | ||
| part_dir = part_dir_dict[ith] | ||
| for img in part: | ||
| basename = os.path.basename(img) | ||
| img_save_path = os.path.join(root_path, 'images', part_dir, | ||
| basename.split('.')[0] + save_img_suffix) | ||
| shutil.copy(img, img_save_path) | ||
| mask_path = 'data/siim-acr-pneumothorax/png_masks/' + basename | ||
| mask = Image.open(mask_path).convert('L') | ||
| mask_save_path = os.path.join( | ||
| root_path, 'masks', part_dir, | ||
| basename.split('.')[0] + save_seg_map_suffix) | ||
| mask.save(mask_save_path) |