-
Notifications
You must be signed in to change notification settings - Fork 2.6k
Commit
This commit does not belong to any branch on this repository, and may belong to a fork outside of the repository.
[Project] Medical semanti seg dataset: Fusc2021 (#2682)
- Loading branch information
1 parent
d934d10
commit ac24111
Showing
8 changed files
with
391 additions
and
0 deletions.
There are no files selected for viewing
136 changes: 136 additions & 0 deletions
136
projects/medical/2d_image/histopathology/fusc2021/README.md
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,136 @@ | ||
| # Foot Ulcer Segmentation Challenge 2021 (FUSC 2021) | ||
|
|
||
| ## Description | ||
|
|
||
| This project supports **`Foot Ulcer Segmentation Challenge 2021 (FUSC 2021) `**, which can be downloaded from [here](https://fusc.grand-challenge.org/). | ||
|
|
||
| ### Dataset Overview | ||
|
|
||
| This chronic wound dataset was collected over 2 years from October 2019 to April 2021 at the center and contains 1,210 foot ulcer images taken from 889 patients during multiple clinical visits. The raw images were taken by Canon SX 620 HS digital camera and iPad Pro under uncontrolled illumination conditions, | ||
| with various backgrounds. The images (shown in Figure 1) are randomly split into 3 subsets: a training set with 810 images, a validation set with 200 images, and a testing set with 200 images. Of course, the annotations of the testing set are kept private. The data collected were de-identified and in accordance with relevant guidelines and regulations and the patient’s informed consent is waived by the institutional review board of the University of Wisconsin-Milwaukee. | ||
|
|
||
| ### Information Statistics | ||
|
|
||
| | Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License | | ||
| | --------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- | | ||
| | [fusc2021](https://fusc.grand-challenge.org/) | lower limb | segmentation | histopathology | 2 | 810/200/200 | yes/yes/no | 2021 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) | | ||
|
|
||
| | Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test | | ||
| | :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: | | ||
| | background | 810 | 98.71 | 200 | 98.78 | - | - | | ||
| | wound | 791 | 1.29 | 195 | 1.22 | - | - | | ||
|
|
||
| Note: | ||
|
|
||
| - `Pct` means percentage of pixels in this category in all pixels. | ||
|
|
||
| ### Visualization | ||
|
|
||
| 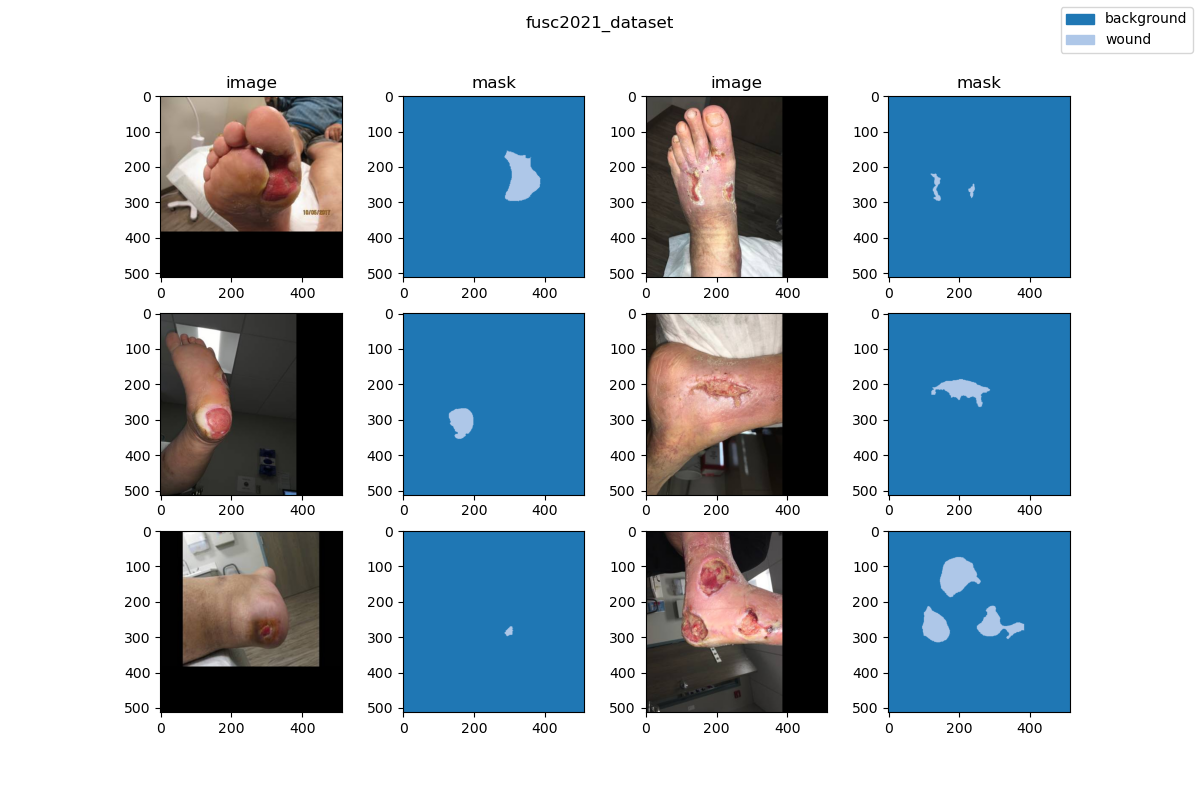 | ||
|
|
||
| ### Dataset Citation | ||
|
|
||
| ``` | ||
| @article{s41598-020-78799-w, | ||
| title={Fully automatic wound segmentation with deep convolutional neural networks}, | ||
| author={Chuanbo Wang and D. M. Anisuzzaman and Victor Williamson and Mrinal Kanti Dhar and Behrouz Rostami and Jeffrey Niezgoda and Sandeep Gopalakrishnan and Zeyun Yu}, | ||
| journal={Scientific Reports}, | ||
| volume={10}, | ||
| number={1}, | ||
| pages={21897}, | ||
| year={2020} | ||
| } | ||
| ``` | ||
|
|
||
| ### Prerequisites | ||
|
|
||
| - Python v3.8 | ||
| - PyTorch v1.10.0 | ||
| - pillow(PIL) v9.3.0 | ||
| - scikit-learn(sklearn) v1.2.0 | ||
| - [MIM](https://github.com/open-mmlab/mim) v0.3.4 | ||
| - [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4 | ||
| - [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher | ||
| - [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5 | ||
|
|
||
| All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `fusc2021/` root directory, run the following line to add the current directory to `PYTHONPATH`: | ||
|
|
||
| ```shell | ||
| export PYTHONPATH=`pwd`:$PYTHONPATH | ||
| ``` | ||
|
|
||
| ### Dataset Preparing | ||
|
|
||
| - download dataset from [here](https://fusc.grand-challenge.org/) and decompress data to path `'data/'`. | ||
| - run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below. | ||
| - run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly. | ||
|
|
||
| ```none | ||
| mmsegmentation | ||
| ├── mmseg | ||
| ├── projects | ||
| │ ├── medical | ||
| │ │ ├── 2d_image | ||
| │ │ │ ├── histopathology | ||
| │ │ │ │ ├── fusc2021 | ||
| │ │ │ │ │ ├── configs | ||
| │ │ │ │ │ ├── datasets | ||
| │ │ │ │ │ ├── tools | ||
| │ │ │ │ │ ├── data | ||
| │ │ │ │ │ │ ├── train.txt | ||
| │ │ │ │ │ │ ├── val.txt | ||
| │ │ │ │ │ │ ├── images | ||
| │ │ │ │ │ │ │ ├── train | ||
| │ │ │ │ | │ │ │ ├── xxx.png | ||
| │ │ │ │ | │ │ │ ├── ... | ||
| │ │ │ │ | │ │ │ └── xxx.png | ||
| │ │ │ │ │ │ ├── masks | ||
| │ │ │ │ │ │ │ ├── train | ||
| │ │ │ │ | │ │ │ ├── xxx.png | ||
| │ │ │ │ | │ │ │ ├── ... | ||
| │ │ │ │ | │ │ │ └── xxx.png | ||
| ``` | ||
|
|
||
| ### Training commands | ||
|
|
||
| To train models on a single server with one GPU. (default) | ||
|
|
||
| ```shell | ||
| mim train mmseg ./configs/${CONFIG_FILE} | ||
| ``` | ||
|
|
||
| ### Testing commands | ||
|
|
||
| To test models on a single server with one GPU. (default) | ||
|
|
||
| ```shell | ||
| mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH} | ||
| ``` | ||
|
|
||
| <!-- List the results as usually done in other model's README. [Example](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/configs/fcn#results-and-models) | ||
| You should claim whether this is based on the pre-trained weights, which are converted from the official release; or it's a reproduced result obtained from retraining the model in this project. --> | ||
|
|
||
| ## Checklist | ||
|
|
||
| - [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`. | ||
|
|
||
| - [x] Finish the code | ||
| - [x] Basic docstrings & proper citation | ||
| - [ ] Test-time correctness | ||
| - [x] A full README | ||
|
|
||
| - [ ] Milestone 2: Indicates a successful model implementation. | ||
|
|
||
| - [ ] Training-time correctness | ||
|
|
||
| - [ ] Milestone 3: Good to be a part of our core package! | ||
|
|
||
| - [ ] Type hints and docstrings | ||
| - [ ] Unit tests | ||
| - [ ] Code polishing | ||
| - [ ] Metafile.yml | ||
|
|
||
| - [ ] Move your modules into the core package following the codebase's file hierarchy structure. | ||
|
|
||
| - [ ] Refactor your modules into the core package following the codebase's file hierarchy structure. |
17 changes: 17 additions & 0 deletions
17
...histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_fusc2021-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,17 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict(imports='datasets.fusc2021_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.0001) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict(num_classes=2), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
17 changes: 17 additions & 0 deletions
17
.../histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_fusc2021-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,17 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict(imports='datasets.fusc2021_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.001) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict(num_classes=2), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
17 changes: 17 additions & 0 deletions
17
...e/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_fusc2021-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,17 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict(imports='datasets.fusc2021_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.01) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict(num_classes=2), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
18 changes: 18 additions & 0 deletions
18
...hology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_fusc2021-512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,18 @@ | ||
| _base_ = [ | ||
| 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py', | ||
| 'mmseg::_base_/default_runtime.py', | ||
| 'mmseg::_base_/schedules/schedule_20k.py' | ||
| ] | ||
| custom_imports = dict(imports='datasets.fusc2021_dataset') | ||
| img_scale = (512, 512) | ||
| data_preprocessor = dict(size=img_scale) | ||
| optimizer = dict(lr=0.01) | ||
| optim_wrapper = dict(optimizer=optimizer) | ||
| model = dict( | ||
| data_preprocessor=data_preprocessor, | ||
| decode_head=dict( | ||
| num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1), | ||
| auxiliary_head=None, | ||
| test_cfg=dict(mode='whole', _delete_=True)) | ||
| vis_backends = None | ||
| visualizer = dict(vis_backends=vis_backends) |
42 changes: 42 additions & 0 deletions
42
projects/medical/2d_image/histopathology/fusc2021/configs/fusc2021_512x512.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,42 @@ | ||
| dataset_type = 'FUSC2021Dataset' | ||
| data_root = 'data/' | ||
| img_scale = (512, 512) | ||
| train_pipeline = [ | ||
| dict(type='LoadImageFromFile'), | ||
| dict(type='LoadAnnotations'), | ||
| dict(type='Resize', scale=img_scale, keep_ratio=False), | ||
| dict(type='RandomFlip', prob=0.5), | ||
| dict(type='PhotoMetricDistortion'), | ||
| dict(type='PackSegInputs') | ||
| ] | ||
| test_pipeline = [ | ||
| dict(type='LoadImageFromFile'), | ||
| dict(type='Resize', scale=img_scale, keep_ratio=False), | ||
| dict(type='LoadAnnotations'), | ||
| dict(type='PackSegInputs') | ||
| ] | ||
| train_dataloader = dict( | ||
| batch_size=16, | ||
| num_workers=4, | ||
| persistent_workers=True, | ||
| sampler=dict(type='InfiniteSampler', shuffle=True), | ||
| dataset=dict( | ||
| type=dataset_type, | ||
| data_root=data_root, | ||
| ann_file='train.txt', | ||
| data_prefix=dict(img_path='images/', seg_map_path='masks/'), | ||
| pipeline=train_pipeline)) | ||
| val_dataloader = dict( | ||
| batch_size=1, | ||
| num_workers=4, | ||
| persistent_workers=True, | ||
| sampler=dict(type='DefaultSampler', shuffle=False), | ||
| dataset=dict( | ||
| type=dataset_type, | ||
| data_root=data_root, | ||
| ann_file='val.txt', | ||
| data_prefix=dict(img_path='images/', seg_map_path='masks/'), | ||
| pipeline=test_pipeline)) | ||
| test_dataloader = val_dataloader | ||
| val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice']) | ||
| test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice']) |
30 changes: 30 additions & 0 deletions
30
projects/medical/2d_image/histopathology/fusc2021/datasets/fusc2021_dataset.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,30 @@ | ||
| from mmseg.datasets import BaseSegDataset | ||
| from mmseg.registry import DATASETS | ||
|
|
||
|
|
||
| @DATASETS.register_module() | ||
| class FUSC2021Dataset(BaseSegDataset): | ||
| """FUSC2021Dataset dataset. | ||
| In segmentation map annotation for FUSC2021Dataset, 0 stands for background | ||
| , which is included in 2 categories. ``reduce_zero_label`` | ||
| is fixed to False. The ``img_suffix`` is fixed to '.png' and | ||
| ``seg_map_suffix`` is fixed to '.png'. | ||
| Args: | ||
| img_suffix (str): Suffix of images. Default: '.png' | ||
| seg_map_suffix (str): Suffix of segmentation maps. Default: '.png' | ||
| reduce_zero_label (bool): Whether to mark label zero as ignored. | ||
| Default to False.. | ||
| """ | ||
| METAINFO = dict(classes=('background', 'wound')) | ||
|
|
||
| def __init__(self, | ||
| img_suffix='.png', | ||
| seg_map_suffix='.png', | ||
| reduce_zero_label=False, | ||
| **kwargs) -> None: | ||
| super().__init__( | ||
| img_suffix=img_suffix, | ||
| seg_map_suffix=seg_map_suffix, | ||
| reduce_zero_label=reduce_zero_label, | ||
| **kwargs) |
114 changes: 114 additions & 0 deletions
114
projects/medical/2d_image/histopathology/fusc2021/tools/prepare_dataset.py
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,114 @@ | ||
| import glob | ||
| import os | ||
|
|
||
| import numpy as np | ||
| from PIL import Image | ||
|
|
||
| root_path = 'data/' | ||
| img_suffix = '.png' | ||
| seg_map_suffix = '.png' | ||
| save_img_suffix = '.png' | ||
| save_seg_map_suffix = '.png' | ||
| src_img_train_dir = os.path.join( | ||
| root_path, 'wound-segmentation/data/' + | ||
| 'Foot Ulcer Segmentation Challenge/train/images') | ||
| src_img_val_dir = os.path.join( | ||
| root_path, 'wound-segmentation/data/' + | ||
| 'Foot Ulcer Segmentation Challenge/validation/images') | ||
| src_img_test_dir = os.path.join( | ||
| root_path, 'wound-segmentation/data/' + | ||
| 'Foot Ulcer Segmentation Challenge/test/images') | ||
| src_mask_train_dir = os.path.join( | ||
| root_path, 'wound-segmentation/data/' + | ||
| 'Foot Ulcer Segmentation Challenge/train/labels') | ||
| src_mask_val_dir = os.path.join( | ||
| root_path, 'wound-segmentation/data/' + | ||
| 'Foot Ulcer Segmentation Challenge/validation/labels') | ||
|
|
||
| tgt_img_train_dir = os.path.join(root_path, 'images/train/') | ||
| tgt_mask_train_dir = os.path.join(root_path, 'masks/train/') | ||
| tgt_img_val_dir = os.path.join(root_path, 'images/val/') | ||
| tgt_mask_val_dir = os.path.join(root_path, 'masks/val/') | ||
| tgt_img_test_dir = os.path.join(root_path, 'images/test/') | ||
| os.system('mkdir -p ' + tgt_img_train_dir) | ||
| os.system('mkdir -p ' + tgt_img_val_dir) | ||
| os.system('mkdir -p ' + tgt_img_test_dir) | ||
| os.system('mkdir -p ' + tgt_mask_train_dir) | ||
| os.system('mkdir -p ' + tgt_mask_val_dir) | ||
|
|
||
|
|
||
| def filter_suffix_recursive(src_dir, suffix): | ||
| # filter out file names and paths in source directory | ||
| suffix = '.' + suffix if '.' not in suffix else suffix | ||
| file_paths = glob.glob( | ||
| os.path.join(src_dir, '**', '*' + suffix), recursive=True) | ||
| file_names = [_.split('/')[-1] for _ in file_paths] | ||
| return sorted(file_paths), sorted(file_names) | ||
|
|
||
|
|
||
| def convert_label(img, convert_dict): | ||
| arr = np.zeros_like(img, dtype=np.uint8) | ||
| for c, i in convert_dict.items(): | ||
| arr[img == c] = i | ||
| return arr | ||
|
|
||
|
|
||
| def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'): | ||
| if not os.path.exists(tgt_dir): | ||
| os.makedirs(tgt_dir) | ||
|
|
||
| src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix) | ||
|
|
||
| for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)): | ||
| tgt_name = src_name.replace(suffix, save_img_suffix) | ||
| tgt_path = os.path.join(tgt_dir, tgt_name) | ||
| num = len(src_paths) | ||
| img = np.array(Image.open(src_path)) | ||
| if len(img.shape) == 2: | ||
| pil = Image.fromarray(img).convert(convert) | ||
| elif len(img.shape) == 3: | ||
| pil = Image.fromarray(img) | ||
| else: | ||
| raise ValueError('Input image not 2D/3D: ', img.shape) | ||
|
|
||
| pil.save(tgt_path) | ||
| print(f'processed {i+1}/{num}.') | ||
|
|
||
|
|
||
| def convert_label_pics_into_pngs(src_dir, | ||
| tgt_dir, | ||
| suffix, | ||
| convert_dict={ | ||
| 0: 0, | ||
| 255: 1 | ||
| }): | ||
| if not os.path.exists(tgt_dir): | ||
| os.makedirs(tgt_dir) | ||
|
|
||
| src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix) | ||
| num = len(src_paths) | ||
| for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)): | ||
| tgt_name = src_name.replace(suffix, save_seg_map_suffix) | ||
| tgt_path = os.path.join(tgt_dir, tgt_name) | ||
|
|
||
| img = np.array(Image.open(src_path).convert('L')) | ||
| img = convert_label(img, convert_dict) | ||
| Image.fromarray(img).save(tgt_path) | ||
| print(f'processed {i+1}/{num}.') | ||
|
|
||
|
|
||
| if __name__ == '__main__': | ||
|
|
||
| convert_pics_into_pngs( | ||
| src_img_train_dir, tgt_img_train_dir, suffix=img_suffix) | ||
|
|
||
| convert_pics_into_pngs(src_img_val_dir, tgt_img_val_dir, suffix=img_suffix) | ||
|
|
||
| convert_pics_into_pngs( | ||
| src_img_test_dir, tgt_img_test_dir, suffix=img_suffix) | ||
|
|
||
| convert_label_pics_into_pngs( | ||
| src_mask_train_dir, tgt_mask_train_dir, suffix=seg_map_suffix) | ||
|
|
||
| convert_label_pics_into_pngs( | ||
| src_mask_val_dir, tgt_mask_val_dir, suffix=seg_map_suffix) |