
diff --git a/docs/zh_cn/advanced_guides/add_datasets.md b/docs/zh_cn/advanced_guides/add_datasets.md
index 4ea14934ed2..22fbf3462fc 100644
--- a/docs/zh_cn/advanced_guides/add_datasets.md
+++ b/docs/zh_cn/advanced_guides/add_datasets.md
@@ -1,4 +1,62 @@
-# 新增自定义数据集(待更新)
+# 新增自定义数据集
+
+## 新增自定义数据集
+
+在这里,我们展示如何构建一个新的数据集。
+
+1. 创建一个新文件 `mmseg/datasets/example.py`
+
+ ```python
+ from mmseg.registry import DATASETS
+ from .basesegdataset import BaseSegDataset
+
+
+ @DATASETS.register_module()
+ class ExampleDataset(BaseSegDataset):
+
+ METAINFO = dict(
+ classes=('xxx', 'xxx', ...),
+ palette=[[x, x, x], [x, x, x], ...])
+
+ def __init__(self, aeg1, arg2):
+ pass
+ ```
+
+2. 在 `mmseg/datasets/__init__.py` 中导入模块
+
+ ```python
+ from .example import ExampleDataset
+ ```
+
+3. 通过创建一个新的数据集配置文件 `configs/_base_/datasets/example_dataset.py` 来使用它
+
+ ```python
+ dataset_type = 'ExampleDataset'
+ data_root = 'data/example/'
+ ...
+ ```
+
+4. 在 `mmseg/utils/class_names.py` 中补充数据集元信息
+
+ ```python
+ def example_classes():
+ return [
+ 'xxx', 'xxx',
+ ...
+ ]
+
+ def example_palette():
+ return [
+ [x, x, x], [x, x, x],
+ ...
+ ]
+ dataset_aliases ={
+ 'example': ['example', ...],
+ ...
+ }
+ ```
+
+**注意:** 如果新数据集不满足 mmseg 的要求,则需要在 `tools/dataset_converters/` 中准备一个数据集预处理脚本
## 通过重新组织数据来定制数据集
@@ -26,30 +84,17 @@
一个训练对将由 img_dir/ann_dir 里同样首缀的文件组成。
-如果给定 `split` 参数,只有部分在 img_dir/ann_dir 里的文件会被加载。
-我们可以对被包括在 split 文本里的文件指定前缀。
+有些数据集不会发布测试集或测试集的标注,如果没有测试集的标注,我们就无法在本地进行评估模型,因此我们在配置文件中将验证集设置为默认测试集。
-除此以外,一个 split 文本如下所示:
-
-```none
-xxx
-zzz
-```
+关于如何构建自己的数据集或实现新的数据集类,请参阅[数据集指南](./datasets.md)以获取更多详细信息。
-只有
-
-`data/my_dataset/img_dir/train/xxx{img_suffix}`,
-`data/my_dataset/img_dir/train/zzz{img_suffix}`,
-`data/my_dataset/ann_dir/train/xxx{seg_map_suffix}`,
-`data/my_dataset/ann_dir/train/zzz{seg_map_suffix}` 将被加载。
-
-注意:标注是跟图像同样的形状 (H, W),其中的像素值的范围是 `[0, num_classes - 1]`。
+**注意:** 标注是跟图像同样的形状 (H, W),其中的像素值的范围是 `[0, num_classes - 1]`。
您也可以使用 [pillow](https://pillow.readthedocs.io/en/stable/handbook/concepts.html#palette) 的 `'P'` 模式去创建包含颜色的标注。
## 通过混合数据去定制数据集
MMSegmentation 同样支持混合数据集去训练。
-当前它支持拼接 (concat), 重复 (repeat) 和多图混合 (multi-image mix)数据集。
+当前它支持拼接 (concat), 重复 (repeat) 和多图混合 (multi-image mix) 数据集。
### 重复数据集
@@ -58,79 +103,29 @@ MMSegmentation 同样支持混合数据集去训练。
```python
dataset_A_train = dict(
- type='RepeatDataset',
- times=N,
- dataset=dict( # 这是 Dataset_A 数据集的原始配置
- type='Dataset_A',
- ...
- pipeline=train_pipeline
- )
+ type='RepeatDataset',
+ times=N,
+ dataset=dict( # 这是 Dataset_A 数据集的原始配置
+ type='Dataset_A',
+ ...
+ pipeline=train_pipeline
)
+)
```
### 拼接数据集
-有2种方式去拼接数据集。
-
-1. 如果您想拼接的数据集是同样的类型,但有不同的标注文件,
- 您可以按如下操作去拼接数据集的配置文件:
-
- 1. 您也许可以拼接两个标注文件夹 `ann_dir`
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = ['anno_dir_1', 'anno_dir_2'],
- pipeline=train_pipeline
- )
- ```
-
- 2. 您也可以去拼接两个 `split` 文件列表
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = 'anno_dir',
- split = ['split_1.txt', 'split_2.txt'],
- pipeline=train_pipeline
- )
- ```
+如果要拼接不同的数据集,可以按如下方式连接数据集配置。
- 3. 您也可以同时拼接 `ann_dir` 文件夹和 `split` 文件列表
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = ['anno_dir_1', 'anno_dir_2'],
- split = ['split_1.txt', 'split_2.txt'],
- pipeline=train_pipeline
- )
- ```
-
- 在这样的情况下, `ann_dir_1` 和 `ann_dir_2` 分别对应于 `split_1.txt` 和 `split_2.txt`
-
-2. 如果您想拼接不同的数据集,您可以如下去拼接数据集的配置文件:
-
- ```python
- dataset_A_train = dict()
- dataset_B_train = dict()
-
- data = dict(
- imgs_per_gpu=2,
- workers_per_gpu=2,
- train = [
- dataset_A_train,
- dataset_B_train
- ],
- val = dataset_A_val,
- test = dataset_A_test
- )
- ```
+```python
+dataset_A_train = dict()
+dataset_B_train = dict()
+concatenate_dataset = dict(
+ type='ConcatDataset',
+ datasets=[dataset_A_train, dataset_B_train])
+```
-一个更复杂的例子如下:分别重复 `Dataset_A` 和 `Dataset_B` N 次和 M 次,然后再去拼接重复后的数据集
+下面是一个更复杂的示例,它分别重复 `Dataset_A` 和 `Dataset_B` N 次和 M 次,然后连接重复的数据集。
```python
dataset_A_train = dict(
@@ -159,41 +154,36 @@ dataset_B_train = dict(
pipeline=train_pipeline
)
)
-data = dict(
- imgs_per_gpu=2,
- workers_per_gpu=2,
- train = [
- dataset_A_train,
- dataset_B_train
- ],
- val = dataset_A_val,
- test = dataset_A_test
-)
+train_dataloader = dict(
+ dataset=dict(
+ type='ConcatDataset',
+ datasets=[dataset_A_train, dataset_B_train]))
+
+val_dataloader = dict(dataset=dataset_A_val)
+test_dataloader = dict(dataset=dataset_A_test)
```
+您可以参考 mmengine 的基础数据集[教程](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/basedataset.html)以了解更多详细信息
+
### 多图混合集
-我们使用 `MultiImageMixDataset` 作为包装(wrapper)去混合多个数据集的图片。
-`MultiImageMixDataset`可以被类似mosaic和mixup的多图混合数据増广使用。
+我们使用 `MultiImageMixDataset` 作为包装(wrapper)去混合多个数据集的图片。
+`MultiImageMixDataset`可以被类似 mosaic 和 mixup 的多图混合数据増广使用。
-`MultiImageMixDataset`与`Mosaic`数据増广一起使用的例子:
+`MultiImageMixDataset` 与 `Mosaic` 数据増广一起使用的例子:
```python
train_pipeline = [
dict(type='RandomMosaic', prob=1),
dict(type='Resize', img_scale=(1024, 512), keep_ratio=True),
dict(type='RandomFlip', prob=0.5),
- dict(type='Normalize', **img_norm_cfg),
- dict(type='DefaultFormatBundle'),
- dict(type='Collect', keys=['img', 'gt_semantic_seg']),
+ dict(type='PackSegInputs')
]
train_dataset = dict(
type='MultiImageMixDataset',
dataset=dict(
- classes=classes,
- palette=palette,
type=dataset_type,
reduce_zero_label=False,
img_dir=data_root + "images/train",
diff --git a/docs/zh_cn/advanced_guides/add_metrics.md b/docs/zh_cn/advanced_guides/add_metrics.md
index 3a371e357e8..0637b447284 100644
--- a/docs/zh_cn/advanced_guides/add_metrics.md
+++ b/docs/zh_cn/advanced_guides/add_metrics.md
@@ -1 +1,81 @@
-# 新增评测指标 (待更新)
+# 新增评测指标
+
+## 使用 MMSegmentation 的源代码进行开发
+
+在这里,我们用 `CustomMetric` 作为例子来展示如何开发一个新的评测指标。
+
+1. 创建一个新文件 `mmseg/evaluation/metrics/custom_metric.py`。
+
+ ```python
+ from typing import List, Sequence
+
+ from mmengine.evaluator import BaseMetric
+
+ from mmseg.registry import METRICS
+
+
+ @METRICS.register_module()
+ class CustomMetric(BaseMetric):
+
+ def __init__(self, arg1, arg2):
+ """
+ The metric first processes each batch of data_samples and predictions,
+ and appends the processed results to the results list. Then it
+ collects all results together from all ranks if distributed training
+ is used. Finally, it computes the metrics of the entire dataset.
+ """
+
+ def process(self, data_batch: dict, data_samples: Sequence[dict]) -> None:
+ pass
+
+ def compute_metrics(self, results: list) -> dict:
+ pass
+
+ def evaluate(self, size: int) -> dict:
+ pass
+ ```
+
+ 在上面的示例中,`CustomMetric` 是 `BaseMetric` 的子类。它有三个方法:`process`,`compute_metrics` 和 `evaluate`。
+
+ - `process()` 处理一批数据样本和预测。处理后的结果需要显示地传给 `self.results` ,将在处理所有数据样本后用于计算指标。更多细节请参考 [MMEngine 文档](https://github.com/open-mmlab/mmengine/blob/main/docs/zh_cn/design/evaluation.md)
+
+ - `compute_metrics()` 用于从处理后的结果中计算指标。
+
+ - `evaluate()` 是一个接口,用于计算指标并返回结果。它将由 `ValLoop` 或 `TestLoop` 在 `Runner` 中调用。在大多数情况下,您不需要重写此方法,但如果您想做一些额外的工作,可以重写它。
+
+ **注意:** 您可以在[这里](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/loops.py#L366) 找到 `Runner` 调用 `evaluate()` 方法的过程。`Runner` 是训练和测试过程的执行器,您可以在[训练引擎文档](./engine.md)中找到有关它的详细信息。
+
+2. 在 `mmseg/evaluation/metrics/__init__.py` 中导入新的指标。
+
+ ```python
+ from .custom_metric import CustomMetric
+ __all__ = ['CustomMetric', ...]
+ ```
+
+3. 在配置文件中设置新的评测指标
+
+ ```python
+ val_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ test_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ ```
+
+## 使用发布版本的 MMSegmentation 进行开发
+
+上面的示例展示了如何使用 MMSegmentation 的源代码开发新指标。如果您想使用 MMSegmentation 的发布版本开发新指标,可以按照以下步骤操作。
+
+1. 创建一个新文件 `/Path/to/metrics/custom_metric.py`,实现 `process`,`compute_metrics` 和 `evaluate` 方法,`evaluate` 方法是可选的。
+
+2. 在代码或配置文件中导入新的指标。
+
+ ```python
+ from path.to.metrics import CustomMetric
+ ```
+
+ 或者
+
+ ```python
+ custom_imports = dict(imports=['/Path/to/metrics'], allow_failed_imports=False)
+
+ val_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ test_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ ```
diff --git a/docs/zh_cn/advanced_guides/add_models.md b/docs/zh_cn/advanced_guides/add_models.md
index 2f0a5af0d18..e05c07c8bac 100644
--- a/docs/zh_cn/advanced_guides/add_models.md
+++ b/docs/zh_cn/advanced_guides/add_models.md
@@ -166,7 +166,7 @@ loss_decode=dict(type='MyLoss', loss_weight=1.0))
### 添加新的数据预处理器(data preprocessor)
-在 MMSegmentation 1.x 版本中,我们使用 [SegDataPreProcessor](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/data_preprocessor.py#L13) 将数据复制到目标设备,并将数据预处理为默认的模型输入格式。这里我们将展示如何开发一个新的数据预处理器。
+在 MMSegmentation 1.x 版本中,我们使用 [SegDataPreProcessor](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/data_preprocessor.py#L13) 将数据复制到目标设备,并将数据预处理为默认的模型输入格式。这里我们将展示如何开发一个新的数据预处理器。
1. 创建一个新文件 `mmseg/models/my_datapreprocessor.py`。
diff --git a/docs/zh_cn/advanced_guides/contribute_dataset.md b/docs/zh_cn/advanced_guides/contribute_dataset.md
new file mode 100644
index 00000000000..4222de32a6c
--- /dev/null
+++ b/docs/zh_cn/advanced_guides/contribute_dataset.md
@@ -0,0 +1,461 @@
+# 在 mmsegmentation projects 中贡献一个标准格式的数据集
+
+- 在开始您的贡献流程前,请先阅读[《OpenMMLab 贡献代码指南》](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html),以详细的了解 OpenMMLab 代码库的代码贡献流程。
+- 该教程以 [Gaofen Image Dataset (GID)](https://www.sciencedirect.com/science/article/pii/S0034425719303414) 高分 2 号卫星所拍摄的遥感图像语义分割数据集作为样例,来演示在 mmsegmentation 中的数据集贡献流程。
+
+## 步骤 1: 配置 mmsegmentation 开发所需必要环境
+
+- 开发所必需的环境安装请参考[中文快速入门指南](https://github.com/open-mmlab/mmsegmentation/blob/main/docs/zh_cn/get_started.md)或[英文 get_started](https://github.com/open-mmlab/mmsegmentation/blob/main/docs/en/get_started.md)。
+
+- 如果您已安装了最新版的 pytorch、mmcv、mmengine,那么您可以跳过步骤 1 至[步骤 2](<#[步骤-2](#%E6%AD%A5%E9%AA%A4-2%E4%BB%A3%E7%A0%81%E8%B4%A1%E7%8C%AE%E5%89%8D%E7%9A%84%E5%87%86%E5%A4%87%E5%B7%A5%E4%BD%9C)>)。
+
+- **注:** 在此处无需安装 mmsegmentation,只需安装开发 mmsegmentation 所必需的 pytorch、mmcv、mmengine 等即可。
+
+**新建虚拟环境(如已有合适的开发环境,可跳过)**
+
+- 从[官方网站](https://docs.conda.io/en/latest/miniconda.html)下载并安装 Miniconda
+- 创建一个 conda 环境,并激活
+
+```shell
+conda create --name openmmlab python=3.8 -y
+conda activate openmmlab
+```
+
+**安装 pytorch (如环境下已安装 pytorch,可跳过)**
+
+- 参考 [official instructions](https://pytorch.org/get-started/locally/) 安装 **PyTorch**
+
+**使用 mim 安装 mmcv、mmengine**
+
+- 使用 [MIM](https://github.com/open-mmlab/mim) 安装 [MMCV](https://github.com/open-mmlab/mmcv)
+
+```shell
+pip install -U openmim
+mim install mmengine
+mim install "mmcv>=2.0.0"
+```
+
+## 步骤 2:代码贡献前的准备工作
+
+### 2.1 Fork mmsegmentation 仓库
+
+- 通过浏览器打开[mmsegmentation 官方仓库](https://github.com/open-mmlab/mmsegmentation/tree/main)。
+- 登录您的 GitHub 账户,以下步骤均需在 GitHub 登录的情况下进行。
+- Fork mmsegmentation 仓库
+ 
+- Fork 之后,mmsegmentation 仓库将会出现在您的个人仓库中。
+
+### 2.2 在您的代码编写软件中 git clone mmsegmentation
+
+这里以 VSCODE 为例
+
+- 打开 VSCODE,新建终端窗口并激活您在[步骤 1 ](#%E6%AD%A5%E9%AA%A4-1-%E9%85%8D%E7%BD%AE-mmsegmentation-%E5%BC%80%E5%8F%91%E6%89%80%E9%9C%80%E5%BF%85%E8%A6%81%E7%8E%AF%E5%A2%83)中所安装的虚拟环境。
+- 在您 GitHub 的个人仓库中找到您 Fork 的 mmsegmentation 仓库,复制其链接。
+ 
+- 在终端中执行命令
+ ```bash
+ git clone {您所复制的个人仓库的链接}
+ ```
+ 
+ **注:** 如提示以下信息,请在 GitHub 中添加 [SSH 秘钥](https://docs.github.com/en/authentication/connecting-to-github-with-ssh/generating-a-new-ssh-key-and-adding-it-to-the-ssh-agent)
+ 
+- 进入 mmsegmentation 目录(之后的操作均在 mmsegmentation 目录下)。
+ ```bash
+ cd mmsegmentation
+ ```
+- 在终端中执行以下命令,添加官方仓库为上游仓库。
+ ```bash
+ git remote add upstream git@github.com:open-mmlab/mmsegmentation.git
+ ```
+- 使用以下命令检查 remote 是否添加成功。
+ ```bash
+ git remote -v
+ ```
+ 
+
+### 2.3 切换目录至 mmsegmentation 并从源码安装mmsegmentation
+
+在`mmsegmentation`目录下执行`pip install -v -e .`,通过源码构建方式安装 mmsegmentaion 库。
+安装完成后,您将能看到如下图所示的文件树。
+

+
+### 2.4 切换分支为 dev-1.x
+
+正如您在[ mmsegmentation 官网](https://github.com/open-mmlab/mmsegmentation/tree/main)所见,该仓库有许多分支,默认分支`main`为稳定的发行版本,以及用于贡献者进行开发的`dev-1.x`分支。`dev-1.x`分支是贡献者们用来提交创意和 PR 的分支,`dev-1.x`分支的内容会被周期性的合入到`main`分支。
+
+
+回到 VSCODE 中,在终端执行命令
+
+```bash
+git checkout dev-1.x
+```
+
+### 2.5 创新属于自己的新分支
+
+在基于`dev-1.x`分支下,使用如下命令,创建属于您自己的分支。
+
+```bash
+# git checkout -b 您的GitHubID/您的分支想要实现的功能的名字
+# git checkout -b AI-Tianlong/support_GID_dataset
+git checkout -b {您的GitHubID/您的分支想要实现的功能的名字}
+```
+
+### 2.6 配置 pre-commit
+
+OpenMMLab 仓库对代码质量有着较高的要求,所有提交的 PR 必须要通过代码格式检查。pre-commit 详细配置参阅[配置 pre-commit](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html#pre-commit)。
+
+## 步骤 3:在`mmsegmentation/projects`下贡献您的代码
+
+**先对 GID 数据集进行分析**
+
+这里以贡献高分 2 号遥感图像语义分割数据集 GID 为例,GID 数据集是由我国自主研发的高分 2 号卫星所拍摄的光学遥感图像所创建,经图像预处理后共提供了 150 张 6800x7200 像素的 RGB 三通道遥感图像。并提供了两种不同类别数的数据标注,一种是包含 5 类有效物体的 RGB 标签,另一种是包含 15 类有效物体的 RGB 标签。本教程将针对 5 类标签进行数据集贡献流程讲解。
+
+GID 的 5 类有效标签分别为:0-背景-\[0,0,0\](mask 标签值-标签名称-RGB 标签值)、1-建筑-\[255,0,0\]、2-农田-\[0,255,0\]、3-森林-\[0,0,255\]、4-草地-\[255,255,0\]、5-水-\[0,0,255\]。在语义分割任务中,标签是与原图尺寸一致的单通道图像,标签图像中的像素值为真实样本图像中对应像素所包含的物体的类别。GID 数据集提供的是具有 RGB 三通道的彩色标签,为了模型的训练需要将 RGB 标签转换为 mask 标签。并且由于图像尺寸为 6800x7200 像素,对于神经网络的训练来有些过大,所以将每张图像裁切成了没有重叠的 512x512 的图像以便进行训练。
+

+
+### 3.1 在`mmsegmentation/projects`下创建新的项目文件夹
+
+在`mmsegmentation/projects`下创建文件夹`gid_dataset`
+
+
+### 3.2 贡献您的数据集代码
+
+为了最终能将您在 projects 中贡献的代码更加顺畅的移入核心库中(对代码要求质量更高),非常建议按照核心库的目录来编辑您的数据集文件。
+关于数据集有 4 个必要的文件:
+
+- **1** `mmseg/datasets/gid.py` 定义了数据集的尾缀、CLASSES、PALETTE、reduce_zero_label等
+- **2** `configs/_base_/gid.py` GID 数据集的配置文件,定义了数据集的`dataset_type`(数据集类型,`mmseg/datasets/gid.py`中注册的数据集的类名)、`data_root`(数据集所在的根目录,建议将数据集通过软连接的方式将数据集放至`mmsegmentation/data`)、`train_pipline`(训练的数据流)、`test_pipline`(测试和验证时的数据流)、`img_rations`(多尺度预测时的多尺度配置)、`tta_pipeline`(多尺度预测)、`train_dataloader`(训练集的数据加载器)、`val_dataloader`(验证集的数据加载器)、`test_dataloader`(测试集的数据加载器)、`val_evaluator`(验证集的评估器)、`test_evaluator`(测试集的评估器)。
+- **3** 使用了 GID 数据集的模型训练配置文件
+ 这个是可选的,但是强烈建议您添加。在核心库中,所贡献的数据集需要和参考文献中所提出的结果精度对齐,为了后期将您贡献的代码合并入核心库。如您的算力充足,最好能提供对应的模型配置文件在您贡献的数据集上所验证的结果以及相应的权重文件,并撰写较为详细的README.md文档。[示例参考结果](https://github.com/open-mmlab/mmsegmentation/tree/main/configs/deeplabv3plus#mapillary-vistas-v12)
+ 
+- **4** 使用如下命令格式: 撰写`docs/zh_cn/user_guides/2_dataset_prepare.md`来添加您的数据集介绍,包括但不限于数据集的下载方式,数据集目录结构、数据集生成等一些必要性的文字性描述和运行命令。以更好地帮助用户能更快的实现数据集的准备工作。
+
+### 3.3 贡献`tools/dataset_converters/gid.py`
+
+由于 GID 数据集是由未经过切分的 6800x7200 图像所构成的数据集,并且没有划分训练集、验证集与测试集。以及其标签为 RGB 彩色标签,需要将标签转换为单通道的 mask label。为了方便训练,首先将 GID 数据集进行裁切和标签转换,并进行数据集划分,构建为 mmsegmentation 所支持的格式。
+
+```python
+# tools/dataset_converters/gid.py
+import argparse
+import glob
+import math
+import os
+import os.path as osp
+from PIL import Image
+
+import mmcv
+import numpy as np
+from mmengine.utils import ProgressBar, mkdir_or_exist
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='Convert GID dataset to mmsegmentation format')
+ parser.add_argument('dataset_img_path', help='GID images folder path')
+ parser.add_argument('dataset_label_path', help='GID labels folder path')
+ parser.add_argument('--tmp_dir', help='path of the temporary directory')
+ parser.add_argument('-o', '--out_dir', help='output path', default='data/gid')
+ parser.add_argument(
+ '--clip_size',
+ type=int,
+ help='clipped size of image after preparation',
+ default=256)
+ parser.add_argument(
+ '--stride_size',
+ type=int,
+ help='stride of clipping original images',
+ default=256)
+ args = parser.parse_args()
+ return args
+
+GID_COLORMAP = dict(
+ Background=(0, 0, 0), #0-背景-黑色
+ Building=(255, 0, 0), #1-建筑-红色
+ Farmland=(0, 255, 0), #2-农田-绿色
+ Forest=(0, 0, 255), #3-森林-蓝色
+ Meadow=(255, 255, 0),#4-草地-黄色
+ Water=(0, 0, 255)#5-水-蓝色
+)
+palette = list(GID_COLORMAP.values())
+classes = list(GID_COLORMAP.keys())
+
+#############用列表来存一个 RGB 和一个类别的对应################
+def colormap2label(palette):
+ colormap2label_list = np.zeros(256**3, dtype = np.longlong)
+ for i, colormap in enumerate(palette):
+ colormap2label_list[(colormap[0] * 256 + colormap[1])*256+colormap[2]] = i
+ return colormap2label_list
+
+#############给定那个列表,和vis_png然后生成masks_png################
+def label_indices(RGB_label, colormap2label_list):
+ RGB_label = RGB_label.astype('int32')
+ idx = (RGB_label[:, :, 0] * 256 + RGB_label[:, :, 1]) * 256 + RGB_label[:, :, 2]
+ # print(idx.shape)
+ return colormap2label_list[idx]
+
+def RGB2mask(RGB_label, colormap2label_list):
+ # RGB_label = np.array(Image.open(RGB_label).convert('RGB')) #打开RGB_png
+ mask_label = label_indices(RGB_label, colormap2label_list) # .numpy()
+ return mask_label
+
+colormap2label_list = colormap2label(palette)
+
+def clip_big_image(image_path, clip_save_dir, args, to_label=False):
+ """
+ Original image of GID dataset is very large, thus pre-processing
+ of them is adopted. Given fixed clip size and stride size to generate
+ clipped image, the intersection of width and height is determined.
+ For example, given one 6800 x 7200 original image, the clip size is
+ 256 and stride size is 256, thus it would generate 29 x 27 = 783 images
+ whose size are all 256 x 256.
+
+ """
+
+ image = mmcv.imread(image_path, channel_order='rgb')
+ # image = mmcv.bgr2gray(image)
+
+ h, w, c = image.shape
+ clip_size = args.clip_size
+ stride_size = args.stride_size
+
+ num_rows = math.ceil((h - clip_size) / stride_size) if math.ceil(
+ (h - clip_size) /
+ stride_size) * stride_size + clip_size >= h else math.ceil(
+ (h - clip_size) / stride_size) + 1
+ num_cols = math.ceil((w - clip_size) / stride_size) if math.ceil(
+ (w - clip_size) /
+ stride_size) * stride_size + clip_size >= w else math.ceil(
+ (w - clip_size) / stride_size) + 1
+
+ x, y = np.meshgrid(np.arange(num_cols + 1), np.arange(num_rows + 1))
+ xmin = x * clip_size
+ ymin = y * clip_size
+
+ xmin = xmin.ravel()
+ ymin = ymin.ravel()
+ xmin_offset = np.where(xmin + clip_size > w, w - xmin - clip_size,
+ np.zeros_like(xmin))
+ ymin_offset = np.where(ymin + clip_size > h, h - ymin - clip_size,
+ np.zeros_like(ymin))
+ boxes = np.stack([
+ xmin + xmin_offset, ymin + ymin_offset,
+ np.minimum(xmin + clip_size, w),
+ np.minimum(ymin + clip_size, h)
+ ], axis=1)
+
+ if to_label:
+ image = RGB2mask(image, colormap2label_list) #这里得改一下
+
+ for count, box in enumerate(boxes):
+ start_x, start_y, end_x, end_y = box
+ clipped_image = image[start_y:end_y,
+ start_x:end_x] if to_label else image[
+ start_y:end_y, start_x:end_x, :]
+ img_name = osp.basename(image_path).replace('.tif', '')
+ img_name = img_name.replace('_label', '')
+ if count % 3 == 0:
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(
+ clip_save_dir.replace('train', 'val'),
+ f'{img_name}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+ else:
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(
+ clip_save_dir,
+ f'{img_name}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+ count += 1
+
+def main():
+ args = parse_args()
+
+ """
+ According to this paper: https://ieeexplore.ieee.org/document/9343296/
+ select 15 images contained in GID, , which cover the whole six
+ categories, to generate train set and validation set.
+
+ According to Paper: https://ieeexplore.ieee.org/document/9343296/
+
+ """
+
+ if args.out_dir is None:
+ out_dir = osp.join('data', 'gid')
+ else:
+ out_dir = args.out_dir
+
+ print('Making directories...')
+ mkdir_or_exist(osp.join(out_dir, 'img_dir', 'train'))
+ mkdir_or_exist(osp.join(out_dir, 'img_dir', 'val'))
+ mkdir_or_exist(osp.join(out_dir, 'ann_dir', 'train'))
+ mkdir_or_exist(osp.join(out_dir, 'ann_dir', 'val'))
+
+ src_path_list = glob.glob(os.path.join(args.dataset_img_path, '*.tif'))
+ print(f'Find {len(src_path_list)} pictures')
+
+ prog_bar = ProgressBar(len(src_path_list))
+
+ dst_img_dir = osp.join(out_dir, 'img_dir', 'train')
+ dst_label_dir = osp.join(out_dir, 'ann_dir', 'train')
+
+ for i, img_path in enumerate(src_path_list):
+ label_path = osp.join(args.dataset_label_path, osp.basename(img_path.replace('.tif', '_label.tif')))
+
+ clip_big_image(img_path, dst_img_dir, args, to_label=False)
+ clip_big_image(label_path, dst_label_dir, args, to_label=True)
+ prog_bar.update()
+
+ print('Done!')
+
+if __name__ == '__main__':
+ main()
+```
+
+### 3.4 贡献`mmseg/datasets/gid.py`
+
+可参考[`projects/mapillary_dataset/mmseg/datasets/mapillary.py`](https://github.com/open-mmlab/mmsegmentation/blob/main/projects/mapillary_dataset/mmseg/datasets/mapillary.py)并在此基础上修改相应变量以适配您的数据集。
+
+```python
+# mmseg/datasets/gid.py
+# Copyright (c) OpenMMLab. All rights reserved.
+from mmseg.datasets.basesegdataset import BaseSegDataset
+from mmseg.registry import DATASETS
+
+# 注册数据集类
+@DATASETS.register_module()
+class GID_Dataset(BaseSegDataset):
+ """Gaofen Image Dataset (GID)
+
+ Dataset paper link:
+ https://www.sciencedirect.com/science/article/pii/S0034425719303414
+ https://x-ytong.github.io/project/GID.html
+
+ GID 6 classes: background(others), built-up, farmland, forest, meadow, water
+
+ In This example, select 10 images from GID dataset as training set,
+ and select 5 images as validation set.
+ The selected images are listed as follows:
+
+ GF2_PMS1__L1A0000647767-MSS1
+ GF2_PMS1__L1A0001064454-MSS1
+ GF2_PMS1__L1A0001348919-MSS1
+ GF2_PMS1__L1A0001680851-MSS1
+ GF2_PMS1__L1A0001680853-MSS1
+ GF2_PMS1__L1A0001680857-MSS1
+ GF2_PMS1__L1A0001757429-MSS1
+ GF2_PMS2__L1A0000607681-MSS2
+ GF2_PMS2__L1A0000635115-MSS2
+ GF2_PMS2__L1A0000658637-MSS2
+ GF2_PMS2__L1A0001206072-MSS2
+ GF2_PMS2__L1A0001471436-MSS2
+ GF2_PMS2__L1A0001642620-MSS2
+ GF2_PMS2__L1A0001787089-MSS2
+ GF2_PMS2__L1A0001838560-MSS2
+
+ The ``img_suffix`` is fixed to '.tif' and ``seg_map_suffix`` is
+ fixed to '.tif' for GID.
+ """
+ METAINFO = dict(
+ classes=('Others', 'Built-up', 'Farmland', 'Forest',
+ 'Meadow', 'Water'),
+
+ palette=[[0, 0, 0], [255, 0, 0], [0, 255, 0], [0, 255, 255],
+ [255, 255, 0], [0, 0, 255]])
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=None,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
+```
+
+### 3.5 贡献使用 GID 的训练 config file
+
+```python
+_base_ = [
+ '../../../configs/_base_/models/deeplabv3plus_r50-d8.py',
+ './_base_/datasets/gid.py',
+ '../../../configs/_base_/default_runtime.py',
+ '../../../configs/_base_/schedules/schedule_240k.py'
+]
+custom_imports = dict(
+ imports=['projects.gid_dataset.mmseg.datasets.gid'])
+
+crop_size = (256, 256)
+data_preprocessor = dict(size=crop_size)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ pretrained='open-mmlab://resnet101_v1c',
+ backbone=dict(depth=101),
+ decode_head=dict(num_classes=6),
+ auxiliary_head=dict(num_classes=6))
+
+```
+
+### 3.6 撰写`docs/zh_cn/user_guides/2_dataset_prepare.md`
+
+**Gaofen Image Dataset (GID)**
+
+- GID 数据集可在[此处](https://x-ytong.github.io/project/Five-Billion-Pixels.html)进行下载。
+- GID 数据集包含 150 张 6800x7200 的大尺寸图像,标签为 RGB 标签。
+- 此处选择 15 张图像生成训练集和验证集,该 15 张图像包含了所有六类信息。所选的图像名称如下:
+
+```None
+ GF2_PMS1__L1A0000647767-MSS1
+ GF2_PMS1__L1A0001064454-MSS1
+ GF2_PMS1__L1A0001348919-MSS1
+ GF2_PMS1__L1A0001680851-MSS1
+ GF2_PMS1__L1A0001680853-MSS1
+ GF2_PMS1__L1A0001680857-MSS1
+ GF2_PMS1__L1A0001757429-MSS1
+ GF2_PMS2__L1A0000607681-MSS2
+ GF2_PMS2__L1A0000635115-MSS2
+ GF2_PMS2__L1A0000658637-MSS2
+ GF2_PMS2__L1A0001206072-MSS2
+ GF2_PMS2__L1A0001471436-MSS2
+ GF2_PMS2__L1A0001642620-MSS2
+ GF2_PMS2__L1A0001787089-MSS2
+ GF2_PMS2__L1A0001838560-MSS2
+```
+
+执行以下命令进行裁切及标签的转换,需要修改为您所存储 15 张图像及标签的路径。
+
+```
+python projects/gid_dataset/tools/dataset_converters/gid.py [15 张图像的路径] [15 张标签的路径]
+```
+
+完成裁切后的 GID 数据结构如下:
+
+```none
+mmsegmentation
+├── mmseg
+├── tools
+├── configs
+├── data
+│ ├── gid
+│ │ ├── ann_dir
+| │ │ │ ├── train
+| │ │ │ ├── val
+│ │ ├── img_dir
+| │ │ │ ├── train
+| │ │ │ ├── val
+
+```
+
+### 3.7 贡献的代码及文档通过`pre-commit`检查
+
+使用命令
+
+```bash
+git add .
+git commit -m "添加描述"
+git push
+```
+
+### 3.8 在 GitHub 中向 mmsegmentation 提交 PR
+
+具体步骤可见[《OpenMMLab 贡献代码指南》](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html)
diff --git a/docs/zh_cn/advanced_guides/data_flow.md b/docs/zh_cn/advanced_guides/data_flow.md
index 0716d36d1b4..20dbe07e75d 100644
--- a/docs/zh_cn/advanced_guides/data_flow.md
+++ b/docs/zh_cn/advanced_guides/data_flow.md
@@ -16,7 +16,7 @@ val_cfg = dict(type='ValLoop')
test_cfg = dict(type='TestLoop')
```
-在上图中,红色线表示 [train_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#train_step) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#train_step))*** ,在每次训练迭代中,数据加载器(dataloader)从存储中加载图像并传输到数据预处理器(data preprocessor),数据预处理器会将图像放到特定的设备上,并将数据堆叠到批处理中,之后模型接受批处理数据作为输入,最后将模型的输出发送给优化器(optimizer)。蓝色线表示 [val_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#val_step) 和 [test_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#test_step) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#test_step))*** 。这两个过程的数据流除了模型输出与 `train_step` 不同外,其余均和 `train_step` 类似。由于在评估时模型参数会被冻结,因此模型的输出将被传递给 [Evaluator](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/evaluation.md#ioumetric) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/evaluation.md#ioumetric))***
+在上图中,红色线表示 [train_step](./models.md#train_step),在每次训练迭代中,数据加载器(dataloader)从存储中加载图像并传输到数据预处理器(data preprocessor),数据预处理器会将图像放到特定的设备上,并将数据堆叠到批处理中,之后模型接受批处理数据作为输入,最后将模型的输出发送给优化器(optimizer)。蓝色线表示 [val_step](./models.md#val_step) 和 [test_step](./models.md#test_step)。这两个过程的数据流除了模型输出与 `train_step` 不同外,其余均和 `train_step` 类似。由于在评估时模型参数会被冻结,因此模型的输出将被传递给 [Evaluator](./evaluation.md#ioumetric)。
来计算指标。
## MMSegmentation 中的数据流约定
@@ -28,7 +28,7 @@ test_cfg = dict(type='TestLoop')
数据加载器(DataLoader)是 MMEngine 的训练和测试流程中的一个重要组件。
从概念上讲,它源于 [PyTorch](https://pytorch.org/) 并保持一致。DataLoader 从文件系统加载数据,原始数据通过数据准备流程后被发送给数据预处理器。
-MMSegmentation 在 [PackSegInputs](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/transforms/formatting.py#L12) 中定义了默认数据格式, 它是 `train_pipeline` 和 `test_pipeline` 的最后一个组件。有关数据转换 `pipeline` 的更多信息,请参阅[数据转换文档](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/transforms.html)。 ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/transforms.html))***
+MMSegmentation 在 [PackSegInputs](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/transforms/formatting.py#L12) 中定义了默认数据格式, 它是 `train_pipeline` 和 `test_pipeline` 的最后一个组件。有关数据转换 `pipeline` 的更多信息,请参阅[数据转换文档](./transforms.md)。
在没有任何修改的情况下,PackSegInputs 的返回值通常是一个包含 `inputs` 和 `data_samples` 的 `dict`。以下伪代码展示了 mmseg 中数据加载器输出的数据类型,它是从数据集中获取的一批数据样本,数据加载器将它们打包成一个字典列表。`inputs` 是输入进模型的张量列表,`data_samples` 包含了输入图像的 meta information 和相应的 ground truth。
@@ -39,11 +39,11 @@ dict(
)
```
-**注意:** [SegDataSample](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/structures/seg_data_sample.py) 是 MMSegmentation 的数据结构接口,用于连接不同组件。`SegDataSample` 实现了抽象数据元素 `mmengine.structures.BaseDataElement`,更多信息请在 [MMEngine](https://github.com/open-mmlab/mmengine) 中参阅 [SegDataSample 文档](https://mmsegmentation.readthedocs.io/zh_CN/1.x/advanced_guides/structures.html)和[数据元素文档](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/data_element.html)。
+**注意:** [SegDataSample](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/structures/seg_data_sample.py) 是 MMSegmentation 的数据结构接口,用于连接不同组件。`SegDataSample` 实现了抽象数据元素 `mmengine.structures.BaseDataElement`,更多信息请在 [MMEngine](https://github.com/open-mmlab/mmengine) 中参阅 [SegDataSample 文档](./structures.md)和[数据元素文档](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/data_element.html)。
### 数据预处理器到模型
-虽然在[上面的图](##数据流概述)中分开绘制了数据预处理器和模型,但数据预处理器是模型的一部分,因此可以在[模型教程](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/models.html)中找到数据预处理器章节。 ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/models.html))***
+虽然在[上面的图](##数据流概述)中分开绘制了数据预处理器和模型,但数据预处理器是模型的一部分,因此可以在[模型教程](./models.md)中找到数据预处理器章节。
数据预处理器的返回值是一个包含 `inputs` 和 `data_samples` 的字典,其中 `inputs` 是批处理图像的 4D 张量,`data_samples` 中添加了一些用于数据预处理的额外元信息。当传递给网络时,字典将被解包为两个值。 以下伪代码展示了数据预处理器的返回值和模型的输入值。
@@ -61,22 +61,22 @@ class Network(BaseSegmentor):
pass
```
-**注意:** 模型的前向传播有 3 种模式,由输入参数 mode 控制,更多信息请参阅[模型教程](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md)。 ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md))***
+**注意:** 模型的前向传播有 3 种模式,由输入参数 mode 控制,更多信息请参阅[模型教程](./models.md)。
### 模型输出
-如[模型教程](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#forward) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#forward))*** 所提到的 3 种前向传播具有 3 种输出。
+如[模型教程](./models.md#forward) ***([中文链接待更新](./models.md#forward))*** 所提到的 3 种前向传播具有 3 种输出。
`train_step` 和 `test_step`(或 `val_step`)分别对应于 `'loss'` 和 `'predict'`。
-在 `test_step` 或 `val_step` 中,推理结果会被传递给 `Evaluator` 。您可以参阅[评估文档](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/evaluation.html) ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/evaluation.html))*** 来获取更多关于 `Evaluator` 的信息。
+在 `test_step` 或 `val_step` 中,推理结果会被传递给 `Evaluator` 。您可以参阅[评估文档](./evaluation.md)来获取更多关于 `Evaluator` 的信息。
-在推理后,MMSegmentation 中的 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/segmentors/base.py#L15) 会对推理结果进行简单的后处理以打包推理结果。神经网络生成的分割 logits,经过 `argmax` 操作后的分割 mask 和 ground truth(如果存在)将被打包到类似 `SegDataSample` 的实例。 [postprocess_result](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/segmentors/base.py#L132) 的返回值是一个 **`SegDataSample`的`List`**。下图显示了这些 `SegDataSample` 实例的关键属性。
+在推理后,MMSegmentation 中的 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/segmentors/base.py#L15) 会对推理结果进行简单的后处理以打包推理结果。神经网络生成的分割 logits,经过 `argmax` 操作后的分割 mask 和 ground truth(如果存在)将被打包到类似 `SegDataSample` 的实例。 [postprocess_result](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/segmentors/base.py#L132) 的返回值是一个 **`SegDataSample`的`List`**。下图显示了这些 `SegDataSample` 实例的关键属性。

-与数据预处理器一致,损失函数也是模型的一部分,它是[解码头](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/decode_heads/decode_head.py#L142)的属性之一。
+与数据预处理器一致,损失函数也是模型的一部分,它是[解码头](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/decode_heads/decode_head.py#L142)的属性之一。
-在 MMSegmentation 中,`decode_head` 的 [loss_by_feat](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/decode_heads/decode_head.py#L291) 方法是用于计算损失的统一接口。
+在 MMSegmentation 中,`decode_head` 的 [loss_by_feat](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/decode_heads/decode_head.py#L291) 方法是用于计算损失的统一接口。
参数:
@@ -87,4 +87,4 @@ class Network(BaseSegmentor):
- dict\[str, Tensor\]:一个损失组件的字典
-**注意:** `train_step` 将损失传递进 OptimWrapper 以更新模型中的权重,更多信息请参阅 [train_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#train_step)。 ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#train_step))***
+**注意:** `train_step` 将损失传递进 OptimWrapper 以更新模型中的权重,更多信息请参阅 [train_step](./models.md#train_step)。
diff --git a/docs/zh_cn/advanced_guides/datasets.md b/docs/zh_cn/advanced_guides/datasets.md
index 546e97f70d6..b45f2d22bb6 100644
--- a/docs/zh_cn/advanced_guides/datasets.md
+++ b/docs/zh_cn/advanced_guides/datasets.md
@@ -1,10 +1,10 @@
# 数据集
-在 MMSegmentation 算法库中, 所有 Dataset 类的功能有两个: 加载[预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/2_dataset_prepare.md) 之后的数据集的信息, 和将数据送入[数据集变换流水线](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/basesegdataset.py#L141) 中, 进行[数据变换操作](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/transforms.md). 加载的数据集信息包括两类: 元信息 (meta information), 数据集本身的信息, 例如数据集总共的类别, 和它们对应调色盘信息: 数据信息 (data information) 是指每组数据中图片和对应标签的路径. 下文中介绍了 MMSegmentation 1.x 中数据集的常用接口, 和 mmseg 数据集基类中数据信息加载与修改数据集类别的逻辑, 以及数据集与数据变换流水线 (pipeline) 的关系.
+在 MMSegmentation 算法库中, 所有 Dataset 类的功能有两个: 加载[预处理](../user_guides/2_dataset_prepare.md) 之后的数据集的信息, 和将数据送入[数据集变换流水线](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L141) 中, 进行[数据变换操作](./transforms.md). 加载的数据集信息包括两类: 元信息 (meta information), 数据集本身的信息, 例如数据集总共的类别, 和它们对应调色盘信息: 数据信息 (data information) 是指每组数据中图片和对应标签的路径. 下文中介绍了 MMSegmentation 1.x 中数据集的常用接口, 和 mmseg 数据集基类中数据信息加载与修改数据集类别的逻辑, 以及数据集与数据变换流水线 (pipeline) 的关系.
## 常用接口
-以 Cityscapes 为例, 介绍数据集常用接口. 如需运行以下示例, 请在当前工作目录下的 `data` 目录下载并[预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/2_dataset_prepare.md#cityscapes) Cityscapes 数据集.
+以 Cityscapes 为例, 介绍数据集常用接口. 如需运行以下示例, 请在当前工作目录下的 `data` 目录下载并[预处理](../user_guides/2_dataset_prepare.md#cityscapes) Cityscapes 数据集.
实例化 Cityscapes 训练数据集:
@@ -96,7 +96,7 @@ print(dataset.metainfo)
'reduce_zero_label': False}
```
-数据集 `__getitem__` 方法的返回值, 是经过数据增强的样本数据的输出, 同样也是一个字典, 包括两个字段, `'inputs'` 字段是当前样本经过数据增强操作的图像, 类型为 torch.Tensor, `'data_samples'` 字段存放的数据类型是 MMSegmentation 1.x 新添加的数据结构 [`Segdatasample`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/structures.md), 其中`gt_sem_seg` 字段是经过数据增强的标签数据.
+数据集 `__getitem__` 方法的返回值, 是经过数据增强的样本数据的输出, 同样也是一个字典, 包括两个字段, `'inputs'` 字段是当前样本经过数据增强操作的图像, 类型为 torch.Tensor, `'data_samples'` 字段存放的数据类型是 MMSegmentation 1.x 新添加的数据结构 [`Segdatasample`](./structures.md), 其中`gt_sem_seg` 字段是经过数据增强的标签数据.
```python
print(dataset[0])
@@ -166,13 +166,13 @@ print(dataset[0])
## BaseSegDataset
-由于 MMSegmentation 中的所有数据集的基本功能均包括(1) 加载[数据集预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/user_guides/2_dataset_prepare.md) 之后的数据信息和 (2) 将数据送入数据变换流水线中进行数据变换, 因此在 MMSegmentation 中将其中的共同接口抽象成 [`BaseSegDataset`](https://mmsegmentation.readthedocs.io/en/dev-1.x/api.html?highlight=BaseSegDataset#mmseg.datasets.BaseSegDataset),它继承自 [MMEngine 的 `BaseDataset`](https://github.com/open-mmlab/mmengine/blob/main/docs/en/advanced_tutorials/basedataset.md), 遵循 OpenMMLab 数据集初始化统一流程, 支持高效的内部数据存储格式, 支持数据集拼接、数据集重复采样等功能.
+由于 MMSegmentation 中的所有数据集的基本功能均包括(1) 加载[数据集预处理](../user_guides/2_dataset_prepare.md) 之后的数据信息和 (2) 将数据送入数据变换流水线中进行数据变换, 因此在 MMSegmentation 中将其中的共同接口抽象成 [`BaseSegDataset`](https://mmsegmentation.readthedocs.io/zh_CN/latest/api.html?highlight=BaseSegDataset#mmseg.datasets.BaseSegDataset),它继承自 [MMEngine 的 `BaseDataset`](https://github.com/open-mmlab/mmengine/blob/main/docs/en/advanced_tutorials/basedataset.md), 遵循 OpenMMLab 数据集初始化统一流程, 支持高效的内部数据存储格式, 支持数据集拼接、数据集重复采样等功能.
在 MMSegmentation BaseSegDataset 中重新定义了**数据信息加载方法**(`load_data_list`)和并新增了 `get_label_map` 方法用来**修改数据集的类别信息**.
### 数据信息加载
-数据信息加载的内容是样本数据的图片路径和标签路径, 具体实现在 MMSegmentation 的 BaseSegDataset 的 [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/163277bfe0fa8fefb63ee5137917fafada1b301c/mmseg/datasets/basesegdataset.py#L231) 中.
-主要有两种获取图片和标签的路径方法, 如果当数据集目录按以下目录结构组织, [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/163277bfe0fa8fefb63ee5137917fafada1b301c/mmseg/datasets/basesegdataset.py#L231)) 会根据数据路径和后缀来解析.
+数据信息加载的内容是样本数据的图片路径和标签路径, 具体实现在 MMSegmentation 的 BaseSegDataset 的 [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L231) 中.
+主要有两种获取图片和标签的路径方法, 如果当数据集目录按以下目录结构组织, [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L231)) 会根据数据路径和后缀来解析.
```
├── data
@@ -322,7 +322,7 @@ print(dataset.metainfo)
'reduce_zero_label': False}
```
-可以看到, 数据集元信息的类别和默认 Cityscapes 不同. 并且, 定义了标签重映射的字段 `label_map` 用来修改每个分割掩膜上的像素的类别索引, 分割标签类别会根据 `label_map`, 将类别重映射, [具体实现](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/basesegdataset.py#L151):
+可以看到, 数据集元信息的类别和默认 Cityscapes 不同. 并且, 定义了标签重映射的字段 `label_map` 用来修改每个分割掩膜上的像素的类别索引, 分割标签类别会根据 `label_map`, 将类别重映射, [具体实现](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L151):
```python
gt_semantic_seg_copy = gt_semantic_seg.copy()
diff --git a/docs/zh_cn/advanced_guides/engine.md b/docs/zh_cn/advanced_guides/engine.md
index a5746fcec89..79b4c8d2296 100644
--- a/docs/zh_cn/advanced_guides/engine.md
+++ b/docs/zh_cn/advanced_guides/engine.md
@@ -61,21 +61,21 @@ OpenMMLab 将模型训练和测试过程抽象为 `Runner`, 插入钩子可以
- 默认钩子 (default hooks)
-它们实现了训练时所必需的功能, 在配置文件中用 `default_hooks` 定义传给 `Runner`, `Runner` 通过 [`register_default_hooks`](https://github.com/open-mmlab/mmengine/blob/090104df21acd05a8aadae5a0d743a7da3314f6f/mmengine/runner/runner.py#L1780) 方法注册.
+它们实现了训练时所必需的功能, 在配置文件中用 `default_hooks` 定义传给 `Runner`, `Runner` 通过 [`register_default_hooks`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/runner.py#L1780) 方法注册.
钩子有对应的优先级, 优先级越高, 越早被执行器调用. 如果优先级一样, 被调用的顺序和钩子注册的顺序一致.
不建议用户修改默认钩子的优先级, 可以参考 [mmengine hooks 文档](https://github.com/open-mmlab/mmengine/blob/main/docs/zh_cn/tutorials/hook.md) 了解钩子优先级的定义.
下面是 MMSegmentation 中所用到的默认钩子:
-| 钩子 | 功能 | 优先级 |
-| :-----------------------------------------------------------------------------------------------------------------------: | :---------------------------------------------------------------------------------------------: | :---------------: |
-| [IterTimerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/iter_timer_hook.py) | 记录 iteration 花费的时间. | NORMAL (50) |
-| [LoggerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/logger_hook.py) | 从 `Runner` 里不同的组件中收集日志记录, 并将其输出到终端, JSON 文件, tensorboard, wandb 等下游. | BELOW_NORMAL (60) |
-| [ParamSchedulerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/param_scheduler_hook.py) | 更新优化器里面的一些超参数, 例如学习率的动量. | LOW (70) |
-| [CheckpointHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py) | 规律性地保存 checkpoint 文件. | VERY_LOW (90) |
-| [DistSamplerSeedHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/sampler_seed_hook.py) | 确保分布式采样器 shuffle 是打开的. | NORMAL (50) |
-| [SegVisualizationHook](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/visualization/local_visualizer.py) | 可视化验证和测试过程里的预测结果. | NORMAL (50) |
+| 钩子 | 功能 | 优先级 |
+| :--------------------------------------------------------------------------------------------------------------------: | :---------------------------------------------------------------------------------------------: | :---------------: |
+| [IterTimerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/iter_timer_hook.py) | 记录 iteration 花费的时间. | NORMAL (50) |
+| [LoggerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/logger_hook.py) | 从 `Runner` 里不同的组件中收集日志记录, 并将其输出到终端, JSON 文件, tensorboard, wandb 等下游. | BELOW_NORMAL (60) |
+| [ParamSchedulerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/param_scheduler_hook.py) | 更新优化器里面的一些超参数, 例如学习率的动量. | LOW (70) |
+| [CheckpointHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py) | 规律性地保存 checkpoint 文件. | VERY_LOW (90) |
+| [DistSamplerSeedHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/sampler_seed_hook.py) | 确保分布式采样器 shuffle 是打开的. | NORMAL (50) |
+| [SegVisualizationHook](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/visualization/local_visualizer.py) | 可视化验证和测试过程里的预测结果. | NORMAL (50) |
-MMSegmentation 会在 [`defualt_hooks`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/schedules/schedule_160k.py#L19-L25) 里面注册一些训练所必需功能的钩子::
+MMSegmentation 会在 [`defualt_hooks`](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/schedules/schedule_160k.py#L19-L25) 里面注册一些训练所必需功能的钩子::
```python
default_hooks = dict(
@@ -94,6 +94,7 @@ default_hooks = dict(
以 `default_hooks` 里面的 `logger` 和 `checkpoint` 为例, 我们来介绍如何修改 `default_hooks` 中默认的钩子.
(1) 模型保存配置
+
`default_hooks` 使用 `checkpoint` 字段来初始化[模型保存钩子 (CheckpointHook)](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py#L19).
```python
@@ -104,6 +105,7 @@ checkpoint = dict(type='CheckpointHook', interval=1)
更多相关参数的细节可以参考[这里](https://mmengine.readthedocs.io/zh_CN/latest/api/generated/mmengine.hooks.CheckpointHook.html#checkpointhook).
(2) 日志配置
+
`日志钩子 (LoggerHook)` 被用来收集 `执行器 (Runner)` 里面不同组件的日志信息然后写入终端, JSON 文件, tensorboard 和 wandb 等地方.
```python
@@ -126,7 +128,7 @@ visualizer = dict(
- 自定义钩子 (custom hooks)
-自定义钩子在配置通过 `custom_hooks` 定义, `Runner` 通过 [`register_custom_hooks`](https://github.com/open-mmlab/mmengine/blob/090104df21acd05a8aadae5a0d743a7da3314f6f/mmengine/runner/runner.py#L1852) 方法注册.
+自定义钩子在配置通过 `custom_hooks` 定义, `Runner` 通过 [`register_custom_hooks`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/runner.py#L1820) 方法注册.
自定义钩子优先级需要在配置文件里设置, 如果没有设置, 则会被默认设置为 `NORMAL`. 下面是部分 MMEngine 中实现的自定义钩子:
| 钩子 | 用法 |
@@ -145,7 +147,7 @@ custom_hooks = [
### SegVisualizationHook
-MMSegmentation 实现了 [`SegVisualizationHook`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/engine/hooks/visualization_hook.py#L17), 用来在验证和测试时可视化预测结果.
+MMSegmentation 实现了 [`SegVisualizationHook`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/engine/hooks/visualization_hook.py#L17), 用来在验证和测试时可视化预测结果.
`SegVisualizationHook` 重写了基类 `Hook` 中的 `_after_iter` 方法, 在验证或测试时, 根据指定的迭代次数间隔调用 `visualizer` 的 `add_datasample` 方法绘制语义分割结果, 具体实现如下:
```python
@@ -181,7 +183,7 @@ class SegVisualizationHook(Hook):
```
-关于可视化更多的细节可以查看[这里](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/visualization.md).
+关于可视化更多的细节可以查看[这里](../user_guides/visualization.md).
## 优化器
@@ -234,7 +236,7 @@ optim_wrapper = dict(type='AmpOptimWrapper', optimizer=optimizer)
在模型训练中, 如果想在优化器里为不同参数分别设置优化策略, 例如设置不同的学习率、权重衰减等超参数, 可以通过设置配置文件里 `optim_wrapper` 中的 `paramwise_cfg` 来实现.
-下面的配置文件以 [ViT `optim_wrapper`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/vit/vit_vit-b16-ln_mln_upernet_8xb2-160k_ade20k-512x512.py#L15-L27) 为例介绍 `paramwise_cfg` 参数使用.
+下面的配置文件以 [ViT `optim_wrapper`](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/vit/vit_vit-b16-ln_mln_upernet_8xb2-160k_ade20k-512x512.py#L15-L27) 为例介绍 `paramwise_cfg` 参数使用.
训练时将 `pos_embed`, `mask_token`, `norm` 模块的 weight decay 参数的系数设置成 0.
即: 在训练时, 这些模块的 weight decay 将被变为 `weight_decay * decay_mult`=0.
@@ -257,9 +259,9 @@ optim_wrapper = dict(
### 优化器封装构造器
-默认的优化器封装构造器 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/376251961da47ea8254ab808ae5c51e1430f18dc/mmengine/optim/optimizer/default_constructor.py#L19) 根据输入的 `optim_wrapper` 和 `optim_wrapper` 中定义的 `paramwise_cfg` 来构建训练中使用的优化器. 当 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/376251961da47ea8254ab808ae5c51e1430f18dc/mmengine/optim/optimizer/default_constructor.py#L19) 功能不能满足需求时, 可以自定义优化器封装构造器来实现超参数的配置.
+默认的优化器封装构造器 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/optim/optimizer/default_constructor.py#L19) 根据输入的 `optim_wrapper` 和 `optim_wrapper` 中定义的 `paramwise_cfg` 来构建训练中使用的优化器. 当 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/optim/optimizer/default_constructor.py#L19) 功能不能满足需求时, 可以自定义优化器封装构造器来实现超参数的配置.
-MMSegmentation 中的实现了 [`LearningRateDecayOptimizerConstructor`](https://github.com/open-mmlab/mmsegmentation/blob/b21df463d47447f33c28d9a4f46136ad64d34a40/mmseg/engine/optimizers/layer_decay_optimizer_constructor.py#L104), 可以对以 ConvNeXt, BEiT 和 MAE 为骨干网络的模型训练时, 骨干网络的模型参数的学习率按照定义的衰减比例(`decay_rate`)逐层递减, 在配置文件中的配置如下:
+MMSegmentation 中的实现了 [`LearningRateDecayOptimizerConstructor`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/engine/optimizers/layer_decay_optimizer_constructor.py#L104), 可以对以 ConvNeXt, BEiT 和 MAE 为骨干网络的模型训练时, 骨干网络的模型参数的学习率按照定义的衰减比例(`decay_rate`)逐层递减, 在配置文件中的配置如下:
```python
optim_wrapper = dict(
diff --git a/docs/zh_cn/advanced_guides/models.md b/docs/zh_cn/advanced_guides/models.md
index 408a57863c4..6eb22517a46 100644
--- a/docs/zh_cn/advanced_guides/models.md
+++ b/docs/zh_cn/advanced_guides/models.md
@@ -30,17 +30,17 @@
## 基本接口
-MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmenter](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/models/segmentors/base.py#L15) 类,主要提供 `forward`、`train_step`、`val_step` 和 `test_step` 接口。接下来将详细介绍这些接口。
+MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/models/segmentors/base.py#L15) 类,主要提供 `forward`、`train_step`、`val_step` 和 `test_step` 接口。接下来将详细介绍这些接口。
### forward
-  +
+  编码器解码器数据流
-
编码器解码器数据流
-  +
+  级联编码器解码器数据流
级联编码器解码器数据流
@@ -115,7 +115,7 @@ MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmenter](https://github.co
-Dict\[str, `torch.Tensor`\]:用于记录日志的张量的`字典`。
-  +
+  train_step 数据流
train_step 数据流
@@ -132,7 +132,7 @@ MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmenter](https://github.co
- `list` - 给定数据的预测结果。
-  +
+  test_step/val_step 数据流
test_step/val_step 数据流
diff --git a/docs/zh_cn/advanced_guides/training_tricks.md b/docs/zh_cn/advanced_guides/training_tricks.md
index a33c0ea9cfd..e5b8e4dae1e 100644
--- a/docs/zh_cn/advanced_guides/training_tricks.md
+++ b/docs/zh_cn/advanced_guides/training_tricks.md
@@ -1,4 +1,4 @@
-# 训练技巧(待更新)
+# 训练技巧
MMSegmentation 支持如下训练技巧:
@@ -9,17 +9,17 @@ MMSegmentation 支持如下训练技巧:
在 MMSegmentation 里面,您也可以在配置文件里添加如下行来让解码头组件的学习率是主干组件的10倍。
```python
-optimizer=dict(
+optim_wrapper=dict(
paramwise_cfg = dict(
custom_keys={
'head': dict(lr_mult=10.)}))
```
-通过这种修改,任何被分组到 `'head'` 的参数的学习率都将乘以10。您也可以参照 [MMCV 文档](https://mmcv.readthedocs.io/en/latest/api.html#mmcv.runner.DefaultOptimizerConstructor) 获取更详细的信息。
+通过这种修改,任何被分组到 `'head'` 的参数的学习率都将乘以10。您也可以参照 [MMEngine 文档](https://mmengine.readthedocs.io/zh_CN/latest/tutorials/optim_wrapper.html#id6) 获取更详细的信息。
## 在线难样本挖掘 (Online Hard Example Mining, OHEM)
-对于训练时采样,我们在 [这里](https://github.com/open-mmlab/mmsegmentation/tree/master/mmseg/core/seg/sampler) 做了像素采样器。
+MMSegmentation 中实现了像素采样器,训练时可以对特定像素进行采样,例如 OHEM(Online Hard Example Mining),可以解决样本不平衡问题,
如下例子是使用 PSPNet 训练并采用 OHEM 策略的配置:
```python
@@ -58,38 +58,17 @@ model=dict(
```python
_base_ = './fcn_unet_s5-d16_64x64_40k_drive.py'
model = dict(
- decode_head=dict(loss_decode=[dict(type='CrossEntropyLoss', loss_name='loss_ce', loss_weight=1.0),
- dict(type='DiceLoss', loss_name='loss_dice', loss_weight=3.0)]),
- auxiliary_head=dict(loss_decode=[dict(type='CrossEntropyLoss', loss_name='loss_ce',loss_weight=1.0),
- dict(type='DiceLoss', loss_name='loss_dice', loss_weight=3.0)]),
- )
+ decode_head=dict(loss_decode=[
+ dict(type='CrossEntropyLoss', loss_name='loss_ce', loss_weight=1.0),
+ dict(type='DiceLoss', loss_name='loss_dice', loss_weight=3.0)
+ ]),
+ auxiliary_head=dict(loss_decode=[
+ dict(type='CrossEntropyLoss', loss_name='loss_ce', loss_weight=1.0),
+ dict(type='DiceLoss', loss_name='loss_dice', loss_weight=3.0)
+ ]),
+)
```
通过这种方式,确定训练过程中损失函数的权重 `loss_weight` 和在训练日志里的名字 `loss_name`。
-注意: `loss_name` 的名字必须带有 `loss_` 前缀,这样它才能被包括在反传的图里。
-
-## 在损失函数中忽略特定的 label 类别
-
-默认设置 `avg_non_ignore=False`, 即每个像素都用来计算损失函数。尽管其中的一些像素属于需要被忽略的类别。
-
-对于训练时损失函数的计算,我们目前支持使用 `avg_non_ignore` 和 `ignore_index` 来忽略 label 特定的类别。 这样损失函数将只在非忽略类别像素中求平均值,会获得更好的表现。这里是[相关 PR](https://github.com/open-mmlab/mmsegmentation/pull/1409)。以 `unet` 使用 `Cityscapes` 数据集训练为例,
-在计算损失函数时,忽略 label 为0的背景,并且仅在不被忽略的像素上计算均值。配置文件写为:
-
-```python
-_base_ = './fcn_unet_s5-d16_4x4_512x1024_160k_cityscapes.py'
-model = dict(
- decode_head=dict(
- ignore_index=0,
- loss_decode=dict(
- type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0, avg_non_ignore=True),
- auxiliary_head=dict(
- ignore_index=0,
- loss_decode=dict(
- type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0, avg_non_ignore=True)),
- ))
-```
-
-通过这种方式,确定训练过程中损失函数的权重 `loss_weight` 和在训练日志里的名字 `loss_name`。
-
-注意: `loss_name` 的名字必须带有 `loss_` 前缀,这样它才能被包括在反传的图里。
+注意: `loss_name` 的名字必须带有 `loss_` 前缀,这样它才能被包括在计算图里。
diff --git a/docs/zh_cn/get_started.md b/docs/zh_cn/get_started.md
index 38e93e9cb4e..ca375f3d9d6 100644
--- a/docs/zh_cn/get_started.md
+++ b/docs/zh_cn/get_started.md
@@ -4,7 +4,7 @@
本教程中,我们将会演示如何使用 PyTorch 准备环境。
-MMSegmentation 可以在 Linux, Windows 和 macOS 系统上运行,并且需要安装 Python 3.6+, CUDA 9.2+ 和 PyTorch 1.5+
+MMSegmentation 可以在 Linux, Windows 和 macOS 系统上运行,并且需要安装 Python 3.7+, CUDA 10.2+ 和 PyTorch 1.8+
**注意:**
如果您已经安装了 PyTorch, 可以跳过该部分,直接到[下一小节](##安装)。否则,您可以按照以下步骤操作。
@@ -43,7 +43,7 @@ conda install pytorch torchvision cpuonly -c pytorch
```shell
pip install -U openmim
mim install mmengine
-mim install "mmcv>=2.0.0rc1"
+mim install "mmcv>=2.0.0"
```
**步骤 1.** 安装 MMSegmentation
@@ -51,7 +51,7 @@ mim install "mmcv>=2.0.0rc1"
情况 a: 如果您想立刻开发和运行 mmsegmentation,您可通过源码安装:
```shell
-git clone -b dev-1.x https://github.com/open-mmlab/mmsegmentation.git
+git clone -b main https://github.com/open-mmlab/mmsegmentation.git
cd mmsegmentation
pip install -v -e .
# '-v' 表示详细模式,更多的输出
@@ -62,7 +62,7 @@ pip install -v -e .
情况 b: 如果您把 mmsegmentation 作为依赖库或者第三方库,可以通过 pip 安装:
```shell
-pip install "mmsegmentation>=1.0.0rc0"
+pip install "mmsegmentation>=1.0.0"
```
### 验证是否安装成功
@@ -87,8 +87,7 @@ python demo/image_demo.py demo/demo.png configs/pspnet/pspnet_r50-d8_4xb2-40k_ci
您将在当前文件夹中看到一个新图像 `result.jpg`,其中所有目标都覆盖了分割 mask
-选项 (b). 如果您通过 pip 安装 mmsegmentation, 打开您的 python
-解释器,复制粘贴以下代码:
+选项 (b). 如果您通过 pip 安装 mmsegmentation, 打开您的 python 解释器,复制粘贴以下代码:
```python
from mmseg.apis import inference_model, init_model, show_result_pyplot
@@ -111,8 +110,8 @@ show_result_pyplot(model, img, result, show=True, out_file='result.jpg', opacity
# 在一段视频上测试并可视化分割结果
video = mmcv.VideoReader('video.mp4')
for frame in video:
- result = inference_segmentor(model, frame)
- show_result_pyplot(model, result, wait_time=1)
+ result = inference_model(model, frame)
+ show_result_pyplot(model, frame, result, wait_time=1)
```
您可以修改上面的代码来测试单个图像或视频,这两个选项都可以验证安装是否成功。
@@ -137,15 +136,15 @@ MMCV 包含 C++ 和 CUDA 扩展,因此与 PyTorch 的依赖方式比较复杂
为了使用 pip 而不是 MIM 安装 MMCV, 请参考 [MMCV 安装指南](https://mmcv.readthedocs.io/en/latest/get_started/installation.html). 这需要手动指定一个基于 PyTorch 版本及其 CUDA 版本的 find-url.
-例如,以下命令可为 PyTorch 1.10.x and CUDA 11.3 安装 mmcv==2.0.0rc1
+例如,以下命令可为 PyTorch 1.10.x and CUDA 11.3 安装 mmcv==2.0.0
```shell
-pip install mmcv==2.0.0rc1 -f https://download.openmmlab.com/mmcv/dist/cu113/torch1.10/index.html
+pip install mmcv==2.0.0 -f https://download.openmmlab.com/mmcv/dist/cu113/torch1.10/index.html
```
#### 在仅有 CPU 的平台安装
-MMSegmentation 可以在仅有 CPU 的版本上运行。在 CPU 模式,您可以训练(需要 MMCV-Lite 版本 >= 2.0.0rc0),测试和推理模型。
+MMSegmentation 可以在仅有 CPU 的版本上运行。在 CPU 模式,您可以训练(需要 MMCV 版本 >= 2.0.0),测试和推理模型。
#### 在 Google Colab 上安装
@@ -156,7 +155,7 @@ MMSegmentation 可以在仅有 CPU 的版本上运行。在 CPU 模式,您可
```shell
!pip3 install openmim
!mim install mmengine
-!mim install "mmcv>=2.0.0rc1"
+!mim install "mmcv>=2.0.0"
```
**Step 2.** 通过源码安装 MMSegmentation
@@ -164,7 +163,7 @@ MMSegmentation 可以在仅有 CPU 的版本上运行。在 CPU 模式,您可
```shell
!git clone https://github.com/open-mmlab/mmsegmentation.git
%cd mmsegmentation
-!git checkout dev-1.x
+!git checkout main
!pip install -e .
```
@@ -173,7 +172,7 @@ MMSegmentation 可以在仅有 CPU 的版本上运行。在 CPU 模式,您可
```python
import mmseg
print(mmseg.__version__)
-# 示例输出: 1.0.0rc0
+# 示例输出: 1.0.0
```
**注意:**
@@ -195,6 +194,16 @@ docker build -t mmsegmentation docker/
docker run --gpus all --shm-size=8g -it -v {DATA_DIR}:/mmsegmentation/data mmsegmentation
```
+### 可选依赖
+
+#### 安装 GDAL
+
+[GDAL](https://gdal.org/) 是一个用于栅格和矢量地理空间数据格式的转换库。安装 GDAL 可以读取复杂格式和极大的遥感图像。
+
+```shell
+conda install GDAL
+```
+
## 问题解答
如果您在安装过程中遇到了其他问题,请第一时间查阅 [FAQ](notes/faq.md) 文件。如果没有找到答案,您也可以在 GitHub 上提出 [issue](https://github.com/open-mmlab/mmsegmentation/issues/new/choose)
diff --git a/docs/zh_cn/imgs/qq_group_qrcode.jpg b/docs/zh_cn/imgs/qq_group_qrcode.jpg
deleted file mode 100644
index 417347449fe..00000000000
Binary files a/docs/zh_cn/imgs/qq_group_qrcode.jpg and /dev/null differ
diff --git a/docs/zh_cn/imgs/seggroup_qrcode.jpg b/docs/zh_cn/imgs/seggroup_qrcode.jpg
deleted file mode 100644
index 9684582ee1c..00000000000
Binary files a/docs/zh_cn/imgs/seggroup_qrcode.jpg and /dev/null differ
diff --git a/docs/zh_cn/index.rst b/docs/zh_cn/index.rst
index e66c178689d..ce5e49977dc 100644
--- a/docs/zh_cn/index.rst
+++ b/docs/zh_cn/index.rst
@@ -23,7 +23,7 @@
:maxdepth: 1
:caption: 迁移指引
- migration.md
+ migration/index.rst
.. toctree::
:caption: 接口文档(英文)
diff --git a/docs/zh_cn/migration/interface.md b/docs/zh_cn/migration/interface.md
index cd16d2cbc68..42f91bf50ac 100644
--- a/docs/zh_cn/migration/interface.md
+++ b/docs/zh_cn/migration/interface.md
@@ -2,7 +2,7 @@
## 引言
-本指南介绍了 MMSegmentation 0.x 和 MMSegmentation1.x 在行为和 API 方面的基本区别,以及这些如何都与您的迁移过程相关。
+本指南介绍了 MMSegmentation 0.x 和 MMSegmentation1.x 在表现和 API 方面的基本区别,以及这些与迁移过程的关系。
## 新的依赖
@@ -12,11 +12,11 @@ MMSegmentation 1.x 依赖于一些新的软件包,您可以准备一个新的
1. [MMEngine](https://github.com/open-mmlab/mmengine):MMEngine 是 OpenMMLab 2.0 架构的核心,我们将许多与计算机视觉无关的内容从 MMCV 拆分到 MMEngine 中。
-2. [MMCV](https://github.com/open-mmlab/mmcv):OpenMMLab 的计算机视觉包。这不是一个新的依赖,但您需要将其升级到 **2.0.0rc1** 以上的版本。
+2. [MMCV](https://github.com/open-mmlab/mmcv):OpenMMLab 的计算机视觉包。这不是一个新的依赖,但您需要将其升级到 **2.0.0** 或以上的版本。
-3. [MMClassification](https://github.com/open-mmlab/mmclassification)(可选):OpenMMLab 的图像分类工具箱和基准。这不是一个新的依赖,但您需要将其升级到 **1.0.0rc0** 以上的版本。
+3. [MMClassification](https://github.com/open-mmlab/mmclassification)(可选):OpenMMLab 的图像分类工具箱和基准。这不是一个新的依赖,但您需要将其升级到 **1.0.0rc6** 版本。
-4. [MMDetection](https://github.com/open-mmlab/mmdetection)(可选): OpenMMLab 的目标检测工具箱和基准。这不是一个新的依赖,但您需要将其升级到 **3.0.0rc0** 以上的版本。
+4. [MMDetection](https://github.com/open-mmlab/mmdetection)(可选): OpenMMLab 的目标检测工具箱和基准。这不是一个新的依赖,但您需要将其升级到 **3.0.0** 或以上的版本。
## 启动训练
@@ -46,7 +46,7 @@ OpenMMLab 2.0 的主要改进是发布了 MMEngine,它为启动训练任务的
--resume='auto' |
-| 培训练期间是否不评估检查点 |
+训练期间是否不评估检查点 |
--no-validate |
--cfg-options val_cfg=None val_dataloader=None val_evaluator=None |
@@ -102,11 +102,11 @@ OpenMMLab 2.0 的主要改进是发布了 MMEngine,它为启动训练任务的
- `mean`(Sequence,可选):R、G、B 通道的像素平均值。默认为 None。
-- `std`(Sequence,可选):R、G、B通道的像素标准差。默认为 None。
+- `std`(Sequence,可选):R、G、B 通道的像素标准差。默认为 None。
- `size`(Sequence,可选):固定的填充大小。
-- `size_divisor`(int,可选):填充大小的除法因子。
+- `size_divisor`(int,可选):填充图像可以被当前值整除。
- `seg_pad_val`(float,可选):分割图的填充值。默认值:255。
@@ -154,14 +154,14 @@ train_dataloader = dict(
batch_size=4,
num_workers=4,
dataset=dict(...),
- sampler=dict(type='DefaultSampler', shuffle=True) # necessary
+ sampler=dict(type='DefaultSampler', shuffle=True) # 必须
)
val_dataloader = dict(
batch_size=4,
num_workers=4,
dataset=dict(...),
- sampler=dict(type='DefaultSampler', shuffle=False) # necessary
+ sampler=dict(type='DefaultSampler', shuffle=False) # 必须
)
test_dataloader = val_dataloader
@@ -417,10 +417,10 @@ runner = dict(type='IterBasedRunner', max_iters=20000)
```python
-# The `val_interval` is the original `evaluation.interval`.
+# `val_interval` 是旧版本的 `evaluation.interval`。
train_cfg = dict(type='IterBasedTrainLoop', max_iters=20000, val_interval=2000)
-val_cfg = dict(type='ValLoop') # Use the default validation loop.
-test_cfg = dict(type='TestLoop') # Use the default test loop.
+val_cfg = dict(type='ValLoop') # 使用默认的验证循环。
+test_cfg = dict(type='TestLoop') # 使用默认的测试循环。
```
|
@@ -438,22 +438,22 @@ test_cfg = dict(type='TestLoop') # Use the default test loop.
```python
default_hooks = dict(
- # record the time of every iterations.
+ # 记录每次迭代的时间。
timer=dict(type='IterTimerHook'),
- # print log every 50 iterations.
+ # 每50次迭代打印一次日志。
logger=dict(type='LoggerHook', interval=50, log_metric_by_epoch=False),
- # enable the parameter scheduler.
+ # 启用参数调度程序。
param_scheduler=dict(type='ParamSchedulerHook'),
- # save checkpoint every 2000 iterations.
+ # 每2000次迭代保存一次检查点。
checkpoint=dict(type='CheckpointHook', by_epoch=False, interval=2000),
- # set sampler seed in distributed environment.
+ # 在分布式环境中设置采样器种子。
sampler_seed=dict(type='DistSamplerSeedHook'),
- # validation results visualization.
+ # 验证结果可视化。
visualization=dict(type='SegVisualizationHook'))
```
@@ -505,13 +505,13 @@ visualizer = dict(
```python
env_cfg = dict(
- # whether to enable cudnn benchmark
+ # 是否启用 cudnn_benchmark
cudnn_benchmark=False,
- # set multi process parameters
+ # 设置多进程参数
mp_cfg=dict(mp_start_method='fork', opencv_num_threads=0),
- # set distributed parameters
+ # 设置分布式参数
dist_cfg=dict(backend='nccl'),
)
```
diff --git a/docs/zh_cn/migration/package.md b/docs/zh_cn/migration/package.md
index d8d2245bed0..19e5f18c9c4 100644
--- a/docs/zh_cn/migration/package.md
+++ b/docs/zh_cn/migration/package.md
@@ -1,6 +1,6 @@
-#包结构更改
+# 包结构更改
-本节包含您对 MMSeg 0.x 和 1.x 之间的变化感到好奇的内容。
+本节包含您对 MMSeg 0.x 和 1.x 之间的变化可能感到好奇的内容。
@@ -49,7 +49,7 @@
## `mmseg.ops`
-`ops` 包包含 `encoding` 和 `wrappers`,它们被移到了 `mmseg.models.utils` 中。
+`ops` 包含 `encoding` 和 `wrappers`,它们被移到了 `mmseg.models.utils` 中。
## 增加的包
@@ -110,4 +110,4 @@ OpenMMLab 2.0 将 `BaseDataset` 定义为数据集的函数和接口,MMSegment
### `mmseg.models`
-`models` 没有太大变化,只是从以前的 `mmseg.ops` 中添加了 `encoding` 和 `wrappers`
+`models` 没有太大变化,只是从以前的 `mmseg.ops` 添加了 `encoding` 和 `wrappers`
diff --git a/docs/zh_cn/notes/faq.md b/docs/zh_cn/notes/faq.md
index 09fde025fde..bf3b3417802 100644
--- a/docs/zh_cn/notes/faq.md
+++ b/docs/zh_cn/notes/faq.md
@@ -1,8 +1,122 @@
-# 常见问题解答(FAQ)(待更新)
+# 常见问题解答(FAQ)
-我们在这里列出了使用时的一些常见问题及其相应的解决方案。 如果您发现有一些问题被遗漏,请随时提 PR 丰富这个列表。 如果您无法在此获得帮助,请使用 [issue模板](https://github.com/open-mmlab/mmsegmentation/blob/master/.github/ISSUE_TEMPLATE/error-report.md/)创建问题,但是请在模板中填写所有必填信息,这有助于我们更快定位问题。
+我们在这里列出了使用时的一些常见问题及其相应的解决方案。 如果您发现有一些问题被遗漏,请随时提 PR 丰富这个列表。 如果您无法在此获得帮助,请使用 [issue 模板](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/.github/ISSUE_TEMPLATE/error-report.md/)创建问题,但是请在模板中填写所有必填信息,这有助于我们更快定位问题。
+
+## 安装
+
+兼容的 MMSegmentation 和 MMCV 版本如下。请安装正确版本的 MMCV 以避免安装问题。
+
+| MMSegmentation version | MMCV version | MMEngine version | MMClassification (optional) version | MMDetection (optional) version |
+| :--------------------: | :----------------------------: | :---------------: | :---------------------------------: | :----------------------------: |
+| dev-1.x branch | mmcv >= 2.0.0 | MMEngine >= 0.7.4 | mmpretrain>=1.0.0rc7 | mmdet >= 3.0.0 |
+| main branch | mmcv >= 2.0.0 | MMEngine >= 0.7.4 | mmpretrain>=1.0.0rc7 | mmdet >= 3.0.0 |
+| 1.1.2 | mmcv >= 2.0.0 | MMEngine >= 0.7.4 | mmpretrain>=1.0.0rc7 | mmdet >= 3.0.0 |
+| 1.1.1 | mmcv >= 2.0.0 | MMEngine >= 0.7.4 | mmpretrain>=1.0.0rc7 | mmdet >= 3.0.0 |
+| 1.1.0 | mmcv >= 2.0.0 | MMEngine >= 0.7.4 | mmpretrain>=1.0.0rc7 | mmdet >= 3.0.0 |
+| 1.0.0 | mmcv >= 2.0.0rc4 | MMEngine >= 0.7.1 | mmcls==1.0.0rc6 | mmdet >= 3.0.0 |
+| 1.0.0rc6 | mmcv >= 2.0.0rc4 | MMEngine >= 0.5.0 | mmcls>=1.0.0rc0 | mmdet >= 3.0.0rc6 |
+| 1.0.0rc5 | mmcv >= 2.0.0rc4 | MMEngine >= 0.2.0 | mmcls>=1.0.0rc0 | mmdet>=3.0.0rc6 |
+| 1.0.0rc4 | mmcv == 2.0.0rc3 | MMEngine >= 0.1.0 | mmcls>=1.0.0rc0 | mmdet>=3.0.0rc4, \<=3.0.0rc5 |
+| 1.0.0rc3 | mmcv == 2.0.0rc3 | MMEngine >= 0.1.0 | mmcls>=1.0.0rc0 | mmdet>=3.0.0rc4, \<=3.0.0rc5 |
+| 1.0.0rc2 | mmcv == 2.0.0rc3 | MMEngine >= 0.1.0 | mmcls>=1.0.0rc0 | mmdet>=3.0.0rc4, \<=3.0.0rc5 |
+| 1.0.0rc1 | mmcv >= 2.0.0rc1, \<=2.0.0rc3> | MMEngine >= 0.1.0 | mmcls>=1.0.0rc0 | Not required |
+| 1.0.0rc0 | mmcv >= 2.0.0rc1, \<=2.0.0rc3> | MMEngine >= 0.1.0 | mmcls>=1.0.0rc0 | Not required |
+
+如果您已经安装了版本不合适的 mmcv,请先运行`pip uninstall mmcv`卸载已安装的 mmcv,如您先前安装的为 mmcv-full(存在于 OpenMMLab 1.x),请运行`pip uninstall mmcv-full`进行卸载。
+
+- 如出现 "No module named 'mmcv'"
+ 1. 使用`pip uninstall mmcv`卸载环境中现有的 mmcv
+ 2. 按照[安装说明](../get_started.md)安装对应的 mmcv
## 如何获知模型训练时需要的显卡数量
-- 看模型的config文件的命名。可以参考[学习配置文件](https://github.com/open-mmlab/mmsegmentation/blob/master/docs/zh_cn/tutorials/config.md)中的`配置文件命名风格`部分。比如,对于名字为`segformer_mit-b0_8x1_1024x1024_160k_cityscapes.py`的config文件,`8x1`代表训练其对应的模型需要的卡数为8,每张卡中的batch size为1。
-- 看模型的log文件。点开该模型的log文件,并在其中搜索`nGPU`,在`nGPU`后的数字个数即训练时所需的卡数。比如,在log文件中搜索`nGPU`得到`nGPU 0,1,2,3,4,5,6,7`的记录,则说明训练该模型需要使用八张卡。
+- 看模型的 config 文件命名。可以参考[了解配置文件](../user_guides/1_config.md)中的`配置文件命名风格`部分。比如,对于名字为`segformer_mit-b0_8xb1-160k_cityscapes-1024x1024.py`的 config 文件,`8xb1`代表训练其对应的模型需要的卡数为 8,每张卡中的 batch size 为 1。
+- 看模型的 log 文件。点开该模型的 log 文件,并在其中搜索`nGPU`,在`nGPU`后的数字个数即训练时所需的卡数。比如,在 log 文件中搜索`nGPU`得到`nGPU 0,1,2,3,4,5,6,7`的记录,则说明训练该模型需要使用八张卡。
+
+## auxiliary head 是什么
+
+简单来说,这是一个提高准确率的深度监督技术。在训练阶段,`decode_head`用于输出语义分割的结果,`auxiliary_head` 只是增加了一个辅助损失,其产生的分割结果对你的模型结果没有影响,仅在在训练中起作用。您可以阅读这篇[论文](https://arxiv.org/pdf/1612.01105.pdf)了解更多信息。
+
+## 运行测试脚本时如何输出绘制分割掩膜的图像
+
+在测试脚本中,我们提供了`--out`参数来控制是否输出保存预测的分割掩膜图像。您可以运行以下命令输出测试结果:
+
+```shell
+python tools/test.py ${CONFIG_FILE} ${CHECKPOINT_FILE} --out ${OUTPUT_DIR}
+```
+
+更多用例细节可查阅[文档](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/user_guides/4_train_test.md#%E6%B5%8B%E8%AF%95%E5%B9%B6%E4%BF%9D%E5%AD%98%E5%88%86%E5%89%B2%E7%BB%93%E6%9E%9C),[PR #2712](https://github.com/open-mmlab/mmsegmentation/pull/2712) 以及[迁移文档](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/migration/interface.md#%E6%B5%8B%E8%AF%95%E5%90%AF%E5%8A%A8)了解相关说明。
+
+## 如何处理二值分割任务?
+
+MMSegmentation 使用 `num_classes` 和 `out_channels` 来控制模型最后一层 `self.conv_seg` 的输出。更多细节可以参考 [这里](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/decode_heads/decode_head.py)。
+
+`num_classes` 应该和数据集本身类别个数一致,当是二值分割时,数据集只有前景和背景两类,所以 `num_classes` 为 2. `out_channels` 控制模型最后一层的输出的通道数,通常和 `num_classes` 相等,但当二值分割时候,可以有两种处理方法, 分别是:
+
+- 设置 `out_channels=2`,在训练时以 Cross Entropy Loss 作为损失函数,在推理时使用 `F.softmax()` 归一化 logits 值,然后通过 `argmax()` 得到每个像素的预测结果。
+
+- 设置 `out_channels=1`,在训练时以 Binary Cross Entropy Loss 作为损失函数,在推理时使用 `F.sigmoid()` 和 `threshold` 得到预测结果,`threshold` 默认为 0.3。
+
+对于实现上述两种计算二值分割的方法,需要在 `decode_head` 和 `auxiliary_head` 的配置里修改。下面是对样例 [pspnet_unet_s5-d16.py](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/models/pspnet_unet_s5-d16.py) 做出的对应修改。
+
+- (1) `num_classes=2`, `out_channels=2` 并在 `CrossEntropyLoss` 里面设置 `use_sigmoid=False`。
+
+```python
+decode_head=dict(
+ type='PSPHead',
+ in_channels=64,
+ in_index=4,
+ num_classes=2,
+ out_channels=2,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0)),
+auxiliary_head=dict(
+ type='FCNHead',
+ in_channels=128,
+ in_index=3,
+ num_classes=2,
+ out_channels=2,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=0.4)),
+```
+
+- (2) `num_classes=2`, `out_channels=1` 并在 `CrossEntropyLoss` 里面设置 `use_sigmoid=True`.
+
+```python
+decode_head=dict(
+ type='PSPHead',
+ in_channels=64,
+ in_index=4,
+ num_classes=2,
+ out_channels=1,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=True, loss_weight=1.0)),
+auxiliary_head=dict(
+ type='FCNHead',
+ in_channels=128,
+ in_index=3,
+ num_classes=2,
+ out_channels=1,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=True, loss_weight=0.4)),
+```
+
+## `reduce_zero_label` 的作用
+
+数据集中 `reduce_zero_label` 参数类型为布尔类型,默认为 False,它的功能是为了忽略数据集 label 0。具体做法是将 label 0 改为 255,其余 label 相应编号减 1,同时 decode head 里将 255 设为 ignore index,即不参与 loss 计算。
+以下是 `reduce_zero_label` 具体实现逻辑:
+
+```python
+if self.reduce_zero_label:
+ # avoid using underflow conversion
+ gt_semantic_seg[gt_semantic_seg == 0] = 255
+ gt_semantic_seg = gt_semantic_seg - 1
+ gt_semantic_seg[gt_semantic_seg == 254] = 255
+```
+
+关于您的数据集是否需要使用 reduce_zero_label,有以下两类情况:
+
+- 例如在 [Potsdam](https://github.com/open-mmlab/mmsegmentation/blob/1.x/docs/en/user_guides/2_dataset_prepare.md#isprs-potsdam) 数据集上,有 0-不透水面、1-建筑、2-低矮植被、3-树、4-汽车、5-杂乱,六类。但该数据集提供了两种 RGB 标签,一种为图像边缘处有黑色像素的标签,另一种是没有黑色边缘的标签。对于有黑色边缘的标签,在 [dataset_converters.py](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/tools/dataset_converters/potsdam.py)中,其将黑色边缘转换为 label 0,其余标签分别为 1-不透水面、2-建筑、3-低矮植被、4-树、5-汽车、6-杂乱,那么此时,就应该在数据集 [potsdam.py](https://github.com/open-mmlab/mmsegmentation/blob/ff95416c3b5ce8d62b9289f743531398efce534f/mmseg/datasets/potsdam.py#L23) 中将`reduce_zero_label=True`。如果使用的是没有黑色边缘的标签,那么 mask label 中只有 0-5,此时就应该使`reduce_zero_label=False`。需要结合您的实际情况来使用。
+- 例如在第 0 类为 background 类别的数据集上,如果您最终是需要将背景和您的其余类别分开时,是不需要使用`reduce_zero_label`的,此时在数据集中应该将其设置为`reduce_zero_label=False`
+
+**注意:** 使用 `reduce_zero_label` 请确认数据集原始类别个数,如果只有两类,需要关闭 `reduce_zero_label` 即设置 `reduce_zero_label=False`。
diff --git a/docs/zh_cn/overview.md b/docs/zh_cn/overview.md
index 7dce105a813..ed147956d08 100644
--- a/docs/zh_cn/overview.md
+++ b/docs/zh_cn/overview.md
@@ -42,7 +42,7 @@ MMSeg 主要包含了 apis, structures, datasets, models, engine, evaluation 和
以下是详细步骤,将带您一步步学习如何使用 MMSegmentation :
-1. 有关安装说明,请参阅 [开始你的第一步](getting_started.md)。
+1. 有关安装说明,请参阅 [开始你的第一步](get_started.md)。
2. 对于初学者来说,MMSegmentation 是开始语义分割之旅的最好选择,因为这里实现了许多 SOTA 模型以及经典的模型 [model](model_zoo.md) 。另外,将各类组件和高级 API 結合使用,可以更便捷的执行分割任务。关于 MMSegmentation 的基本用法,请参考下面的教程:
@@ -62,8 +62,8 @@ MMSeg 主要包含了 apis, structures, datasets, models, engine, evaluation 和
4. MMSegmentation 也为用户自定义和一些前沿的研究提供了教程,请参考下面的教程来建立你自己的分割项目:
- [添加新的模型](advanced_guides/add_models.md)
- - [添加新的数据集](advanced_guides/add_dataset.md)
- - [添加新的 transform](advanced_guides/add_transform.md)
+ - [添加新的数据集](advanced_guides/add_datasets.md)
+ - [添加新的 transform](advanced_guides/add_transforms.md)
- [自定义 runtime](advanced_guides/customize_runtime.md)
5. 如果您更熟悉 MMSegmentation v0.x , 以下是 MMSegmentation v0.x 迁移到 v1.x 的文档
diff --git a/docs/zh_cn/user_guides/2_dataset_prepare.md b/docs/zh_cn/user_guides/2_dataset_prepare.md
index c9c3606977d..5532624bef4 100644
--- a/docs/zh_cn/user_guides/2_dataset_prepare.md
+++ b/docs/zh_cn/user_guides/2_dataset_prepare.md
@@ -1,3 +1,750 @@
-## 准备数据集(待更新)
+# 教程2:准备数据集
-中文版文档支持中,请先阅读[英文版本](../../en/user_guides/2_dataset_prepare.md)
+我们建议将数据集根目录符号链接到 `$MMSEGMENTATION/data`。
+如果您的目录结构不同,您可能需要更改配置文件中相应的路径。
+对于中国境内的用户,我们也推荐通过开源数据平台 [OpenDataLab](https://opendatalab.com/) 来下载dsdl标准数据,以获得更好的下载和使用体验,这里有一个下载dsdl数据集并进行训练的案例[DSDLReadme](../../../configs/dsdl/README.md),欢迎尝试。
+
+```none
+mmsegmentation
+├── mmseg
+├── tools
+├── configs
+├── data
+│ ├── cityscapes
+│ │ ├── leftImg8bit
+│ │ │ ├── train
+│ │ │ ├── val
+│ │ ├── gtFine
+│ │ │ ├── train
+│ │ │ ├── val
+│ ├── VOCdevkit
+│ │ ├── VOC2012
+│ │ │ ├── JPEGImages
+│ │ │ ├── SegmentationClass
+│ │ │ ├── ImageSets
+│ │ │ │ ├── Segmentation
+│ │ ├── VOC2010
+│ │ │ ├── JPEGImages
+│ │ │ ├── SegmentationClassContext
+│ │ │ ├── ImageSets
+│ │ │ │ ├── SegmentationContext
+│ │ │ │ │ ├── train.txt
+│ │ │ │ │ ├── val.txt
+│ │ │ ├── trainval_merged.json
+│ │ ├── VOCaug
+│ │ │ ├── dataset
+│ │ │ │ ├── cls
+│ ├── ade
+│ │ ├── ADEChallengeData2016
+│ │ │ ├── annotations
+│ │ │ │ ├── training
+│ │ │ │ ├── validation
+│ │ │ ├── images
+│ │ │ │ ├── training
+│ │ │ │ ├── validation
+│ ├── coco_stuff10k
+│ │ ├── images
+│ │ │ ├── train2014
+│ │ │ ├── test2014
+│ │ ├── annotations
+│ │ │ ├── train2014
+│ │ │ ├── test2014
+│ │ ├── imagesLists
+│ │ │ ├── train.txt
+│ │ │ ├── test.txt
+│ │ │ ├── all.txt
+│ ├── coco_stuff164k
+│ │ ├── images
+│ │ │ ├── train2017
+│ │ │ ├── val2017
+│ │ ├── annotations
+│ │ │ ├── train2017
+│ │ │ ├── val2017
+│ ├── CHASE_DB1
+│ │ ├── images
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ ├── annotations
+│ │ │ ├── training
+│ │ │ ├── validation
+│ ├── DRIVE
+│ │ ├── images
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ ├── annotations
+│ │ │ ├── training
+│ │ │ ├── validation
+│ ├── HRF
+│ │ ├── images
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ ├── annotations
+│ │ │ ├── training
+│ │ │ ├── validation
+│ ├── STARE
+│ │ ├── images
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ ├── annotations
+│ │ │ ├── training
+│ │ │ ├── validation
+| ├── dark_zurich
+| │ ├── gps
+| │ │ ├── val
+| │ │ └── val_ref
+| │ ├── gt
+| │ │ └── val
+| │ ├── LICENSE.txt
+| │ ├── lists_file_names
+| │ │ ├── val_filenames.txt
+| │ │ └── val_ref_filenames.txt
+| │ ├── README.md
+| │ └── rgb_anon
+| │ | ├── val
+| │ | └── val_ref
+| ├── NighttimeDrivingTest
+| | ├── gtCoarse_daytime_trainvaltest
+| | │ └── test
+| | │ └── night
+| | └── leftImg8bit
+| | | └── test
+| | | └── night
+│ ├── loveDA
+│ │ ├── img_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ │ │ ├── test
+│ │ ├── ann_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ ├── potsdam
+│ │ ├── img_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ │ ├── ann_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ ├── vaihingen
+│ │ ├── img_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ │ ├── ann_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ ├── iSAID
+│ │ ├── img_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ │ │ ├── test
+│ │ ├── ann_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ ├── synapse
+│ │ ├── img_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ │ ├── ann_dir
+│ │ │ ├── train
+│ │ │ ├── val
+│ ├── REFUGE
+│ │ ├── images
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ │ ├── test
+│ │ ├── annotations
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ │ ├── test
+│ ├── mapillary
+│ │ ├── training
+│ │ │ ├── images
+│ │ │ ├── v1.2
+| │ │ │ ├── instances
+| │ │ │ ├── labels
+| │ │ │ └── panoptic
+│ │ │ ├── v2.0
+| │ │ │ ├── instances
+| │ │ │ ├── labels
+| │ │ │ ├── panoptic
+| │ │ │ └── polygons
+│ │ ├── validation
+│ │ │ ├── images
+| │ │ ├── v1.2
+| │ │ │ ├── instances
+| │ │ │ ├── labels
+| │ │ │ └── panoptic
+│ │ │ ├── v2.0
+| │ │ │ ├── instances
+| │ │ │ ├── labels
+| │ │ │ ├── panoptic
+| │ │ │ └── polygons
+│ ├── bdd100k
+│ │ ├── images
+│ │ │ └── 10k
+| │ │ │ ├── test
+| │ │ │ ├── train
+| │ │ │ └── val
+│ │ └── labels
+│ │ │ └── sem_seg
+| │ │ │ ├── colormaps
+| │ │ │ │ ├──train
+| │ │ │ │ └──val
+| │ │ │ ├── masks
+| │ │ │ │ ├──train
+| │ │ │ │ └──val
+| │ │ │ ├── polygons
+| │ │ │ │ ├──sem_seg_train.json
+| │ │ │ │ └──sem_seg_val.json
+| │ │ │ └── rles
+| │ │ │ │ ├──sem_seg_train.json
+| │ │ │ │ └──sem_seg_val.json
+│ ├── nyu
+│ │ ├── images
+│ │ │ ├── train
+│ │ │ ├── test
+│ │ ├── annotations
+│ │ │ ├── train
+│ │ │ ├── test
+```
+
+## 用 MIM 下载数据集
+
+通过使用 [OpenXLab](https://openxlab.org.cn/datasets),您可以直接下载开源数据集。通过平台的搜索功能,您可以快速轻松地找到他们正在寻找的数据集。使用平台上的格式化数据集,您可以高效地跨数据集执行任务。
+
+如果您使用 MIM 下载,请确保版本大于 v0.3.8。您可以使用以下命令进行更新、安装、登录和数据集下载:
+
+```shell
+# upgrade your MIM
+pip install -U openmim
+
+# install OpenXLab CLI tools
+pip install -U openxlab
+# log in OpenXLab
+openxlab login
+
+# download ADE20K by MIM
+mim download mmsegmentation --dataset ade20k
+```
+
+## Cityscapes
+
+Cityscapes [官方网站](https://www.cityscapes-dataset.com/)可以下载 Cityscapes 数据集,按照官网要求注册并登陆后,数据可以在[这里](https://www.cityscapes-dataset.com/downloads/)找到。
+
+按照惯例,`**labelTrainIds.png` 用于 cityscapes 训练。
+我们提供了一个基于 [cityscapesscripts](https://github.com/mcordts/cityscapesScripts) 的[脚本](https://github.com/open-mmlab/mmsegmentation/blob/1.x/tools/dataset_converters/cityscapes.py)用于生成 `**labelTrainIds.png`。
+
+```shell
+# --nproc 表示 8 个转换进程,也可以省略。
+python tools/dataset_converters/cityscapes.py data/cityscapes --nproc 8
+```
+
+## Pascal VOC
+
+Pascal VOC 2012 可从[此处](http://host.robots.ox.ac.uk/pascal/VOC/voc2012/VOCtrainval_11-May-2012.tar)下载。
+此外,Pascal VOC 数据集的最新工作通常利用额外的增强数据,可以在[这里](http://www.eecs.berkeley.edu/Research/Projects/CS/vision/grouping/semantic_contours/benchmark.tgz)找到。
+
+如果您想使用增强的 VOC 数据集,请运行以下命令将增强数据的标注转换为正确的格式。
+
+```shell
+# --nproc 表示 8 个转换进程,也可以省略。
+python tools/dataset_converters/voc_aug.py data/VOCdevkit data/VOCdevkit/VOCaug --nproc 8
+```
+
+请参考[拼接数据集文档](../advanced_guides/add_datasets.md#拼接数据集)及 [voc_aug 配置示例](../../../configs/_base_/datasets/pascal_voc12_aug.py)以详细了解如何将它们拼接并合并训练。
+
+## ADE20K
+
+ADE20K 的训练和验证集可以从这个[链接](http://data.csail.mit.edu/places/ADEchallenge/ADEChallengeData2016.zip)下载。
+如果需要下载测试数据集,可以在[官网](http://host.robots.ox.ac.uk/)注册后,下载[测试集](http://host.robots.ox.ac.uk:8080/eval/downloads/VOC2010test.tar)。
+
+## Pascal Context
+
+Pascal Context 的训练和验证集可以从[此处](http://host.robots.ox.ac.uk/pascal/VOC/voc2010/VOCtrainval_03-May-2010.tar)下载。注册后,您也可以从[此处](http://host.robots.ox.ac.uk:8080/eval/downloads/VOC2010test.tar)下载测试集。
+
+从原始数据集中抽出部分数据作为验证集,您可以从[此处](https://codalabuser.blob.core.windows.net/public/trainval_merged.json)下载 trainval_merged.json 文件。
+
+请先安装 [Detail](https://github.com/zhanghang1989/detail-api) 工具然后运行以下命令将标注转换为正确的格式。
+
+```shell
+python tools/dataset_converters/pascal_context.py data/VOCdevkit data/VOCdevkit/VOC2010/trainval_merged.json
+```
+
+## COCO Stuff 10k
+
+数据可以通过 wget 在[这里](http://calvin.inf.ed.ac.uk/wp-content/uploads/data/cocostuffdataset/cocostuff-10k-v1.1.zip)下载。
+
+对于 COCO Stuff 10k 数据集,请运行以下命令下载并转换数据集。
+
+```shell
+# 下载
+mkdir coco_stuff10k && cd coco_stuff10k
+wget http://calvin.inf.ed.ac.uk/wp-content/uploads/data/cocostuffdataset/cocostuff-10k-v1.1.zip
+
+# 解压
+unzip cocostuff-10k-v1.1.zip
+
+# --nproc 表示 8 个转换进程,也可以省略。
+python tools/dataset_converters/coco_stuff10k.py /path/to/coco_stuff10k --nproc 8
+```
+
+按照惯例,`/path/to/coco_stuff164k/annotations/*2014/*_labelTrainIds.png` 中的 mask 标注用于 COCO Stuff 10k 的训练和测试。
+
+## COCO Stuff 164k
+
+对于 COCO Stuff 164k 数据集,请运行以下命令下载并转换增强的数据集。
+
+```shell
+# 下载
+mkdir coco_stuff164k && cd coco_stuff164k
+wget http://images.cocodataset.org/zips/train2017.zip
+wget http://images.cocodataset.org/zips/val2017.zip
+wget http://calvin.inf.ed.ac.uk/wp-content/uploads/data/cocostuffdataset/stuffthingmaps_trainval2017.zip
+
+# 解压
+unzip train2017.zip -d images/
+unzip val2017.zip -d images/
+unzip stuffthingmaps_trainval2017.zip -d annotations/
+
+# --nproc 表示 8 个转换进程,也可以省略。
+python tools/dataset_converters/coco_stuff164k.py /path/to/coco_stuff164k --nproc 8
+```
+
+按照惯例,`/path/to/coco_stuff164k/annotations/*2017/*_labelTrainIds.png` 中的 mask 标注用于 COCO Stuff 164k 的训练和测试。
+
+此数据集的详细信息可在[此处](https://github.com/nightrome/cocostuff#downloads)找到。
+
+## CHASE DB1
+
+CHASE DB1 的训练和验证集可以从[此处](https://staffnet.kingston.ac.uk/~ku15565/CHASE_DB1/assets/CHASEDB1.zip)下载。
+
+请运行以下命令,准备 CHASE DB1 数据集:
+
+```shell
+python tools/dataset_converters/chase_db1.py /path/to/CHASEDB1.zip
+```
+
+该脚本将自动调整数据集目录结构,使其满足 MMSegmentation 数据集加载要求。
+
+## DRIVE
+
+按照[官网](https://drive.grand-challenge.org/)要求,注册并登陆后,便可以下载 DRIVE 的训练和验证数据集。
+
+要将 DRIVE 数据集转换为 MMSegmentation 的格式,请运行以下命令:
+
+```shell
+python tools/dataset_converters/drive.py /path/to/training.zip /path/to/test.zip
+```
+
+该脚本将自动调整数据集目录结构,使其满足 MMSegmentation 数据集加载要求。
+
+## HRF
+
+请下载 [health.zip](https://www5.cs.fau.de/fileadmin/research/datasets/fundus-images/healthy.zip)、[glaucoma.zip](https://www5.cs.fau.de/fileadmin/research/datasets/fundus-images/glaucoma.zip)、[diabetic_retinopathy.zip](https://www5.cs.fau.de/fileadmin/research/datasets/fundus-images/diabetic_retinopathy.zip)、[healthy_manualsegm.zip](https://www5.cs.fau.de/fileadmin/research/datasets/fundus-images/healthy_manualsegm.zip)、[glaucoma_manualsegm.zip](https://www5.cs.fau.de/fileadmin/research/datasets/fundus-images/glaucoma_manualsegm.zip) 和 [diabetic_retinopathy_manualsegm.zip](https://www5.cs.fau.de/fileadmin/research/datasets/fundus-images/diabetic_retinopathy_manualsegm.zip),无需解压,可以直接运行以下命令,准备 HRF 数据集:
+
+```shell
+python tools/dataset_converters/hrf.py /path/to/healthy.zip /path/to/healthy_manualsegm.zip /path/to/glaucoma.zip /path/to/glaucoma_manualsegm.zip /path/to/diabetic_retinopathy.zip /path/to/diabetic_retinopathy_manualsegm.zip
+```
+
+该脚本将自动调整数据集目录结构,使其满足 MMSegmentation 数据集加载要求。
+
+## STARE
+
+请下载 [stare images.tar](http://cecas.clemson.edu/~ahoover/stare/probing/stare-images.tar)、[labels-ah.tar](http://cecas.clemson.edu/~ahoover/stare/probing/labels-ah.tar) 和 [labels-vk.tar](http://cecas.clemson.edu/~ahoover/stare/probing/labels-vk.tar),无需解压,可以直接运行以下命令,准备 STARE 数据集:
+
+```shell
+python tools/dataset_converters/stare.py /path/to/stare-images.tar /path/to/labels-ah.tar /path/to/labels-vk.tar
+```
+
+该脚本将自动调整数据集目录结构,使其满足 MMSegmentation 数据集加载要求。
+
+## Dark Zurich
+
+由于我们只支持在此数据集上的模型测试,因此您只需要下载并解压[验证数据集](https://data.vision.ee.ethz.ch/csakarid/shared/GCMA_UIoU/Dark_Zurich_val_anon.zip)。
+
+## Nighttime Driving
+
+由于我们只支持在此数据集上的模型测试,因此您只需要下载并解压[验证数据集](http://data.vision.ee.ethz.ch/daid/NighttimeDriving/NighttimeDrivingTest.zip)。
+
+## LoveDA
+
+数据可以从[此处](https://drive.google.com/drive/folders/1ibYV0qwn4yuuh068Rnc-w4tPi0U0c-ti?usp=sharing)下载 LaveDA 数据集。
+
+或者可以从 [zenodo](https://zenodo.org/record/5706578#.YZvN7SYRXdF) 下载。下载后,无需解压,直接运行以下命令:
+
+```shell
+# 下载 Train.zip
+wget https://zenodo.org/record/5706578/files/Train.zip
+# 下载 Val.zip
+wget https://zenodo.org/record/5706578/files/Val.zip
+# 下载 Test.zip
+wget https://zenodo.org/record/5706578/files/Test.zip
+```
+
+请对于 LoveDA 数据集,请运行以下命令调整数据集目录。
+
+```shell
+python tools/dataset_converters/loveda.py /path/to/loveDA
+```
+
+可将模型对 LoveDA 的测试集的预测结果上传至到数据集[测试服务器](https://codalab.lisn.upsaclay.fr/competitions/421),查看评测结果。
+
+有关 LoveDA 的更多详细信息,可查看[此处](https://github.com/Junjue-Wang/LoveDA).
+
+## ISPRS Potsdam
+
+[Potsdam](https://www.isprs.org/education/benchmarks/UrbanSemLab/2d-sem-label-potsdam.aspx) 城市语义分割数据集用于 2D 语义分割竞赛 —— Potsdam。
+
+数据集可以在竞赛[主页](https://www.isprs.org/education/benchmarks/UrbanSemLab/default.aspx)上请求获得。
+这里也提供了[BaiduNetdisk](https://pan.baidu.com/s/1K-cLVZnd1X7d8c26FQ-nGg?pwd=mseg),提取码:mseg、 [Google Drive](https://drive.google.com/drive/folders/1w3EJuyUGet6_qmLwGAWZ9vw5ogeG0zLz?usp=sharing)以及[OpenDataLab](https://opendatalab.com/ISPRS_Potsdam/download)。
+实验中需要下载 '2_Ortho_RGB.zip' 和 '5_Labels_all_noBoundary.zip'。
+
+对于 Potsdam 数据集,请运行以下命令调整数据集目录。
+
+```shell
+python tools/dataset_converters/potsdam.py /path/to/potsdam
+```
+
+在我们的默认设置中,将生成 3456 张图像用于训练和 2016 张图像用于验证。
+
+## ISPRS Vaihingen
+
+[Vaihingen](https://www.isprs.org/education/benchmarks/UrbanSemLab/2d-sem-label-vaihingen.aspx) 城市语义分割数据集用于 2D 语义分割竞赛 —— Vaihingen。
+
+数据集可以在竞赛[主页](https://www.isprs.org/education/benchmarks/UrbanSemLab/default.aspx)上请求获得。
+这里也提供了[BaiduNetdisk](https://pan.baidu.com/s/109D3WLrLafsuYtLeerLiiA?pwd=mseg),提取码:mseg 、 [Google Drive](https://drive.google.com/drive/folders/1w3NhvLVA2myVZqOn2pbiDXngNC7NTP_t?usp=sharing)。
+实验中需要下载 'ISPRS_semantic_labeling_Vaihingen.zip' 和 'ISPRS_semantic_labeling_Vaihingen_ground_truth_eroded_COMPLETE.zip'。
+
+对于 Vaihingen 数据集,请运行以下命令调整数据集目录。
+
+```shell
+python tools/dataset_converters/vaihingen.py /path/to/vaihingen
+```
+
+在我们的默认设置(`clip_size`=512, `stride_size`=256)中,将生成 344 张图像用于训练和 398 张图像用于验证。
+
+## iSAID
+
+iSAID 数据集可从 [DOTA-v1.0](https://captain-whu.github.io/DOTA/dataset.html) 下载训练/验证/测试数据集的图像数据,
+
+并从 [iSAID](https://captain-whu.github.io/iSAID/dataset.html)下载训练/验证数据集的标注数据。
+
+该数据集是航空图像实例分割和语义分割任务的大规模数据集。
+
+下载 iSAID 数据集后,您可能需要按照以下结构进行数据集准备。
+
+```none
+├── data
+│ ├── iSAID
+│ │ ├── train
+│ │ │ ├── images
+│ │ │ │ ├── part1.zip
+│ │ │ │ ├── part2.zip
+│ │ │ │ ├── part3.zip
+│ │ │ ├── Semantic_masks
+│ │ │ │ ├── images.zip
+│ │ ├── val
+│ │ │ ├── images
+│ │ │ │ ├── part1.zip
+│ │ │ ├── Semantic_masks
+│ │ │ │ ├── images.zip
+│ │ ├── test
+│ │ │ ├── images
+│ │ │ │ ├── part1.zip
+│ │ │ │ ├── part2.zip
+```
+
+```shell
+python tools/dataset_converters/isaid.py /path/to/iSAID
+```
+
+在我们的默认设置(`patch_width`=896, `patch_height`=896, `overlap_area`=384)中,将生成 33978 张图像用于训练和 11644 张图像用于验证。
+
+## LIP(Look Into Person) dataset
+
+该数据集可以从[此页面](https://lip.sysuhcp.com/overview.php)下载。
+
+请运行以下命令来解压数据集。
+
+```shell
+unzip LIP.zip
+cd LIP
+unzip TrainVal_images.zip
+unzip TrainVal_parsing_annotations.zip
+cd TrainVal_parsing_annotations
+unzip TrainVal_parsing_annotations.zip
+mv train_segmentations ../
+mv val_segmentations ../
+cd ..
+```
+
+LIP 数据集的内容包括:
+
+```none
+├── data
+│ ├── LIP
+│ │ ├── train_images
+│ │ │ ├── 1000_1234574.jpg
+│ │ │ ├── ...
+│ │ ├── train_segmentations
+│ │ │ ├── 1000_1234574.png
+│ │ │ ├── ...
+│ │ ├── val_images
+│ │ │ ├── 100034_483681.jpg
+│ │ │ ├── ...
+│ │ ├── val_segmentations
+│ │ │ ├── 100034_483681.png
+│ │ │ ├── ...
+```
+
+## Synapse dataset
+
+此数据集可以从[此页面](https://www.synapse.org/#!Synapse:syn3193805/wiki/)下载。
+
+遵循 [TransUNet](https://arxiv.org/abs/2102.04306) 的数据准备设定,将原始训练集(30 次扫描)拆分为新的训练集(18 次扫描)和验证集(12 次扫描)。请运行以下命令来准备数据集。
+
+```shell
+unzip RawData.zip
+cd ./RawData/Training
+```
+
+然后创建 `train.txt` 和 `val.txt` 以拆分数据集。
+
+根据 TransUnet,以下是数据集的划分。
+
+train.txt
+
+```none
+img0005.nii.gz
+img0006.nii.gz
+img0007.nii.gz
+img0009.nii.gz
+img0010.nii.gz
+img0021.nii.gz
+img0023.nii.gz
+img0024.nii.gz
+img0026.nii.gz
+img0027.nii.gz
+img0028.nii.gz
+img0030.nii.gz
+img0031.nii.gz
+img0033.nii.gz
+img0034.nii.gz
+img0037.nii.gz
+img0039.nii.gz
+img0040.nii.gz
+```
+
+val.txt
+
+```none
+img0008.nii.gz
+img0022.nii.gz
+img0038.nii.gz
+img0036.nii.gz
+img0032.nii.gz
+img0002.nii.gz
+img0029.nii.gz
+img0003.nii.gz
+img0001.nii.gz
+img0004.nii.gz
+img0025.nii.gz
+img0035.nii.gz
+```
+
+synapse 数据集的内容包括:
+
+```none
+├── Training
+│ ├── img
+│ │ ├── img0001.nii.gz
+│ │ ├── img0002.nii.gz
+│ │ ├── ...
+│ ├── label
+│ │ ├── label0001.nii.gz
+│ │ ├── label0002.nii.gz
+│ │ ├── ...
+│ ├── train.txt
+│ ├── val.txt
+```
+
+然后,使用此命令转换 synapse 数据集。
+
+```shell
+python tools/dataset_converters/synapse.py --dataset-path /path/to/synapse
+```
+
+注意,MMSegmentation 的默认评估指标(例如 mean dice value)是在 2D 切片图像上计算的,这与 [TransUNet](https://arxiv.org/abs/2102.04306) 等一些论文中的 3D 扫描结果是不同的。
+
+## REFUGE
+
+在 [REFUGE Challenge](https://refuge.grand-challenge.org) 官网上注册并下载 [REFUGE 数据集](https://refuge.grand-challenge.org/REFUGE2Download)。
+
+然后,解压 `REFUGE2.zip`,原始数据集的内容包括:
+
+```none
+├── REFUGE2
+│ ├── REFUGE2
+│ │ ├── Annotation-Training400.zip
+│ │ ├── REFUGE-Test400.zip
+│ │ ├── REFUGE-Test-GT.zip
+│ │ ├── REFUGE-Training400.zip
+│ │ ├── REFUGE-Validation400.zip
+│ │ ├── REFUGE-Validation400-GT.zip
+│ ├── __MACOSX
+```
+
+请运行以下命令转换 REFUGE 数据集:
+
+```shell
+python tools/convert_datasets/refuge.py --raw_data_root=/path/to/refuge/REFUGE2/REFUGE2
+```
+
+脚本会将目录结构转换如下:
+
+```none
+│ ├── REFUGE
+│ │ ├── images
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ │ ├── test
+│ │ ├── annotations
+│ │ │ ├── training
+│ │ │ ├── validation
+│ │ │ ├── test
+```
+
+包含 400 张用于训练的图像、400 张用于验证的图像和 400 张用于测试的图像,这与 REFUGE 2018 数据集相同。
+
+## Mapillary Vistas Datasets
+
+- Mapillary Vistas [官方网站](https://www.mapillary.com/dataset/vistas) 可以下载 Mapillary Vistas 数据集,按照官网要求注册并登陆后,数据可以在[这里](https://www.mapillary.com/dataset/vistas)找到。
+
+- Mapillary Vistas 数据集使用 8-bit with color-palette 来存储标签。不需要进行转换操作。
+
+- 假设您已将数据集 zip 文件放在 `mmsegmentation/data/mapillary` 中
+
+- 请运行以下命令来解压数据集。
+
+ ```bash
+ cd data/mapillary
+ unzip An-ZjB1Zm61yAZG0ozTymz8I8NqI4x0MrYrh26dq7kPgfu8vf9ImrdaOAVOFYbJ2pNAgUnVGBmbue9lTgdBOb5BbKXIpFs0fpYWqACbrQDChAA2fdX0zS9PcHu7fY8c-FOvyBVxPNYNFQuM.zip
+ ```
+
+- 解压后,您将获得类似于此结构的 Mapillary Vistas 数据集。语义分割 mask 标签在 `labels` 文件夹中。
+
+ ```none
+ mmsegmentation
+ ├── mmseg
+ ├── tools
+ ├── configs
+ ├── data
+ │ ├── mapillary
+ │ │ ├── training
+ │ │ │ ├── images
+ │ │ │ ├── v1.2
+ | │ │ │ ├── instances
+ | │ │ │ ├── labels
+ | │ │ │ └── panoptic
+ │ │ │ ├── v2.0
+ | │ │ │ ├── instances
+ | │ │ │ ├── labels
+ | │ │ │ ├── panoptic
+ | │ │ │ └── polygons
+ │ │ ├── validation
+ │ │ │ ├── images
+ | │ │ ├── v1.2
+ | │ │ │ ├── instances
+ | │ │ │ ├── labels
+ | │ │ │ └── panoptic
+ │ │ │ ├── v2.0
+ | │ │ │ ├── instances
+ | │ │ │ ├── labels
+ | │ │ │ ├── panoptic
+ | │ │ │ └── polygons
+ ```
+
+- 您可以在配置中使用 `MapillaryDataset_v1` 和 `Mapillary Dataset_v2` 设置数据集版本。
+ 在此处 [V1.2](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/datasets/mapillary_v1.py) 和 [V2.0](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/datasets/mapillary_v2.py) 查看 Mapillary Vistas 数据集配置文件
+
+## LEVIR-CD
+
+[LEVIR-CD](https://justchenhao.github.io/LEVIR/) 大规模遥感建筑变化检测数据集。
+
+数据集可以在[主页](https://justchenhao.github.io/LEVIR/)上请求获得。
+
+数据集的补充版本可以在[主页](https://github.com/S2Looking/Dataset)上请求获得。
+
+请下载数据集的补充版本,然后解压 `LEVIR-CD+.zip`,数据集的内容包括:
+
+```none
+│ ├── LEVIR-CD+
+│ │ ├── train
+│ │ │ ├── A
+│ │ │ ├── B
+│ │ │ ├── label
+│ │ ├── test
+│ │ │ ├── A
+│ │ │ ├── B
+│ │ │ ├── label
+```
+
+对于 LEVIR-CD 数据集,请运行以下命令无重叠裁剪影像:
+
+```shell
+python tools/dataset_converters/levircd.py --dataset-path /path/to/LEVIR-CD+ --out_dir /path/to/LEVIR-CD
+```
+
+裁剪后的影像大小为256x256,与原论文保持一致。
+
+## BDD100K
+
+- 可以从[官方网站](https://bdd-data.berkeley.edu/) 下载 BDD100K数据集(语义分割任务主要是10K数据集),按照官网要求注册并登陆后,数据可以在[这里](https://bdd-data.berkeley.edu/portal.html#download)找到。
+
+- 图像数据对应的名称是是`10K Images`, 语义分割标注对应的名称是`Segmentation`
+
+- 下载后,可以使用以下代码进行解压
+
+ ```bash
+ unzip ~/bdd100k_images_10k.zip -d ~/mmsegmentation/data/
+ unzip ~/bdd100k_sem_seg_labels_trainval.zip -d ~/mmsegmentation/data/
+ ```
+
+就可以得到以下文件结构了:
+
+```none
+mmsegmentation
+├── mmseg
+├── tools
+├── configs
+├── data
+│ ├── bdd100k
+│ │ ├── images
+│ │ │ └── 10k
+| │ │ │ ├── test
+| │ │ │ ├── train
+| │ │ │ └── val
+│ │ └── labels
+│ │ │ └── sem_seg
+| │ │ │ ├── colormaps
+| │ │ │ │ ├──train
+| │ │ │ │ └──val
+| │ │ │ ├── masks
+| │ │ │ │ ├──train
+| │ │ │ │ └──val
+| │ │ │ ├── polygons
+| │ │ │ │ ├──sem_seg_train.json
+| │ │ │ │ └──sem_seg_val.json
+| │ │ │ └── rles
+| │ │ │ │ ├──sem_seg_train.json
+| │ │ │ │ └──sem_seg_val.json
+```
+
+## NYU
+
+- 您可以从 [这个链接](https://drive.google.com/file/d/1wC-io-14RCIL4XTUrQLk6lBqU2AexLVp/view?usp=share_link) 下载 NYU 数据集
+
+- 下载完成后,您可以使用 [tools/dataset_converters/nyu.py](/tools/dataset_converters/nyu.py) 脚本来解压和组织数据到所需的格式
+
+ ```bash
+ python tools/dataset_converters/nyu.py nyu.zip
+ ```
diff --git a/docs/zh_cn/user_guides/3_inference.md b/docs/zh_cn/user_guides/3_inference.md
index d2fe60076f8..0afcb4b05d6 100644
--- a/docs/zh_cn/user_guides/3_inference.md
+++ b/docs/zh_cn/user_guides/3_inference.md
@@ -1,3 +1,244 @@
-## 使用预训练模型推理(待更新)
+# 教程3:使用预训练模型推理
-中文版文档支持中,请先阅读[英文版本](../../en/user_guides/3_inference.md)
+MMSegmentation 在 [Model Zoo](../Model_Zoo.md) 中为语义分割提供了预训练的模型,并支持多个标准数据集,包括 Cityscapes、ADE20K 等。
+本说明将展示如何使用现有模型对给定图像进行推理。
+关于如何在标准数据集上测试现有模型,请参阅本[指南](./4_train_test.md)
+
+MMSegmentation 为用户提供了数个接口,以便轻松使用预训练的模型进行推理。
+
+- [教程3:使用预训练模型推理](#教程3使用预训练模型推理)
+ - [推理器](#推理器)
+ - [基本使用](#基本使用)
+ - [初始化](#初始化)
+ - [可视化预测结果](#可视化预测结果)
+ - [模型列表](#模型列表)
+ - [推理 API](#推理-api)
+ - [mmseg.apis.init_model](#mmsegapisinit_model)
+ - [mmseg.apis.inference_model](#mmsegapisinference_model)
+ - [mmseg.apis.show_result_pyplot](#mmsegapisshow_result_pyplot)
+
+## 推理器
+
+在 MMSegmentation 中,我们提供了最**方便的**方式 `MMSegInferencer` 来使用模型。您只需 3 行代码就可以获得图像的分割掩膜。
+
+### 基本使用
+
+以下示例展示了如何使用 `MMSegInferencer` 对单个图像执行推理。
+
+```
+>>> from mmseg.apis import MMSegInferencer
+>>> # 将模型加载到内存中
+>>> inferencer = MMSegInferencer(model='deeplabv3plus_r18-d8_4xb2-80k_cityscapes-512x1024')
+>>> # 推理
+>>> inferencer('demo/demo.png', show=True)
+```
+
+可视化结果应如下所示:
+
+
+

+
 Abstract
+
+We address the problem of estimating a high quality dense depth map from a single RGB input image. We start out with a baseline encoder-decoder convolutional neural network architecture and pose the question of how the global processing of information can help improve overall depth estimation. To this end, we propose a transformer-based architecture block that divides the depth range into bins whose center value is estimated adaptively per image. The final depth values are estimated as linear combinations of the bin centers. We call our new building block AdaBins. Our results show a decisive improvement over the state-of-the-art on several popular depth datasets across all metrics.We also validate the effectiveness of the proposed block with an ablation study and provide the code and corresponding pre-trained weights of the new state-of-the-art model.
+
+Our main contributions are the following:
+
+- We propose an architecture building block that performs global processing of the scene’s information.We propose to divide the predicted depth range into bins where the bin widths change per image. The final depth estimation is a linear combination of the bin center values.
+- We show a decisive improvement for supervised single image depth estimation across all metrics for the two most popular datasets, NYU and KITTI.
+- We analyze our findings and investigate different modifications on the proposed AdaBins block and study their effect on the accuracy of the depth estimation.
+
+
Abstract
+
+We address the problem of estimating a high quality dense depth map from a single RGB input image. We start out with a baseline encoder-decoder convolutional neural network architecture and pose the question of how the global processing of information can help improve overall depth estimation. To this end, we propose a transformer-based architecture block that divides the depth range into bins whose center value is estimated adaptively per image. The final depth values are estimated as linear combinations of the bin centers. We call our new building block AdaBins. Our results show a decisive improvement over the state-of-the-art on several popular depth datasets across all metrics.We also validate the effectiveness of the proposed block with an ablation study and provide the code and corresponding pre-trained weights of the new state-of-the-art model.
+
+Our main contributions are the following:
+
+- We propose an architecture building block that performs global processing of the scene’s information.We propose to divide the predicted depth range into bins where the bin widths change per image. The final depth estimation is a linear combination of the bin center values.
+- We show a decisive improvement for supervised single image depth estimation across all metrics for the two most popular datasets, NYU and KITTI.
+- We analyze our findings and investigate different modifications on the proposed AdaBins block and study their effect on the accuracy of the depth estimation.
+
+
+

+
 Performance
+
+### NYU and KITTI
+
+| Model | Encoder | Training epoch | Batchsize | Train Resolution | δ1 | δ2 | δ3 | REL | RMS | RMS log | params(M) | Links |
+| ------------- | --------------- | -------------- | --------- | ---------------- | ----- | ----- | ----- | ----- | ----- | ------- | --------- | ----------------------------------------------------------------------------------------------------------------------- |
+| AdaBins_nyu | EfficientNet-B5 | 25 | 16 | 416x544 | 0.903 | 0.984 | 0.997 | 0.103 | 0.364 | 0.044 | 78 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/adabins/adabins_efficient_b5_nyu_third-party-f68d6bd3.pth) |
+| AdaBins_kitti | EfficientNet-B5 | 25 | 16 | 352x764 | 0.964 | 0.995 | 0.999 | 0.058 | 2.360 | 0.088 | 78 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/adabins/adabins_efficient-b5_kitty_third-party-a1aa6f36.pth) |
+
+## Citation
+
+```bibtex
+@article{10.1109/cvpr46437.2021.00400,
+ author = {Bhat, S. A. and Alhashim, I. and Wonka, P.},
+ title = {Adabins: depth estimation using adaptive bins},
+ journal = {2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR)},
+ year = {2021},
+ doi = {10.1109/cvpr46437.2021.00400}
+}
+```
diff --git a/projects/Adabins/backbones/__init__.py b/projects/Adabins/backbones/__init__.py
new file mode 100644
index 00000000000..04ae180be5f
--- /dev/null
+++ b/projects/Adabins/backbones/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .adabins_backbone import AdabinsBackbone
+
+__all__ = ['AdabinsBackbone']
diff --git a/projects/Adabins/backbones/adabins_backbone.py b/projects/Adabins/backbones/adabins_backbone.py
new file mode 100644
index 00000000000..07d73809e39
--- /dev/null
+++ b/projects/Adabins/backbones/adabins_backbone.py
@@ -0,0 +1,141 @@
+import timm
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import ConvModule, build_conv_layer
+from mmengine.model import BaseModule
+
+from mmseg.registry import MODELS
+
+
+class UpSampleBN(nn.Module):
+ """ UpSample module
+ Args:
+ skip_input (int): the input feature
+ output_features (int): the output feature
+ norm_cfg (dict, optional): Config dict for normalization layer.
+ Default: dict(type='BN', requires_grad=True).
+ act_cfg (dict, optional): The activation layer of AAM:
+ Aggregate Attention Module.
+ """
+
+ def __init__(self,
+ skip_input,
+ output_features,
+ norm_cfg=dict(type='BN'),
+ act_cfg=dict(type='LeakyReLU')):
+ super().__init__()
+
+ self._net = nn.Sequential(
+ ConvModule(
+ in_channels=skip_input,
+ out_channels=output_features,
+ kernel_size=3,
+ stride=1,
+ padding=1,
+ bias=True,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ),
+ ConvModule(
+ in_channels=output_features,
+ out_channels=output_features,
+ kernel_size=3,
+ stride=1,
+ padding=1,
+ bias=True,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ))
+
+ def forward(self, x, concat_with):
+ up_x = F.interpolate(
+ x,
+ size=[concat_with.size(2),
+ concat_with.size(3)],
+ mode='bilinear',
+ align_corners=True)
+ f = torch.cat([up_x, concat_with], dim=1)
+ return self._net(f)
+
+
+class Encoder(nn.Module):
+ """ the efficientnet_b5 model
+ Args:
+ basemodel_name (str): the name of base model
+ """
+
+ def __init__(self, basemodel_name):
+ super().__init__()
+ self.original_model = timm.create_model(
+ basemodel_name, pretrained=True)
+ # Remove last layer
+ self.original_model.global_pool = nn.Identity()
+ self.original_model.classifier = nn.Identity()
+
+ def forward(self, x):
+ features = [x]
+ for k, v in self.original_model._modules.items():
+ if k == 'blocks':
+ for ki, vi in v._modules.items():
+ features.append(vi(features[-1]))
+ else:
+ features.append(v(features[-1]))
+ return features
+
+
+@MODELS.register_module()
+class AdabinsBackbone(BaseModule):
+ """ the backbone of the adabins
+ Args:
+ basemodel_name (str):the name of base model
+ num_features (int): the middle feature
+ num_classes (int): the classes number
+ bottleneck_features (int): the bottleneck features
+ conv_cfg (dict): Config dict for convolution layer.
+ """
+
+ def __init__(self,
+ basemodel_name,
+ num_features=2048,
+ num_classes=128,
+ bottleneck_features=2048,
+ conv_cfg=dict(type='Conv')):
+ super().__init__()
+ self.encoder = Encoder(basemodel_name)
+ features = int(num_features)
+ self.conv2 = build_conv_layer(
+ conv_cfg,
+ bottleneck_features,
+ features,
+ kernel_size=1,
+ stride=1,
+ padding=1)
+ self.up1 = UpSampleBN(
+ skip_input=features // 1 + 112 + 64, output_features=features // 2)
+ self.up2 = UpSampleBN(
+ skip_input=features // 2 + 40 + 24, output_features=features // 4)
+ self.up3 = UpSampleBN(
+ skip_input=features // 4 + 24 + 16, output_features=features // 8)
+ self.up4 = UpSampleBN(
+ skip_input=features // 8 + 16 + 8, output_features=features // 16)
+
+ self.conv3 = build_conv_layer(
+ conv_cfg,
+ features // 16,
+ num_classes,
+ kernel_size=3,
+ stride=1,
+ padding=1)
+
+ def forward(self, x):
+ features = self.encoder(x)
+ x_block0, x_block1, x_block2, x_block3, x_block4 = features[
+ 3], features[4], features[5], features[7], features[10]
+ x_d0 = self.conv2(x_block4)
+ x_d1 = self.up1(x_d0, x_block3)
+ x_d2 = self.up2(x_d1, x_block2)
+ x_d3 = self.up3(x_d2, x_block1)
+ x_d4 = self.up4(x_d3, x_block0)
+ out = self.conv3(x_d4)
+ return out
diff --git a/projects/Adabins/configs/_base_/datasets/nyu.py b/projects/Adabins/configs/_base_/datasets/nyu.py
new file mode 100644
index 00000000000..1b49ec7e8de
--- /dev/null
+++ b/projects/Adabins/configs/_base_/datasets/nyu.py
@@ -0,0 +1,32 @@
+dataset_type = 'NYUDataset'
+data_root = 'data/nyu'
+
+test_pipeline = [
+ dict(dict(type='LoadImageFromFile', to_float32=True)),
+ dict(dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3)),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id'))
+]
+
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ test_mode=True,
+ data_prefix=dict(
+ img_path='images/test', depth_map_path='annotations/test'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(
+ type='DepthMetric', max_depth_eval=10.0, crop_type='nyu_crop')
+test_evaluator = val_evaluator
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
diff --git a/projects/Adabins/configs/_base_/default_runtime.py b/projects/Adabins/configs/_base_/default_runtime.py
new file mode 100644
index 00000000000..272b4d24679
--- /dev/null
+++ b/projects/Adabins/configs/_base_/default_runtime.py
@@ -0,0 +1,15 @@
+default_scope = 'mmseg'
+env_cfg = dict(
+ cudnn_benchmark=True,
+ mp_cfg=dict(mp_start_method='fork', opencv_num_threads=0),
+ dist_cfg=dict(backend='nccl'),
+)
+vis_backends = [dict(type='LocalVisBackend')]
+visualizer = dict(
+ type='SegLocalVisualizer', vis_backends=vis_backends, name='visualizer')
+log_processor = dict(by_epoch=False)
+log_level = 'INFO'
+load_from = None
+resume = False
+
+tta_model = dict(type='SegTTAModel')
diff --git a/projects/Adabins/configs/_base_/models/Adabins.py b/projects/Adabins/configs/_base_/models/Adabins.py
new file mode 100644
index 00000000000..35cbd8c5777
--- /dev/null
+++ b/projects/Adabins/configs/_base_/models/Adabins.py
@@ -0,0 +1,35 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255)
+model = dict(
+ type='DepthEstimator',
+ data_preprocessor=data_preprocessor,
+ # pretrained='open-mmlab://resnet50_v1c',
+ backbone=dict(
+ type='AdabinsBackbone',
+ basemodel_name='tf_efficientnet_b5_ap',
+ num_features=2048,
+ num_classes=128,
+ bottleneck_features=2048,
+ ),
+ decode_head=dict(
+ type='AdabinsHead',
+ in_channels=128,
+ n_query_channels=128,
+ patch_size=16,
+ embedding_dim=128,
+ num_heads=4,
+ n_bins=256,
+ min_val=0.001,
+ max_val=10,
+ norm='linear'),
+
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_NYU_416x544.py b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_NYU_416x544.py
new file mode 100644
index 00000000000..5c00ea152bf
--- /dev/null
+++ b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_NYU_416x544.py
@@ -0,0 +1,15 @@
+_base_ = [
+ '../_base_/models/Adabins.py', '../_base_/datasets/nyu.py',
+ '../_base_/default_runtime.py'
+]
+custom_imports = dict(
+ imports=['projects.Adabins.backbones', 'projects.Adabins.decode_head'],
+ allow_failed_imports=False)
+crop_size = (416, 544)
+data_preprocessor = dict(size=crop_size)
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(),
+ decode_head=dict(),
+)
diff --git a/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_kitti_352x704.py b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_kitti_352x704.py
new file mode 100644
index 00000000000..330cdf41a5b
--- /dev/null
+++ b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_kitti_352x704.py
@@ -0,0 +1,12 @@
+_base_ = ['../_base_/models/Adabins.py']
+custom_imports = dict(
+ imports=['projects.Adabins.backbones', 'projects.Adabins.decode_head'],
+ allow_failed_imports=False)
+crop_size = (352, 704)
+data_preprocessor = dict(size=crop_size)
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(),
+ decode_head=dict(min_val=0.001, max_val=80),
+)
diff --git a/projects/Adabins/decode_head/__init__.py b/projects/Adabins/decode_head/__init__.py
new file mode 100644
index 00000000000..c7d62df12bb
--- /dev/null
+++ b/projects/Adabins/decode_head/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .adabins_head import AdabinsHead
+
+__all__ = ['AdabinsHead']
diff --git a/projects/Adabins/decode_head/adabins_head.py b/projects/Adabins/decode_head/adabins_head.py
new file mode 100644
index 00000000000..ee043172ab9
--- /dev/null
+++ b/projects/Adabins/decode_head/adabins_head.py
@@ -0,0 +1,179 @@
+from typing import List, Tuple
+
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import build_conv_layer
+from torch import Tensor
+
+from mmseg.registry import MODELS
+
+
+class PatchTransformerEncoder(nn.Module):
+ """the Patch Transformer Encoder.
+
+ Args:
+ in_channels (int): the channels of input
+ patch_size (int): the path size
+ embedding_dim (int): The feature dimension.
+ num_heads (int): the number of encoder head
+ conv_cfg (dict): Config dict for convolution layer.
+ """
+
+ def __init__(self,
+ in_channels,
+ patch_size=10,
+ embedding_dim=128,
+ num_heads=4,
+ conv_cfg=dict(type='Conv')):
+ super().__init__()
+ encoder_layers = nn.TransformerEncoderLayer(
+ embedding_dim, num_heads, dim_feedforward=1024)
+ self.transformer_encoder = nn.TransformerEncoder(
+ encoder_layers, num_layers=4) # takes shape S,N,E
+
+ self.embedding_convPxP = build_conv_layer(
+ conv_cfg,
+ in_channels,
+ embedding_dim,
+ kernel_size=patch_size,
+ stride=patch_size)
+ self.positional_encodings = nn.Parameter(
+ torch.rand(500, embedding_dim), requires_grad=True)
+
+ def forward(self, x):
+ embeddings = self.embedding_convPxP(x).flatten(
+ 2) # .shape = n,c,s = n, embedding_dim, s
+ embeddings = embeddings + self.positional_encodings[:embeddings.shape[
+ 2], :].T.unsqueeze(0)
+
+ # change to S,N,E format required by transformer
+ embeddings = embeddings.permute(2, 0, 1)
+ x = self.transformer_encoder(embeddings) # .shape = S, N, E
+ return x
+
+
+class PixelWiseDotProduct(nn.Module):
+ """the pixel wise dot product."""
+
+ def __init__(self):
+ super().__init__()
+
+ def forward(self, x, K):
+ n, c, h, w = x.size()
+ _, cout, ck = K.size()
+ assert c == ck, 'Number of channels in x and Embedding dimension ' \
+ '(at dim 2) of K matrix must match'
+ y = torch.matmul(
+ x.view(n, c, h * w).permute(0, 2, 1),
+ K.permute(0, 2, 1)) # .shape = n, hw, cout
+ return y.permute(0, 2, 1).view(n, cout, h, w)
+
+
+@MODELS.register_module()
+class AdabinsHead(nn.Module):
+ """the head of the adabins,include mViT.
+
+ Args:
+ in_channels (int):the channels of the input
+ n_query_channels (int):the channels of the query
+ patch_size (int): the patch size
+ embedding_dim (int):The feature dimension.
+ num_heads (int):the number of head
+ n_bins (int):the number of bins
+ min_val (float): the min width of bin
+ max_val (float): the max width of bin
+ conv_cfg (dict): Config dict for convolution layer.
+ norm (str): the activate method
+ align_corners (bool, optional): Geometrically, we consider the pixels
+ of the input and output as squares rather than points.
+ """
+
+ def __init__(self,
+ in_channels,
+ n_query_channels=128,
+ patch_size=16,
+ embedding_dim=128,
+ num_heads=4,
+ n_bins=100,
+ min_val=0.1,
+ max_val=10,
+ conv_cfg=dict(type='Conv'),
+ norm='linear',
+ align_corners=False,
+ threshold=0):
+ super().__init__()
+ self.out_channels = n_bins
+ self.align_corners = align_corners
+ self.norm = norm
+ self.num_classes = n_bins
+ self.min_val = min_val
+ self.max_val = max_val
+ self.n_query_channels = n_query_channels
+ self.patch_transformer = PatchTransformerEncoder(
+ in_channels, patch_size, embedding_dim, num_heads)
+ self.dot_product_layer = PixelWiseDotProduct()
+ self.threshold = threshold
+ self.conv3x3 = build_conv_layer(
+ conv_cfg,
+ in_channels,
+ embedding_dim,
+ kernel_size=3,
+ stride=1,
+ padding=1)
+ self.regressor = nn.Sequential(
+ nn.Linear(embedding_dim, 256), nn.LeakyReLU(), nn.Linear(256, 256),
+ nn.LeakyReLU(), nn.Linear(256, n_bins))
+ self.conv_out = nn.Sequential(
+ build_conv_layer(conv_cfg, in_channels, n_bins, kernel_size=1),
+ nn.Softmax(dim=1))
+
+ def forward(self, x):
+ # n, c, h, w = x.size()
+ tgt = self.patch_transformer(x.clone()) # .shape = S, N, E
+
+ x = self.conv3x3(x)
+
+ regression_head, queries = tgt[0,
+ ...], tgt[1:self.n_query_channels + 1,
+ ...]
+
+ # Change from S, N, E to N, S, E
+ queries = queries.permute(1, 0, 2)
+ range_attention_maps = self.dot_product_layer(
+ x, queries) # .shape = n, n_query_channels, h, w
+
+ y = self.regressor(regression_head) # .shape = N, dim_out
+ if self.norm == 'linear':
+ y = torch.relu(y)
+ eps = 0.1
+ y = y + eps
+ elif self.norm == 'softmax':
+ return torch.softmax(y, dim=1), range_attention_maps
+ else:
+ y = torch.sigmoid(y)
+ bin_widths_normed = y / y.sum(dim=1, keepdim=True)
+ out = self.conv_out(range_attention_maps)
+
+ bin_widths = (self.max_val -
+ self.min_val) * bin_widths_normed # .shape = N, dim_out
+ bin_widths = F.pad(
+ bin_widths, (1, 0), mode='constant', value=self.min_val)
+ bin_edges = torch.cumsum(bin_widths, dim=1)
+
+ centers = 0.5 * (bin_edges[:, :-1] + bin_edges[:, 1:])
+ n, dim_out = centers.size()
+ centers = centers.view(n, dim_out, 1, 1)
+
+ pred = torch.sum(out * centers, dim=1, keepdim=True)
+ return bin_edges, pred
+
+ def predict(self, inputs: Tuple[Tensor], batch_img_metas: List[dict],
+ test_cfg, **kwargs) -> Tensor:
+ """Forward function for testing, only ``pam_cam`` is used."""
+ pred = self.forward(inputs)[-1]
+ final = torch.clamp(pred, self.min_val, self.max_val)
+
+ final[torch.isinf(final)] = self.max_val
+ final[torch.isnan(final)] = self.min_val
+ return final
diff --git a/projects/CAT-Seg/README.md b/projects/CAT-Seg/README.md
new file mode 100644
index 00000000000..890e461ce4c
--- /dev/null
+++ b/projects/CAT-Seg/README.md
@@ -0,0 +1,92 @@
+# CAT-Seg
+
+> [CAT-Seg: Cost Aggregation for Open-Vocabulary Semantic Segmentation](https://arxiv.org/abs/2303.11797)
+
+## Introduction
+
+
+
+Official Repo
+
+Code Snippet
+
+## Abstract
+
+
+
+Existing works on open-vocabulary semantic segmentation have utilized large-scale vision-language models, such as CLIP, to leverage their exceptional open-vocabulary recognition capabilities. However, the problem of transferring these capabilities learned from image-level supervision to the pixel-level task of segmentation and addressing arbitrary unseen categories at inference makes this task challenging. To address these issues, we aim to attentively relate objects within an image to given categories by leveraging relational information among class categories and visual semantics through aggregation, while also adapting the CLIP representations to the pixel-level task. However, we observe that direct optimization of the CLIP embeddings can harm its open-vocabulary capabilities. In this regard, we propose an alternative approach to optimize the imagetext similarity map, i.e. the cost map, using a novel cost aggregation-based method. Our framework, namely CATSeg, achieves state-of-the-art performance across all benchmarks. We provide extensive ablation studies to validate our choices. [Project page](https://ku-cvlab.github.io/CAT-Seg).
+
+
+
+
Performance
+
+### NYU and KITTI
+
+| Model | Encoder | Training epoch | Batchsize | Train Resolution | δ1 | δ2 | δ3 | REL | RMS | RMS log | params(M) | Links |
+| ------------- | --------------- | -------------- | --------- | ---------------- | ----- | ----- | ----- | ----- | ----- | ------- | --------- | ----------------------------------------------------------------------------------------------------------------------- |
+| AdaBins_nyu | EfficientNet-B5 | 25 | 16 | 416x544 | 0.903 | 0.984 | 0.997 | 0.103 | 0.364 | 0.044 | 78 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/adabins/adabins_efficient_b5_nyu_third-party-f68d6bd3.pth) |
+| AdaBins_kitti | EfficientNet-B5 | 25 | 16 | 352x764 | 0.964 | 0.995 | 0.999 | 0.058 | 2.360 | 0.088 | 78 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/adabins/adabins_efficient-b5_kitty_third-party-a1aa6f36.pth) |
+
+## Citation
+
+```bibtex
+@article{10.1109/cvpr46437.2021.00400,
+ author = {Bhat, S. A. and Alhashim, I. and Wonka, P.},
+ title = {Adabins: depth estimation using adaptive bins},
+ journal = {2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR)},
+ year = {2021},
+ doi = {10.1109/cvpr46437.2021.00400}
+}
+```
diff --git a/projects/Adabins/backbones/__init__.py b/projects/Adabins/backbones/__init__.py
new file mode 100644
index 00000000000..04ae180be5f
--- /dev/null
+++ b/projects/Adabins/backbones/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .adabins_backbone import AdabinsBackbone
+
+__all__ = ['AdabinsBackbone']
diff --git a/projects/Adabins/backbones/adabins_backbone.py b/projects/Adabins/backbones/adabins_backbone.py
new file mode 100644
index 00000000000..07d73809e39
--- /dev/null
+++ b/projects/Adabins/backbones/adabins_backbone.py
@@ -0,0 +1,141 @@
+import timm
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import ConvModule, build_conv_layer
+from mmengine.model import BaseModule
+
+from mmseg.registry import MODELS
+
+
+class UpSampleBN(nn.Module):
+ """ UpSample module
+ Args:
+ skip_input (int): the input feature
+ output_features (int): the output feature
+ norm_cfg (dict, optional): Config dict for normalization layer.
+ Default: dict(type='BN', requires_grad=True).
+ act_cfg (dict, optional): The activation layer of AAM:
+ Aggregate Attention Module.
+ """
+
+ def __init__(self,
+ skip_input,
+ output_features,
+ norm_cfg=dict(type='BN'),
+ act_cfg=dict(type='LeakyReLU')):
+ super().__init__()
+
+ self._net = nn.Sequential(
+ ConvModule(
+ in_channels=skip_input,
+ out_channels=output_features,
+ kernel_size=3,
+ stride=1,
+ padding=1,
+ bias=True,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ),
+ ConvModule(
+ in_channels=output_features,
+ out_channels=output_features,
+ kernel_size=3,
+ stride=1,
+ padding=1,
+ bias=True,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ))
+
+ def forward(self, x, concat_with):
+ up_x = F.interpolate(
+ x,
+ size=[concat_with.size(2),
+ concat_with.size(3)],
+ mode='bilinear',
+ align_corners=True)
+ f = torch.cat([up_x, concat_with], dim=1)
+ return self._net(f)
+
+
+class Encoder(nn.Module):
+ """ the efficientnet_b5 model
+ Args:
+ basemodel_name (str): the name of base model
+ """
+
+ def __init__(self, basemodel_name):
+ super().__init__()
+ self.original_model = timm.create_model(
+ basemodel_name, pretrained=True)
+ # Remove last layer
+ self.original_model.global_pool = nn.Identity()
+ self.original_model.classifier = nn.Identity()
+
+ def forward(self, x):
+ features = [x]
+ for k, v in self.original_model._modules.items():
+ if k == 'blocks':
+ for ki, vi in v._modules.items():
+ features.append(vi(features[-1]))
+ else:
+ features.append(v(features[-1]))
+ return features
+
+
+@MODELS.register_module()
+class AdabinsBackbone(BaseModule):
+ """ the backbone of the adabins
+ Args:
+ basemodel_name (str):the name of base model
+ num_features (int): the middle feature
+ num_classes (int): the classes number
+ bottleneck_features (int): the bottleneck features
+ conv_cfg (dict): Config dict for convolution layer.
+ """
+
+ def __init__(self,
+ basemodel_name,
+ num_features=2048,
+ num_classes=128,
+ bottleneck_features=2048,
+ conv_cfg=dict(type='Conv')):
+ super().__init__()
+ self.encoder = Encoder(basemodel_name)
+ features = int(num_features)
+ self.conv2 = build_conv_layer(
+ conv_cfg,
+ bottleneck_features,
+ features,
+ kernel_size=1,
+ stride=1,
+ padding=1)
+ self.up1 = UpSampleBN(
+ skip_input=features // 1 + 112 + 64, output_features=features // 2)
+ self.up2 = UpSampleBN(
+ skip_input=features // 2 + 40 + 24, output_features=features // 4)
+ self.up3 = UpSampleBN(
+ skip_input=features // 4 + 24 + 16, output_features=features // 8)
+ self.up4 = UpSampleBN(
+ skip_input=features // 8 + 16 + 8, output_features=features // 16)
+
+ self.conv3 = build_conv_layer(
+ conv_cfg,
+ features // 16,
+ num_classes,
+ kernel_size=3,
+ stride=1,
+ padding=1)
+
+ def forward(self, x):
+ features = self.encoder(x)
+ x_block0, x_block1, x_block2, x_block3, x_block4 = features[
+ 3], features[4], features[5], features[7], features[10]
+ x_d0 = self.conv2(x_block4)
+ x_d1 = self.up1(x_d0, x_block3)
+ x_d2 = self.up2(x_d1, x_block2)
+ x_d3 = self.up3(x_d2, x_block1)
+ x_d4 = self.up4(x_d3, x_block0)
+ out = self.conv3(x_d4)
+ return out
diff --git a/projects/Adabins/configs/_base_/datasets/nyu.py b/projects/Adabins/configs/_base_/datasets/nyu.py
new file mode 100644
index 00000000000..1b49ec7e8de
--- /dev/null
+++ b/projects/Adabins/configs/_base_/datasets/nyu.py
@@ -0,0 +1,32 @@
+dataset_type = 'NYUDataset'
+data_root = 'data/nyu'
+
+test_pipeline = [
+ dict(dict(type='LoadImageFromFile', to_float32=True)),
+ dict(dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3)),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id'))
+]
+
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ test_mode=True,
+ data_prefix=dict(
+ img_path='images/test', depth_map_path='annotations/test'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(
+ type='DepthMetric', max_depth_eval=10.0, crop_type='nyu_crop')
+test_evaluator = val_evaluator
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
diff --git a/projects/Adabins/configs/_base_/default_runtime.py b/projects/Adabins/configs/_base_/default_runtime.py
new file mode 100644
index 00000000000..272b4d24679
--- /dev/null
+++ b/projects/Adabins/configs/_base_/default_runtime.py
@@ -0,0 +1,15 @@
+default_scope = 'mmseg'
+env_cfg = dict(
+ cudnn_benchmark=True,
+ mp_cfg=dict(mp_start_method='fork', opencv_num_threads=0),
+ dist_cfg=dict(backend='nccl'),
+)
+vis_backends = [dict(type='LocalVisBackend')]
+visualizer = dict(
+ type='SegLocalVisualizer', vis_backends=vis_backends, name='visualizer')
+log_processor = dict(by_epoch=False)
+log_level = 'INFO'
+load_from = None
+resume = False
+
+tta_model = dict(type='SegTTAModel')
diff --git a/projects/Adabins/configs/_base_/models/Adabins.py b/projects/Adabins/configs/_base_/models/Adabins.py
new file mode 100644
index 00000000000..35cbd8c5777
--- /dev/null
+++ b/projects/Adabins/configs/_base_/models/Adabins.py
@@ -0,0 +1,35 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255)
+model = dict(
+ type='DepthEstimator',
+ data_preprocessor=data_preprocessor,
+ # pretrained='open-mmlab://resnet50_v1c',
+ backbone=dict(
+ type='AdabinsBackbone',
+ basemodel_name='tf_efficientnet_b5_ap',
+ num_features=2048,
+ num_classes=128,
+ bottleneck_features=2048,
+ ),
+ decode_head=dict(
+ type='AdabinsHead',
+ in_channels=128,
+ n_query_channels=128,
+ patch_size=16,
+ embedding_dim=128,
+ num_heads=4,
+ n_bins=256,
+ min_val=0.001,
+ max_val=10,
+ norm='linear'),
+
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_NYU_416x544.py b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_NYU_416x544.py
new file mode 100644
index 00000000000..5c00ea152bf
--- /dev/null
+++ b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_NYU_416x544.py
@@ -0,0 +1,15 @@
+_base_ = [
+ '../_base_/models/Adabins.py', '../_base_/datasets/nyu.py',
+ '../_base_/default_runtime.py'
+]
+custom_imports = dict(
+ imports=['projects.Adabins.backbones', 'projects.Adabins.decode_head'],
+ allow_failed_imports=False)
+crop_size = (416, 544)
+data_preprocessor = dict(size=crop_size)
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(),
+ decode_head=dict(),
+)
diff --git a/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_kitti_352x704.py b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_kitti_352x704.py
new file mode 100644
index 00000000000..330cdf41a5b
--- /dev/null
+++ b/projects/Adabins/configs/adabins/adabins_efficient_b5_4x16_25e_kitti_352x704.py
@@ -0,0 +1,12 @@
+_base_ = ['../_base_/models/Adabins.py']
+custom_imports = dict(
+ imports=['projects.Adabins.backbones', 'projects.Adabins.decode_head'],
+ allow_failed_imports=False)
+crop_size = (352, 704)
+data_preprocessor = dict(size=crop_size)
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(),
+ decode_head=dict(min_val=0.001, max_val=80),
+)
diff --git a/projects/Adabins/decode_head/__init__.py b/projects/Adabins/decode_head/__init__.py
new file mode 100644
index 00000000000..c7d62df12bb
--- /dev/null
+++ b/projects/Adabins/decode_head/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .adabins_head import AdabinsHead
+
+__all__ = ['AdabinsHead']
diff --git a/projects/Adabins/decode_head/adabins_head.py b/projects/Adabins/decode_head/adabins_head.py
new file mode 100644
index 00000000000..ee043172ab9
--- /dev/null
+++ b/projects/Adabins/decode_head/adabins_head.py
@@ -0,0 +1,179 @@
+from typing import List, Tuple
+
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import build_conv_layer
+from torch import Tensor
+
+from mmseg.registry import MODELS
+
+
+class PatchTransformerEncoder(nn.Module):
+ """the Patch Transformer Encoder.
+
+ Args:
+ in_channels (int): the channels of input
+ patch_size (int): the path size
+ embedding_dim (int): The feature dimension.
+ num_heads (int): the number of encoder head
+ conv_cfg (dict): Config dict for convolution layer.
+ """
+
+ def __init__(self,
+ in_channels,
+ patch_size=10,
+ embedding_dim=128,
+ num_heads=4,
+ conv_cfg=dict(type='Conv')):
+ super().__init__()
+ encoder_layers = nn.TransformerEncoderLayer(
+ embedding_dim, num_heads, dim_feedforward=1024)
+ self.transformer_encoder = nn.TransformerEncoder(
+ encoder_layers, num_layers=4) # takes shape S,N,E
+
+ self.embedding_convPxP = build_conv_layer(
+ conv_cfg,
+ in_channels,
+ embedding_dim,
+ kernel_size=patch_size,
+ stride=patch_size)
+ self.positional_encodings = nn.Parameter(
+ torch.rand(500, embedding_dim), requires_grad=True)
+
+ def forward(self, x):
+ embeddings = self.embedding_convPxP(x).flatten(
+ 2) # .shape = n,c,s = n, embedding_dim, s
+ embeddings = embeddings + self.positional_encodings[:embeddings.shape[
+ 2], :].T.unsqueeze(0)
+
+ # change to S,N,E format required by transformer
+ embeddings = embeddings.permute(2, 0, 1)
+ x = self.transformer_encoder(embeddings) # .shape = S, N, E
+ return x
+
+
+class PixelWiseDotProduct(nn.Module):
+ """the pixel wise dot product."""
+
+ def __init__(self):
+ super().__init__()
+
+ def forward(self, x, K):
+ n, c, h, w = x.size()
+ _, cout, ck = K.size()
+ assert c == ck, 'Number of channels in x and Embedding dimension ' \
+ '(at dim 2) of K matrix must match'
+ y = torch.matmul(
+ x.view(n, c, h * w).permute(0, 2, 1),
+ K.permute(0, 2, 1)) # .shape = n, hw, cout
+ return y.permute(0, 2, 1).view(n, cout, h, w)
+
+
+@MODELS.register_module()
+class AdabinsHead(nn.Module):
+ """the head of the adabins,include mViT.
+
+ Args:
+ in_channels (int):the channels of the input
+ n_query_channels (int):the channels of the query
+ patch_size (int): the patch size
+ embedding_dim (int):The feature dimension.
+ num_heads (int):the number of head
+ n_bins (int):the number of bins
+ min_val (float): the min width of bin
+ max_val (float): the max width of bin
+ conv_cfg (dict): Config dict for convolution layer.
+ norm (str): the activate method
+ align_corners (bool, optional): Geometrically, we consider the pixels
+ of the input and output as squares rather than points.
+ """
+
+ def __init__(self,
+ in_channels,
+ n_query_channels=128,
+ patch_size=16,
+ embedding_dim=128,
+ num_heads=4,
+ n_bins=100,
+ min_val=0.1,
+ max_val=10,
+ conv_cfg=dict(type='Conv'),
+ norm='linear',
+ align_corners=False,
+ threshold=0):
+ super().__init__()
+ self.out_channels = n_bins
+ self.align_corners = align_corners
+ self.norm = norm
+ self.num_classes = n_bins
+ self.min_val = min_val
+ self.max_val = max_val
+ self.n_query_channels = n_query_channels
+ self.patch_transformer = PatchTransformerEncoder(
+ in_channels, patch_size, embedding_dim, num_heads)
+ self.dot_product_layer = PixelWiseDotProduct()
+ self.threshold = threshold
+ self.conv3x3 = build_conv_layer(
+ conv_cfg,
+ in_channels,
+ embedding_dim,
+ kernel_size=3,
+ stride=1,
+ padding=1)
+ self.regressor = nn.Sequential(
+ nn.Linear(embedding_dim, 256), nn.LeakyReLU(), nn.Linear(256, 256),
+ nn.LeakyReLU(), nn.Linear(256, n_bins))
+ self.conv_out = nn.Sequential(
+ build_conv_layer(conv_cfg, in_channels, n_bins, kernel_size=1),
+ nn.Softmax(dim=1))
+
+ def forward(self, x):
+ # n, c, h, w = x.size()
+ tgt = self.patch_transformer(x.clone()) # .shape = S, N, E
+
+ x = self.conv3x3(x)
+
+ regression_head, queries = tgt[0,
+ ...], tgt[1:self.n_query_channels + 1,
+ ...]
+
+ # Change from S, N, E to N, S, E
+ queries = queries.permute(1, 0, 2)
+ range_attention_maps = self.dot_product_layer(
+ x, queries) # .shape = n, n_query_channels, h, w
+
+ y = self.regressor(regression_head) # .shape = N, dim_out
+ if self.norm == 'linear':
+ y = torch.relu(y)
+ eps = 0.1
+ y = y + eps
+ elif self.norm == 'softmax':
+ return torch.softmax(y, dim=1), range_attention_maps
+ else:
+ y = torch.sigmoid(y)
+ bin_widths_normed = y / y.sum(dim=1, keepdim=True)
+ out = self.conv_out(range_attention_maps)
+
+ bin_widths = (self.max_val -
+ self.min_val) * bin_widths_normed # .shape = N, dim_out
+ bin_widths = F.pad(
+ bin_widths, (1, 0), mode='constant', value=self.min_val)
+ bin_edges = torch.cumsum(bin_widths, dim=1)
+
+ centers = 0.5 * (bin_edges[:, :-1] + bin_edges[:, 1:])
+ n, dim_out = centers.size()
+ centers = centers.view(n, dim_out, 1, 1)
+
+ pred = torch.sum(out * centers, dim=1, keepdim=True)
+ return bin_edges, pred
+
+ def predict(self, inputs: Tuple[Tensor], batch_img_metas: List[dict],
+ test_cfg, **kwargs) -> Tensor:
+ """Forward function for testing, only ``pam_cam`` is used."""
+ pred = self.forward(inputs)[-1]
+ final = torch.clamp(pred, self.min_val, self.max_val)
+
+ final[torch.isinf(final)] = self.max_val
+ final[torch.isnan(final)] = self.min_val
+ return final
diff --git a/projects/CAT-Seg/README.md b/projects/CAT-Seg/README.md
new file mode 100644
index 00000000000..890e461ce4c
--- /dev/null
+++ b/projects/CAT-Seg/README.md
@@ -0,0 +1,92 @@
+# CAT-Seg
+
+> [CAT-Seg: Cost Aggregation for Open-Vocabulary Semantic Segmentation](https://arxiv.org/abs/2303.11797)
+
+## Introduction
+
+
+
+Official Repo
+
+Code Snippet
+
+## Abstract
+
+
+
+Existing works on open-vocabulary semantic segmentation have utilized large-scale vision-language models, such as CLIP, to leverage their exceptional open-vocabulary recognition capabilities. However, the problem of transferring these capabilities learned from image-level supervision to the pixel-level task of segmentation and addressing arbitrary unseen categories at inference makes this task challenging. To address these issues, we aim to attentively relate objects within an image to given categories by leveraging relational information among class categories and visual semantics through aggregation, while also adapting the CLIP representations to the pixel-level task. However, we observe that direct optimization of the CLIP embeddings can harm its open-vocabulary capabilities. In this regard, we propose an alternative approach to optimize the imagetext similarity map, i.e. the cost map, using a novel cost aggregation-based method. Our framework, namely CATSeg, achieves state-of-the-art performance across all benchmarks. We provide extensive ablation studies to validate our choices. [Project page](https://ku-cvlab.github.io/CAT-Seg).
+
+
+
+
+

+CAT-Seg model structure
+
+

+

 diff --git a/projects/isnet/README.md b/projects/isnet/README.md
index 3a3172a9d99..0a79ad6a4fa 100644
--- a/projects/isnet/README.md
+++ b/projects/isnet/README.md
@@ -96,11 +96,11 @@ A project does not necessarily have to be finished in a single PR, but it's esse
- [ ] Type hints and docstrings
-
+
- [ ] Unit tests
-
+
- [ ] Code polishing
@@ -108,10 +108,10 @@ A project does not necessarily have to be finished in a single PR, but it's esse
- [ ] Metafile.yml
-
+
- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
-
+
- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/mapillary_dataset/README.md b/projects/mapillary_dataset/README.md
index cdc61d53a93..44a1e33ef93 100644
--- a/projects/mapillary_dataset/README.md
+++ b/projects/mapillary_dataset/README.md
@@ -10,7 +10,7 @@ This project implements **`Mapillary Vistas Dataset`**
### Dataset preparing
-Preparing `Mapillary Vistas Dataset` dataset following [Mapillary Vistas Dataset Preparing Guide](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md)
+Preparing `Mapillary Vistas Dataset` dataset following [Mapillary Vistas Dataset Preparing Guide](https://github.com/open-mmlab/mmsegmentation/tree/main/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md)
```none
mmsegmentation
diff --git a/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md b/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md
index fa074543300..c5cbc0f9b80 100644
--- a/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md
+++ b/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md
@@ -47,7 +47,7 @@
```
- You could set Datasets version with `MapillaryDataset_v1` and `MapillaryDataset_v2` in your configs.
- View the Mapillary Vistas Datasets config file here [V1.2](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/datasets/mapillary_v1.py) and [V2.0](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/datasets/mapillary_v2.py)
+ View the Mapillary Vistas Datasets config file here [V1.2](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/datasets/mapillary_v1.py) and [V2.0](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/datasets/mapillary_v2.py)
- **View datasets labels index and palette**
diff --git a/projects/medical/2d_image/ct/cranium/README.md b/projects/medical/2d_image/ct/cranium/README.md
new file mode 100644
index 00000000000..d3fa64ea406
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/README.md
@@ -0,0 +1,142 @@
+# Brain CT Images with Intracranial Hemorrhage Masks (Cranium)
+
+## Description
+
+This project supports **`Brain CT Images with Intracranial Hemorrhage Masks (Cranium)`**, which can be downloaded from [here](https://www.kaggle.com/datasets/vbookshelf/computed-tomography-ct-images).
+
+### Dataset Overview
+
+This dataset consists of head CT (Computed Thomography) images in jpg format. There are 2500 brain window images and 2500 bone window images, for 82 patients. There are approximately 30 image slices per patient. 318 images have associated intracranial image masks. Also included are csv files containing hemorrhage diagnosis data and patient data.
+This is version 1.0.0 of this dataset. A full description of this dataset as well as updated versions can be found here:
+https://physionet.org/content/ct-ich/1.0.0/
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ----------------------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------- |
+| [Cranium](https://www.kaggle.com/datasets/vbookshelf/computed-tomography-ct-images) | head_and_neck | segmentation | ct | 2 | 2501/-/- | yes/-/- | 2020 | [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 2501 | 99.93 | - | - | - | - |
+| hemorrhage | 318 | 0.07 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```
+@article{hssayeni2020computed,
+ title={Computed tomography images for intracranial hemorrhage detection and segmentation},
+ author={Hssayeni, Murtadha and Croock, MS and Salman, AD and Al-khafaji, HF and Yahya, ZA and Ghoraani, B},
+ journal={Intracranial Hemorrhage Segmentation Using A Deep Convolutional Model. Data},
+ volume={5},
+ number={1},
+ pages={179},
+ year={2020}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `cranium/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://www.kaggle.com/datasets/vbookshelf/computed-tomography-ct-images) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── ct
+ │ │ │ │ ├── cranium
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 2000 | 99.93 | 501 | 99.92 | - | - |
+| hemorrhage | 260 | 0.07 | 260 | 0.08 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/ct/cranium/configs/cranium_512x512.py b/projects/medical/2d_image/ct/cranium/configs/cranium_512x512.py
new file mode 100644
index 00000000000..d9b44362a5c
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/cranium_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'CraniumDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_cranium-512x512.py
new file mode 100644
index 00000000000..ac013a215ae
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_cranium-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_cranium-512x512.py
new file mode 100644
index 00000000000..c71110a21f7
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_cranium-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_cranium-512x512.py
new file mode 100644
index 00000000000..abbdac285b2
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_cranium-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_cranium-512x512.py
new file mode 100644
index 00000000000..418595268f9
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_cranium-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/datasets/cranium_dataset.py b/projects/medical/2d_image/ct/cranium/datasets/cranium_dataset.py
new file mode 100644
index 00000000000..d65f1cbfc68
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/datasets/cranium_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class CraniumDataset(BaseSegDataset):
+ """CraniumDataset dataset.
+
+ In segmentation map annotation for CraniumDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'hemorrhage'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/ct/cranium/tools/prepare_dataset.py b/projects/medical/2d_image/ct/cranium/tools/prepare_dataset.py
new file mode 100644
index 00000000000..1aa4e435614
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/tools/prepare_dataset.py
@@ -0,0 +1,66 @@
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+
+def read_single_array_from_pil(path):
+ return np.asarray(Image.open(path))
+
+
+def save_png_from_array(arr, save_path, mode=None):
+ Image.fromarray(arr, mode=mode).save(save_path)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+patients_dir = os.path.join(
+ root_path, 'Cranium/computed-tomography-images-for-' +
+ 'intracranial-hemorrhage-detection-and-segmentation-1.0.0' +
+ '/Patients_CT')
+
+patients = sorted(os.listdir(patients_dir))
+for p in patients:
+ data_dir = os.path.join(patients_dir, p, 'brain')
+ file_names = os.listdir(data_dir)
+ img_w_mask_names = [
+ _.replace('_HGE_Seg', '') for _ in file_names if 'Seg' in _
+ ]
+ img_wo_mask_names = [
+ _ for _ in file_names if _ not in img_w_mask_names and 'Seg' not in _
+ ]
+
+ for file_name in file_names:
+ path = os.path.join(data_dir, file_name)
+ img = read_single_array_from_pil(path)
+ tgt_name = file_name.replace('.jpg', img_suffix)
+ tgt_name = p + '_' + tgt_name
+ if 'Seg' in file_name: # is a mask
+ tgt_name = tgt_name.replace('_HGE_Seg', '')
+ mask_path = os.path.join(tgt_mask_dir, tgt_name)
+ mask = convert_label(img, convert_dict={0: 0, 255: 1})
+ save_png_from_array(mask, mask_path)
+ else:
+ img_path = os.path.join(tgt_img_dir, tgt_name)
+ pil = Image.fromarray(img).convert('RGB')
+ pil.save(img_path)
+
+ if file_name in img_wo_mask_names:
+ mask = np.zeros_like(img, dtype=np.uint8)
+ mask_path = os.path.join(tgt_mask_dir, tgt_name)
+ save_png_from_array(mask, mask_path)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/README.md b/projects/medical/2d_image/dermoscopy/isic2016_task1/README.md
new file mode 100644
index 00000000000..6e44e415ed6
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/README.md
@@ -0,0 +1,149 @@
+# ISIC-2016 Task1
+
+## Description
+
+This project support **`ISIC-2016 Task1 `**, and the dataset used in this project can be downloaded from [here](https://challenge.isic-archive.com/data/#2016).
+
+### Dataset Overview
+
+The overarching goal of the challenge is to develop image analysis tools to enable the automated diagnosis of melanoma from dermoscopic images.
+
+This challenge provides training data (~900 images) for participants to engage in all 3 components of lesion image analysis. A separate test dataset (~350 images) will be provided for participants to generate and submit automated results.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------- | ----------------- | ------------ | ---------- | ------------ | --------------------- | ---------------------- | ------------ | ---------------------------------------------------------------------- |
+| [ISIC-2016 Task1](https://challenge.isic-archive.com/data/#2016) | full body | segmentation | dermoscopy | 2 | 900/-/379- | yes/-/yes | 2016 | [CC-0](https://creativecommons.org/share-your-work/public-domain/cc0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 900 | 82.08 | - | - | 379 | 81.98 |
+| skin lesion | 900 | 17.92 | - | - | 379 | 18.02 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python 3.8
+- PyTorch 1.10.0
+- pillow(PIL) 9.3.0
+- scikit-learn(sklearn) 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of PYTHONPATH, which should point to the project's directory so that Python can locate the module files. In isic2016_task1/ root directory, run the following line to add the current directory to PYTHONPATH:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://challenge.isic-archive.com/data/#2016) and decompression data to path 'data/'.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── dermoscopy
+ │ │ │ │ ├── isic2016_task1
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+```
+
+### Training commands
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+To train on multiple GPUs, e.g. 8 GPUs, run the following command:
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH} --launcher pytorch --gpus 8
+```
+
+### Testing commands
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### ISIC-2016 Task1
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config |
+| :-------------: | :------: | :-------: | :----: | :--: | :---: | :---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py
new file mode 100644
index 00000000000..5638de4d560
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2016-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2016-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py
new file mode 100644
index 00000000000..bf17faa5380
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2016-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2016-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py
new file mode 100644
index 00000000000..f7bfcf6158f
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2016-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2016-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/isic2016-task1_512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/isic2016-task1_512x512.py
new file mode 100644
index 00000000000..029f5d4d7ec
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/isic2016-task1_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ISIC2017Task1'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='test.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/datasets/isic2016-task1_dataset.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/datasets/isic2016-task1_dataset.py
new file mode 100644
index 00000000000..8f11bdd0ba9
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/datasets/isic2016-task1_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ISIC2017Task1(BaseSegDataset):
+ """ISIC2017Task1 dataset.
+
+ In segmentation map annotation for ISIC2017Task1,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('normal', 'skin lesion'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/tools/prepare_dataset.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/tools/prepare_dataset.py
new file mode 100755
index 00000000000..ef4dad54086
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/tools/prepare_dataset.py
@@ -0,0 +1,120 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+
+
+def check_maskid(train_imgs):
+ for i in train_masks:
+ img = Image.open(i)
+ print(np.unique(np.array(img)))
+
+
+def reformulate_file(image_list, mask_list):
+ file_list = []
+ for idx, (imgp,
+ maskp) in enumerate(zip(sorted(image_list), sorted(mask_list))):
+ item = {'image': imgp, 'label': maskp}
+ file_list.append(item)
+ return file_list
+
+
+def check_file_exist(pair_list):
+ rel_path = os.getcwd()
+ for idx, sample in enumerate(pair_list):
+ image_path = sample['image']
+ assert os.path.exists(os.path.join(rel_path, image_path))
+ if 'label' in sample:
+ mask_path = sample['label']
+ assert os.path.exists(os.path.join(rel_path, mask_path))
+ print('all file path ok!')
+
+
+def convert_maskid(mask):
+ # add mask id conversion
+ arr_mask = np.array(mask).astype(np.uint8)
+ arr_mask[arr_mask == 255] = 1
+ return Image.fromarray(arr_mask)
+
+
+def process_dataset(file_lists, part_dir_dict):
+ for ith, part in enumerate(file_lists):
+ part_dir = part_dir_dict[ith]
+ for sample in part:
+ # read image and mask
+ image_path = sample['image']
+ if 'label' in sample:
+ mask_path = sample['label']
+
+ basename = os.path.basename(image_path)
+ targetname = basename.split('.')[0] # from image name
+
+ # check image file
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ targetname + save_img_suffix)
+ if not os.path.exists(img_save_path):
+ if not image_path.endswith('.png'):
+ src = Image.open(image_path)
+ src.save(img_save_path)
+ else:
+ shutil.copy(image_path, img_save_path)
+
+ if mask_path is not None:
+ mask_save_path = os.path.join(root_path, 'masks', part_dir,
+ targetname + save_seg_map_suffix)
+ if not os.path.exists(mask_save_path):
+ # check mask file
+ mask = Image.open(mask_path).convert('L')
+ # convert mask id
+ mask = convert_maskid(mask)
+ if not mask_path.endswith('.png'):
+ mask.save(mask_save_path)
+ else:
+ mask.save(mask_save_path)
+
+ # print image num
+ part_dir_folder = os.path.join(root_path, 'images', part_dir)
+ print(
+ f'{part_dir} has {len(os.listdir(part_dir_folder))} images completed!' # noqa
+ )
+
+
+if __name__ == '__main__':
+
+ root_path = 'data/' # original file
+ img_suffix = '.jpg'
+ seg_map_suffix = '.png'
+ save_img_suffix = '.png'
+ save_seg_map_suffix = '.png'
+
+ train_imgs = glob.glob('data/ISBI2016_ISIC_Part1_Training_Data/*' # noqa
+ + img_suffix)
+ train_masks = glob.glob(
+ 'data/ISBI2016_ISIC_Part1_Training_GroundTruth/*' # noqa
+ + seg_map_suffix)
+
+ test_imgs = glob.glob('data/ISBI2016_ISIC_Part1_Test_Data/*' + img_suffix)
+ test_masks = glob.glob(
+ 'data/ISBI2016_ISIC_Part1_Test_GroundTruth/*' # noqa
+ + seg_map_suffix)
+
+ assert len(train_imgs) == len(train_masks)
+ assert len(test_imgs) == len(test_masks)
+
+ print(f'training images: {len(train_imgs)}, test images: {len(test_imgs)}')
+
+ os.system('mkdir -p ' + root_path + 'images/train/')
+ os.system('mkdir -p ' + root_path + 'images/test/')
+ os.system('mkdir -p ' + root_path + 'masks/train/')
+ os.system('mkdir -p ' + root_path + 'masks/test/')
+
+ train_pair_list = reformulate_file(train_imgs, train_masks)
+ test_pair_list = reformulate_file(test_imgs, test_masks)
+
+ check_file_exist(train_pair_list)
+ check_file_exist(test_pair_list)
+
+ part_dir_dict = {0: 'train/', 1: 'test/'}
+ process_dataset([train_pair_list, test_pair_list], part_dir_dict)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/README.md b/projects/medical/2d_image/dermoscopy/isic2017_task1/README.md
new file mode 100644
index 00000000000..c7cc27096be
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/README.md
@@ -0,0 +1,158 @@
+# ISIC-2017 Task1
+
+## Description
+
+This project support **`ISIC-2017 Task1 `**, and the dataset used in this project can be downloaded from [here](https://challenge.isic-archive.com/data/#2017).
+
+### Dataset Overview
+
+The goal of the challenge is to help participants develop image analysis tools to enable the automated diagnosis of melanoma from dermoscopic images.
+
+This challenge provides training data (~2000 images) for participants to engage in all 3 components of lesion image analysis. A separate public validation dataset (~150 images) and blind held-out test dataset (~600 images) will be provided for participants to generate and submit automated results.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------- | ----------------- | ------------ | ---------- | ------------ | --------------------- | ---------------------- | ------------ | ---------------------------------------------------------------------- |
+| [ISIC-2017 Task1](https://challenge.isic-archive.com/data/#2017) | full body | segmentation | dermoscopy | 2 | 2000/150/600 | yes/yes/yes | 2017 | [CC-0](https://creativecommons.org/share-your-work/public-domain/cc0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 2000 | 82.86 | 150 | 73.88 | 600 | 70.62 |
+| skin lesion | 2000 | 17.14 | 150 | 26.12 | 600 | 29.38 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python 3.8
+- PyTorch 1.10.0
+- pillow(PIL) 9.3.0
+- scikit-learn(sklearn) 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of PYTHONPATH, which should point to the project's directory so that Python can locate the module files. In isic2017_task1/ root directory, run the following line to add the current directory to PYTHONPATH:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://challenge.isic-archive.com/data/#2017) and decompression data to path 'data/'.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── dermoscopy
+ │ │ │ │ ├── isic2017_task1
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── val
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── val
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+```
+
+### Training commands
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+To train on multiple GPUs, e.g. 8 GPUs, run the following command:
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH} --launcher pytorch --gpus 8
+```
+
+### Testing commands
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### ISIC-2017 Task1
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config |
+| :-------------: | :------: | :-------: | :----: | :--: | :---: | :---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py
new file mode 100644
index 00000000000..58d0a125d33
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2017-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2017-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py
new file mode 100644
index 00000000000..3becacf64fb
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2017-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2017-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py
new file mode 100644
index 00000000000..654ef4dc3d5
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2017-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2017-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/isic2017-task1_512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/isic2017-task1_512x512.py
new file mode 100644
index 00000000000..95997a10997
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/isic2017-task1_512x512.py
@@ -0,0 +1,41 @@
+dataset_type = 'ISIC2017Task1'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/train/', seg_map_path='masks/train/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(img_path='images/val/', seg_map_path='masks/val/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/datasets/isic2017-task1_dataset.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/datasets/isic2017-task1_dataset.py
new file mode 100644
index 00000000000..8f11bdd0ba9
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/datasets/isic2017-task1_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ISIC2017Task1(BaseSegDataset):
+ """ISIC2017Task1 dataset.
+
+ In segmentation map annotation for ISIC2017Task1,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('normal', 'skin lesion'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/tools/prepare_dataset.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/tools/prepare_dataset.py
new file mode 100755
index 00000000000..b3643c93590
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/tools/prepare_dataset.py
@@ -0,0 +1,127 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+
+
+def check_maskid(train_imgs):
+ for i in train_masks:
+ img = Image.open(i)
+ print(np.unique(np.array(img)))
+
+
+def reformulate_file(image_list, mask_list):
+ file_list = []
+ for idx, (imgp,
+ maskp) in enumerate(zip(sorted(image_list), sorted(mask_list))):
+ item = {'image': imgp, 'label': maskp}
+ file_list.append(item)
+ return file_list
+
+
+def convert_maskid(mask):
+ # add mask id conversion
+ arr_mask = np.array(mask).astype(np.uint8)
+ arr_mask[arr_mask == 255] = 1
+ return Image.fromarray(arr_mask)
+
+
+def check_file_exist(pair_list):
+ rel_path = os.getcwd()
+ for idx, sample in enumerate(pair_list):
+ image_path = sample['image']
+ assert os.path.exists(os.path.join(rel_path, image_path))
+ if 'label' in sample:
+ mask_path = sample['label']
+ assert os.path.exists(os.path.join(rel_path, mask_path))
+ print('all file path ok!')
+
+
+def process_dataset(file_lists, part_dir_dict):
+ for ith, part in enumerate(file_lists):
+ part_dir = part_dir_dict[ith]
+ for sample in part:
+ # read image and mask
+ image_path = sample['image']
+ if 'label' in sample:
+ mask_path = sample['label']
+
+ basename = os.path.basename(image_path)
+ targetname = basename.split('.')[0] # from image name
+
+ # check image file
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ targetname + save_img_suffix)
+ if not os.path.exists(img_save_path):
+ if not image_path.endswith('.png'):
+ src = Image.open(image_path)
+ src.save(img_save_path)
+ else:
+ shutil.copy(image_path, img_save_path)
+
+ if mask_path is not None:
+ mask_save_path = os.path.join(root_path, 'masks', part_dir,
+ targetname + save_seg_map_suffix)
+ if not os.path.exists(mask_save_path):
+ # check mask file
+ mask = Image.open(mask_path).convert('L')
+ # convert mask id
+ mask = convert_maskid(mask)
+ if not mask_path.endswith('.png'):
+ mask.save(mask_save_path)
+ else:
+ mask.save(mask_save_path)
+
+ # print image num
+ part_dir_folder = os.path.join(root_path, 'images', part_dir)
+ print(
+ f'{part_dir} has {len(os.listdir(part_dir_folder))} images completed!' # noqa
+ )
+
+
+if __name__ == '__main__':
+
+ root_path = 'data/' # original file
+ img_suffix = '.jpg'
+ seg_map_suffix = '.png'
+ save_img_suffix = '.png'
+ save_seg_map_suffix = '.png'
+
+ train_imgs = glob.glob('data/ISIC-2017_Training_Data/*' + img_suffix)
+ train_masks = glob.glob('data/ISIC-2017_Training_Part1_GroundTruth/*' +
+ seg_map_suffix)
+
+ val_imgs = glob.glob('data/ISIC-2017_Validation_Data/*' + img_suffix)
+ val_masks = glob.glob('data/ISIC-2017_Validation_Part1_GroundTruth/*' +
+ seg_map_suffix)
+
+ test_imgs = glob.glob('data/ISIC-2017_Test_v2_Data/*' + img_suffix)
+ test_masks = glob.glob('data/ISIC-2017_Test_v2_Part1_GroundTruth/*' +
+ seg_map_suffix)
+
+ assert len(train_imgs) == len(train_masks)
+ assert len(val_imgs) == len(val_masks)
+ assert len(test_imgs) == len(test_masks)
+
+ os.system('mkdir -p ' + root_path + 'images/train/')
+ os.system('mkdir -p ' + root_path + 'images/val/')
+ os.system('mkdir -p ' + root_path + 'images/test/')
+ os.system('mkdir -p ' + root_path + 'masks/train/')
+ os.system('mkdir -p ' + root_path + 'masks/val/')
+ os.system('mkdir -p ' + root_path + 'masks/test/')
+
+ part_dir_dict = {0: 'train/', 1: 'val/', 2: 'test/'}
+
+ train_pair_list = reformulate_file(train_imgs, train_masks)
+ val_pair_list = reformulate_file(val_imgs, val_masks)
+ test_pair_list = reformulate_file(test_imgs, test_masks)
+
+ check_file_exist(train_pair_list)
+ check_file_exist(val_pair_list)
+ check_file_exist(test_pair_list)
+
+ part_dir_dict = {0: 'train/', 1: 'val/', 2: 'test/'}
+ process_dataset([train_pair_list, val_pair_list, test_pair_list],
+ part_dir_dict)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/README.md b/projects/medical/2d_image/endoscopy/kvasir_seg/README.md
new file mode 100644
index 00000000000..ea597bc4401
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/README.md
@@ -0,0 +1,145 @@
+# Kvasir-Sessile Dataset (Kvasir SEG)
+
+## Description
+
+This project supports **`Kvasir-Sessile Dataset (Kvasir SEG) `**, which can be downloaded from [here](https://opendatalab.com/Kvasir-Sessile_dataset).
+
+## Dataset Overview
+
+The Kvasir-SEG dataset contains polyp images and their corresponding ground truth from the Kvasir Dataset v2. The resolution of the images contained in Kvasir-SEG varies from 332x487 to 1920x1072 pixels.
+
+
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------------- | ----------------- | ------------ | --------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------- |
+| [Kvarsir-SEG](https://opendatalab.com/Kvasir-Sessile_dataset) | abdomen | segmentation | endoscopy | 2 | 196/-/- | yes/-/- | 2020 | [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 196 | 92.31 | - | - | - | - |
+| polyp | 196 | 7.69 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{jha2020kvasir,
+ title={Kvasir-seg: A segmented polyp dataset},
+ author={Jha, Debesh and Smedsrud, Pia H and Riegler, Michael A and Halvorsen, P{\aa}l and Lange, Thomas de and Johansen, Dag and Johansen, H{\aa}vard D},
+ booktitle={International Conference on Multimedia Modeling},
+ pages={451--462},
+ year={2020},
+ organization={Springer}
+ }
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `kvasir_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.com/Kvasir-Sessile_dataset) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── endoscopy
+ │ │ │ │ ├── kvasir_seg
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 156 | 92.28 | 40 | 92.41 | - | - |
+| polyp | 156 | 7.72 | 40 | 7.59 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg .configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..145d5a7a172
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_kvasir-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..3ea05c51090
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..7e064a716aa
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..0fc1d6e99d7
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/kvasir-seg_512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/kvasir-seg_512x512.py
new file mode 100644
index 00000000000..e8b2467f8cf
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/kvasir-seg_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'KvasirSEGDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/datasets/kvasir-seg_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg/datasets/kvasir-seg_dataset.py
new file mode 100644
index 00000000000..9d601328eb6
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/datasets/kvasir-seg_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class KvasirSEGDataset(BaseSegDataset):
+ """KvasirSEGDataset dataset.
+
+ In segmentation map annotation for KvasirSEGDataset, 0 stands for
+ background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix`` is
+ fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'polyp'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/tools/prepare_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg/tools/prepare_dataset.py
new file mode 100644
index 00000000000..74c43e96351
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/tools/prepare_dataset.py
@@ -0,0 +1,87 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '.jpg'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ os.path.join(root_path, 'sessile-main-Kvasir-SEG/images'),
+ tgt_img_dir,
+ suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ os.path.join(root_path, 'sessile-main-Kvasir-SEG/masks'),
+ tgt_mask_dir,
+ suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/README.md b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/README.md
new file mode 100644
index 00000000000..80eb00f51bd
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/README.md
@@ -0,0 +1,145 @@
+# Kvasir-SEG Segmented Polyp Dataset from Aliyun (Kvasir SEG Aliyun)
+
+## Description
+
+This project supports **`Kvasir-SEG Segmented Polyp Dataset from Aliyun (Kvasir SEG Aliyun) `**, which can be downloaded from [here](https://tianchi.aliyun.com/dataset/84385).
+
+### Dataset Overview
+
+Colorectal cancer is the second most common cancer type among women and third most common among men. Polyps are precursors to colorectal cancer and therefore important to detect and remove at an early stage. Polyps are found in nearly half of the individuals at age 50 that undergo a colonoscopy screening, and their frequency increase with age.Polyps are abnormal tissue growth from the mucous membrane, which is lining the inside of the GI tract, and can sometimes be cancerous. Colonoscopy is the gold standard for detection and assessment of these polyps with subsequent biopsy and removal of the polyps. Early disease detection has a huge impact on survival from colorectal cancer. Increasing the detection of polyps has been shown to decrease risk of colorectal cancer. Thus, automatic detection of more polyps at an early stage can play a crucial role in prevention and survival from colorectal cancer.
+
+The Kvasir-SEG dataset is based on the previous Kvasir dataset, which is the first multi-class dataset for gastrointestinal (GI) tract disease detection and classification. It contains annotated polyp images and their corresponding masks. The pixels depicting polyp tissue, the ROI, are represented by the foreground (white mask), while the background (in black) does not contain positive pixels. These images were collected and verified by experienced gastroenterologists from Vestre Viken Health Trust in Norway. The classes include anatomical landmarks, pathological findings and endoscopic procedures.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------ | ----------------- | ------------ | --------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------- |
+| [kvasir-seg](https://tianchi.aliyun.com/dataset/84385) | abdomen | segmentation | endoscopy | 2 | 1000/-/- | yes/-/- | 2020 | [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 1000 | 84.72 | - | - | - | - |
+| polyp | 1000 | 15.28 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{jha2020kvasir,
+ title={Kvasir-seg: A segmented polyp dataset},
+ author={Jha, Debesh and Smedsrud, Pia H and Riegler, Michael A and Halvorsen, P{\aa}l and Lange, Thomas de and Johansen, Dag and Johansen, H{\aa}vard D},
+ booktitle={International Conference on Multimedia Modeling},
+ pages={451--462},
+ year={2020},
+ organization={Springer}
+ }
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `kvasir_seg_aliyun/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://tianchi.aliyun.com/dataset/84385) and decompression data to path 'data/.'.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── endoscopy
+ │ │ │ │ ├── kvasir_seg_aliyun
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 800 | 84.66 | 200 | 84.94 | - | - |
+| polyp | 800 | 15.34 | 200 | 15.06 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..b59db95232b
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..6c526680cd4
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..a192a5bd240
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..5325e1f0807
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/kvasir-seg-aliyun_512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/kvasir-seg-aliyun_512x512.py
new file mode 100644
index 00000000000..5f868804679
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/kvasir-seg-aliyun_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'KvasirSEGAliyunDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/datasets/kvasir-seg-aliyun_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/datasets/kvasir-seg-aliyun_dataset.py
new file mode 100644
index 00000000000..198caf07bcd
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/datasets/kvasir-seg-aliyun_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class KvasirSEGAliyunDataset(BaseSegDataset):
+ """KvasirSEGAliyunDataset dataset.
+
+ In segmentation map annotation for KvasirSEGAliyunDataset,
+ 0 stands for background,which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'polyp'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/tools/prepare_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/tools/prepare_dataset.py
new file mode 100644
index 00000000000..b230e7fef58
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/tools/prepare_dataset.py
@@ -0,0 +1,86 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '.jpg'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path).convert('L'))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+ convert_pics_into_pngs(
+ os.path.join(root_path, 'Kvasir-SEG/images'),
+ tgt_img_dir,
+ suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ os.path.join(root_path, 'Kvasir-SEG/masks'),
+ tgt_mask_dir,
+ suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/README.md b/projects/medical/2d_image/fluorescein_angriogram/vampire/README.md
new file mode 100644
index 00000000000..c2c61c46a0b
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/README.md
@@ -0,0 +1,158 @@
+# Vessel Assessment and Measurement Platform for Images of the REtina
+
+## Description
+
+This project support **`Vessel Assessment and Measurement Platform for Images of the REtina`**, and the dataset used in this project can be downloaded from [here](https://vampire.computing.dundee.ac.uk/vesselseg.html).
+
+### Dataset Overview
+
+In order to promote evaluation of vessel segmentation on ultra-wide field-of-view (UWFV) fluorescein angriogram (FA) frames, we make public 8 frames from two different sequences, the manually annotated images and the result of our automatic vessel segmentation algorithm.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------- | ----------------- | ------------ | ---------------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Vampire](https://vampire.computing.dundee.ac.uk/vesselseg.html) | vessel | segmentation | fluorescein angriogram | 2 | 8/-/- | yes/-/- | 2017 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 8 | 96.75 | - | - | - | - |
+| vessel | 8 | 3.25 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```bibtex
+
+@inproceedings{perez2011improving,
+ title={Improving vessel segmentation in ultra-wide field-of-view retinal fluorescein angiograms},
+ author={Perez-Rovira, Adria and Zutis, K and Hubschman, Jean Pierre and Trucco, Emanuele},
+ booktitle={2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society},
+ pages={2614--2617},
+ year={2011},
+ organization={IEEE}
+}
+
+@article{perez2011rerbee,
+ title={RERBEE: robust efficient registration via bifurcations and elongated elements applied to retinal fluorescein angiogram sequences},
+ author={Perez-Rovira, Adria and Cabido, Raul and Trucco, Emanuele and McKenna, Stephen J and Hubschman, Jean Pierre},
+ journal={IEEE Transactions on Medical Imaging},
+ volume={31},
+ number={1},
+ pages={140--150},
+ year={2011},
+ publisher={IEEE}
+}
+
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `vampire/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://vampire.computing.dundee.ac.uk/vesselseg.html) and decompression data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `python ../../tools/split_seg_dataset.py` to split dataset. For the Bacteria_detection dataset, as there is no test or validation dataset, we sample 20% samples from the whole dataset as the validation dataset and 80% samples for training data and make two filename lists `train.txt` and `val.txt`. As we set the random seed as the hard code, we eliminated the randomness, the dataset split actually can be reproducible.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fluorescein_angriogram
+ │ │ │ │ ├── vampire
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 6 | 97.48 | 2 | 94.54 | - | - |
+| vessel | 6 | 2.52 | 2 | 5.46 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_vampire-512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_vampire-512x512.py
new file mode 100755
index 00000000000..7f5273aaff9
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_vampire-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './vampire_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.vampire_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_vampire-512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_vampire-512x512.py
new file mode 100755
index 00000000000..4382229989b
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_vampire-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './vampire_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.vampire_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=dict(size=img_scale),
+ pretrained=None,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_vampire-512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_vampire-512x512.py
new file mode 100755
index 00000000000..8d93e17627f
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_vampire-512x512.py
@@ -0,0 +1,22 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './vampire_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.vampire_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(
+ num_classes=2,
+ loss_decode=dict(type='CrossEntropyLoss', use_sigmoid=True),
+ out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/vampire_512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/vampire_512x512.py
new file mode 100755
index 00000000000..4eda92f9f22
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/vampire_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'VampireDataset'
+data_root = 'data'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/__init__.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/__init__.py
new file mode 100755
index 00000000000..93f9cbf0506
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/__init__.py
@@ -0,0 +1,3 @@
+from .vampire_dataset import VampireDataset
+
+__all__ = ['VampireDataset']
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/vampire_dataset.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/vampire_dataset.py
new file mode 100755
index 00000000000..4d38040f7f1
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/vampire_dataset.py
@@ -0,0 +1,28 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class VampireDataset(BaseSegDataset):
+ """VampireDataset dataset.
+
+ In segmentation map annotation for VampireDataset, 0 stands for background,
+ which is included in 2 categories. ``reduce_zero_label`` is fixed to
+ False. The ``img_suffix`` is fixed to '.png' and ``seg_map_suffix`` is
+ fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/tools/prepare_dataset.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/tools/prepare_dataset.py
new file mode 100644
index 00000000000..2755b5d28bd
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/tools/prepare_dataset.py
@@ -0,0 +1,44 @@
+import os
+import shutil
+
+from PIL import Image
+
+path = 'data'
+
+if not os.path.exists(os.path.join(path, 'images', 'train')):
+ os.system(f'mkdir -p {os.path.join(path, "images", "train")}')
+
+if not os.path.exists(os.path.join(path, 'masks', 'train')):
+ os.system(f'mkdir -p {os.path.join(path, "masks", "train")}')
+
+origin_data_path = os.path.join(path, 'vesselSegmentation')
+
+imgs_amd14 = os.listdir(os.path.join(origin_data_path, 'AMD14'))
+imgs_ger7 = os.listdir(os.path.join(origin_data_path, 'GER7'))
+
+for img in imgs_amd14:
+ shutil.copy(
+ os.path.join(origin_data_path, 'AMD14', img),
+ os.path.join(path, 'images', 'train', img))
+ # copy GT
+ img_gt = img.replace('.png', '-GT.png')
+ shutil.copy(
+ os.path.join(origin_data_path, 'AMD14-GT', f'{img_gt}'),
+ os.path.join(path, 'masks', 'train', img))
+
+for img in imgs_ger7:
+ shutil.copy(
+ os.path.join(origin_data_path, 'GER7', img),
+ os.path.join(path, 'images', 'train', img))
+ # copy GT
+ img_gt = img.replace('.bmp', '-GT.png')
+ img = img.replace('bmp', 'png')
+ shutil.copy(
+ os.path.join(origin_data_path, 'GER7-GT', img_gt),
+ os.path.join(path, 'masks', 'train', img))
+
+imgs = os.listdir(os.path.join(path, 'images', 'train'))
+for img in imgs:
+ if not img.endswith('.png'):
+ im = Image.open(os.path.join(path, 'images', 'train', img))
+ im.save(os.path.join(path, 'images', 'train', img[:-4] + '.png'))
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/README.md b/projects/medical/2d_image/fundus_photography/dr_hagis/README.md
new file mode 100644
index 00000000000..85d8a3e271c
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/README.md
@@ -0,0 +1,155 @@
+# DR HAGIS: Diabetic Retinopathy, Hypertension, Age-related macular degeneration and Glacuoma ImageS
+
+## Description
+
+This project supports **`DR HAGIS: Diabetic Retinopathy, Hypertension, Age-related macular degeneration and Glacuoma ImageS`**, which can be downloaded from [here](https://paperswithcode.com/dataset/dr-hagis).
+
+### Dataset Overview
+
+The DR HAGIS database has been created to aid the development of vessel extraction algorithms suitable for retinal screening programmes. Researchers are encouraged to test their segmentation algorithms using this database. All thirty-nine fundus images were obtained from a diabetic retinopathy screening programme in the UK. Hence, all images were taken from diabetic patients.
+
+Besides the fundus images, the manual segmentation of the retinal surface vessels is provided by an expert grader. These manually segmented images can be used as the ground truth to compare and assess the automatic vessel extraction algorithms. Masks of the FOV are provided as well to quantify the accuracy of vessel extraction within the FOV only. The images were acquired in different screening centers, therefore reflecting the range of image resolutions, digital cameras and fundus cameras used in the clinic. The fundus images were captured using a Topcon TRC-NW6s, Topcon TRC-NW8 or a Canon CR DGi fundus camera with a horizontal 45 degree field-of-view (FOV). The images are 4752x3168 pixels, 3456x2304 pixels, 3126x2136 pixels, 2896x1944 pixels or 2816x1880 pixels in size.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------- | ----------------- | ------------ | ------------------ | ------------ | --------------------- | ---------------------- | ------------ | ------- |
+| [DR HAGIS](https://paperswithcode.com/dataset/dr-hagis) | head and neck | segmentation | fundus photography | 2 | 40/-/- | yes/-/- | 2017 | - |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 40 | 96.38 | - | - | - | - |
+| vessel | 40 | 3.62 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Usage
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `dr_hagis/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://paperswithcode.com/dataset/dr-hagis) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── dr_hagis
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 32 | 96.21 | 8 | 97.12 | - | - |
+| vessel | 32 | 3.79 | 8 | 2.88 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@article{holm2017dr,
+ title={DR HAGIS—a fundus image database for the automatic extraction of retinal surface vessels from diabetic patients},
+ author={Holm, Sven and Russell, Greg and Nourrit, Vincent and McLoughlin, Niall},
+ journal={Journal of Medical Imaging},
+ volume={4},
+ number={1},
+ pages={014503--014503},
+ year={2017},
+ publisher={Society of Photo-Optical Instrumentation Engineers}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/dr-hagis_512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/dr-hagis_512x512.py
new file mode 100644
index 00000000000..93b96384109
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/dr-hagis_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'DRHAGISDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_dr-hagis-512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_dr-hagis-512x512.py
new file mode 100644
index 00000000000..9d14427c45f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_dr-hagis-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './dr-hagis_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.dr-hagis_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_dr-hagis-512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_dr-hagis-512x512.py
new file mode 100644
index 00000000000..507ec748bf5
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_dr-hagis-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './dr-hagis_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.dr-hagis_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_dr-hagis-512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_dr-hagis-512x512.py
new file mode 100644
index 00000000000..092ae00a7d3
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_dr-hagis-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './dr-hagis_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.dr-hagis_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/datasets/dr-hagis_dataset.py b/projects/medical/2d_image/fundus_photography/dr_hagis/datasets/dr-hagis_dataset.py
new file mode 100644
index 00000000000..9659f0b8d77
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/datasets/dr-hagis_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class DRHAGISDataset(BaseSegDataset):
+ """DRHAGISDataset dataset.
+
+ In segmentation map annotation for DRHAGISDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/dr_hagis/tools/prepare_dataset.py
new file mode 100755
index 00000000000..51f4df7dac2
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/tools/prepare_dataset.py
@@ -0,0 +1,41 @@
+import glob
+import os
+import shutil
+
+import mmengine
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '_manual_orig.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(os.path.join('data/DRHAGIS/**/*' + img_suffix))
+
+mmengine.mkdir_or_exist(root_path + 'images/train/')
+mmengine.mkdir_or_exist(root_path + 'masks/train/')
+
+D3_palette = {0: (0, 0, 0), 1: (1, 1, 1)}
+D3_invert_palette = {v: k for k, v in D3_palette.items()}
+D2_255_convert_dict = {0: 0, 255: 1}
+
+part_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ shutil.copy(
+ img, root_path + 'images/' + part_dir + basename.split('.')[0] +
+ save_img_suffix)
+ mask_path = root_path + 'DRHAGIS/Manual_Segmentations/' + basename.split( # noqa
+ '.')[0] + seg_map_suffix
+ label = np.array(Image.open(mask_path))
+
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix # noqa
+ mask = np.array(Image.open(mask_path)).astype(np.uint8)
+ mask[mask == 255] = 1
+ mask = Image.fromarray(mask)
+ mask.save(save_mask_path)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/README.md b/projects/medical/2d_image/fundus_photography/gamma3/README.md
new file mode 100644
index 00000000000..e834508fcb5
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/README.md
@@ -0,0 +1,167 @@
+# Glaucoma grAding from Multi-Modality imAges Task3
+
+## Description
+
+This project support **`Glaucoma grAding from Multi-Modality imAges Task3`**, and the dataset used in this project can be downloaded from [here](https://aistudio.baidu.com/aistudio/competition/detail/121/0/datasets).
+
+### Dataset Overview
+
+This regular-challenge dataset was provided by Sun Yat-sen Ophthalmic Center, Sun Yat-sen University, Guangzhou, China. The dataset contains 200 fundus color images: 100 pairs in the training set and 100 pairs in the test set.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ----------------------------------------------------------------------------------- | ----------------- | ------------ | --------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [GammaTask3](https://aistudio.baidu.com/aistudio/competition/detail/121/0/datasets) | eye | segmentation | fundus photophy | 3 | 100/-/100 | yes/-/- | 2021 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 100 | 99.02 | - | - | - | - |
+| optic disc | 100 | 0.67 | - | - | - | - |
+| optic cup | 100 | 0.31 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```bibtex
+@article{fu2018joint,
+ title={Joint optic disc and cup segmentation based on multi-label deep network and polar transformation},
+ author={Fu, Huazhu and Cheng, Jun and Xu, Yanwu and Wong, Damon Wing Kee and Liu, Jiang and Cao, Xiaochun},
+ journal={IEEE transactions on medical imaging},
+ volume={37},
+ number={7},
+ pages={1597--1605},
+ year={2018},
+ publisher={IEEE}
+}
+
+@article{sevastopolsky2017optic,
+ title={Optic disc and cup segmentation methods for glaucoma detection with modification of U-Net convolutional neural network},
+ author={Sevastopolsky, Artem},
+ journal={Pattern Recognition and Image Analysis},
+ volume={27},
+ pages={618--624},
+ year={2017},
+ publisher={Springer}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `gammm3/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://aistudio.baidu.com/aistudio/competition/detail/121/0/datasets) and decompression data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── gamma3
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 80 | 99.01 | 20 | 99.07 | - | - |
+| optic disc | 80 | 0.68 | 20 | 0.63 | - | - |
+| optic cup | 80 | 0.32 | 20 | 0.31 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_gamma3-512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_gamma3-512x512.py
new file mode 100644
index 00000000000..0daac51e10f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_gamma3-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './gamma3_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.gamma3_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_gamma3-512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_gamma3-512x512.py
new file mode 100644
index 00000000000..8a25cd0d266
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_gamma3-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './gamma3_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.gamma3_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_gamma3-512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_gamma3-512x512.py
new file mode 100644
index 00000000000..ea648438672
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_gamma3-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './gamma3_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.gamma3_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/gamma3_512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/gamma3_512x512.py
new file mode 100644
index 00000000000..d23ab55ca71
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/gamma3_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'Gamma3Dataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/datasets/gamma3_dataset.py b/projects/medical/2d_image/fundus_photography/gamma3/datasets/gamma3_dataset.py
new file mode 100644
index 00000000000..56cbdd63e61
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/datasets/gamma3_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class Gamma3Dataset(BaseSegDataset):
+ """Gamma3Dataset dataset.
+
+ In segmentation map annotation for Gamma3Dataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'disc', 'cup'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/gamma3/tools/prepare_dataset.py
new file mode 100644
index 00000000000..eb820b6b740
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/tools/prepare_dataset.py
@@ -0,0 +1,107 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**/*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 2,
+ 128: 1,
+ 255: 0
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ os.path.join(
+ root_path,
+ 'task3_disc_cup_segmentation/training/fundus color images/'),
+ tgt_img_train_dir,
+ suffix=img_suffix)
+
+ convert_pics_into_pngs(
+ os.path.join(
+ root_path,
+ 'task3_disc_cup_segmentation/testing/fundus color images/'),
+ tgt_img_test_dir,
+ suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ os.path.join(root_path,
+ 'task3_disc_cup_segmentation/training/Disc_Cup_Mask/'),
+ tgt_mask_train_dir,
+ suffix=seg_map_suffix,
+ convert_dict={
+ 0: 2,
+ 128: 1,
+ 255: 0
+ })
+ # original: [0, 128, 255] for ['optic cup', 'optic disc', 'background']
+ # converted: [0, 1, 2] for ['background', 'optic disc', 'optic cup']
diff --git a/projects/medical/2d_image/fundus_photography/orvs/README.md b/projects/medical/2d_image/fundus_photography/orvs/README.md
new file mode 100644
index 00000000000..6f09203ac4d
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/README.md
@@ -0,0 +1,140 @@
+# ORVS (Online Retinal image for Vessel Segmentation (ORVS))
+
+## Description
+
+This project supports **`ORVS (Online Retinal image for Vessel Segmentation (ORVS))`**, which can be downloaded from [here](https://opendatalab.org.cn/ORVS).
+
+### Dataset Overview
+
+The ORVS dataset is a newly established collaboration between the Department of Computer Science and the Department of Vision Science at the University of Calgary. The dataset contains 49 images collected from a clinic in Calgary, Canada, consisting of 42 training images and 7 testing images. All images were obtained using a Zeiss Visucam 200 with a 30-degree field of view (FOV). The image size is 1444×1444 pixels with 24 bits per pixel. The images are stored in JPEG format with low compression, which is common in ophthalmic practice. All images were manually traced by an expert who has been working in the field of retinal image analysis and has been trained to mark all pixels belonging to retinal vessels. The Windows Paint 3D tool was used for manual image annotation.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------ | ----------------- | ------------ | ------------------ | ------------ | --------------------- | ---------------------- | ------------ | ------- |
+| [Bactteria detection](https://opendatalab.org.cn/ORVS) | bacteria | segmentation | fundus photography | 2 | 130/-/72 | yes/-/yes | 2020 | - |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 130 | 94.83 | - | - | 72 | 94.25 |
+| vessel | 130 | 5.17 | - | - | 72 | 5.75 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `orvs/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- Clone this [repository](https://github.com/AbdullahSarhan/ICPRVessels), then move `Vessels-Datasets` to `data/`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── orvs
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@inproceedings{sarhan2021transfer,
+ title={Transfer learning through weighted loss function and group normalization for vessel segmentation from retinal images},
+ author={Sarhan, Abdullah and Rokne, Jon and Alhajj, Reda and Crichton, Andrew},
+ booktitle={2020 25th International Conference on Pattern Recognition (ICPR)},
+ pages={9211--9218},
+ year={2021},
+ organization={IEEE}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_orvs-512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_orvs-512x512.py
new file mode 100644
index 00000000000..662f837158a
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_orvs-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './orvs_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.orvs_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_orvs-512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_orvs-512x512.py
new file mode 100644
index 00000000000..c47cdb6b24d
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_orvs-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './orvs_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.orvs_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_orvs-512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_orvs-512x512.py
new file mode 100644
index 00000000000..1097aade286
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_orvs-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './orvs_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.orvs_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/orvs_512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/orvs_512x512.py
new file mode 100644
index 00000000000..a5594dec388
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/orvs_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ORVSDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='test.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/orvs/datasets/orvs_dataset.py b/projects/medical/2d_image/fundus_photography/orvs/datasets/orvs_dataset.py
new file mode 100644
index 00000000000..e915ae4cd2b
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/datasets/orvs_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ORVSDataset(BaseSegDataset):
+ """ORVSDataset dataset.
+
+ In segmentation map annotation for ORVSDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/orvs/tools/prepare_dataset.py
new file mode 100755
index 00000000000..f902d871010
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/tools/prepare_dataset.py
@@ -0,0 +1,55 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix_list = ['.jpg', '.png', '.tif']
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(
+ os.path.join('data/Vessels-Datasets/*/Train/Original/Images/*' +
+ img_suffix))
+x_test = glob.glob(
+ os.path.join('data/Vessels-Datasets/*/Test/Original/Images/*' +
+ img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/test/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+os.system('mkdir -p ' + root_path + 'masks/test/')
+
+part_dir_dict = {0: 'train/', 1: 'test/'}
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ type_name = img.split('/')[-5]
+ basename = type_name + '_' + os.path.basename(img)
+ save_img_path = root_path + 'images/' + part_dir + basename.split(
+ '.')[0] + save_img_suffix
+ Image.open(img).save(save_img_path)
+
+ for seg_map_suffix in seg_map_suffix_list:
+ if os.path.exists('/'.join(img.split('/')[:-1]).replace(
+ 'Images', 'Labels')):
+ mask_path = img.replace('Images', 'Labels').replace(
+ img_suffix, seg_map_suffix)
+ else:
+ mask_path = img.replace('Images', 'labels').replace(
+ img_suffix, seg_map_suffix)
+ if os.path.exists(mask_path):
+ break
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix
+ masks = np.array(Image.open(mask_path).convert('L')).astype(np.uint8)
+ if len(np.unique(masks)) == 2 and 1 in np.unique(masks):
+ print(np.unique(masks))
+ pass
+ else:
+ masks[masks < 128] = 0
+ masks[masks >= 128] = 1
+ masks = Image.fromarray(masks)
+ masks.save(save_mask_path)
diff --git a/projects/medical/2d_image/fundus_photography/rite/README.md b/projects/medical/2d_image/fundus_photography/rite/README.md
new file mode 100644
index 00000000000..0aea9b00d17
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/README.md
@@ -0,0 +1,135 @@
+# Retinal Images vessel Tree Extraction (RITE)
+
+## Description
+
+This project supports **`Retinal Images vessel Tree Extraction (RITE) `**, which can be downloaded from [here](https://opendatalab.com/RITE).
+
+### Dataset Overview
+
+The RITE (Retinal Images vessel Tree Extraction) is a database that enables comparative studies on segmentation or classification of arteries and veins on retinal fundus images, which is established based on the public available DRIVE database (Digital Retinal Images for Vessel Extraction). RITE contains 40 sets of images, equally separated into a training subset and a test subset, the same as DRIVE. The two subsets are built from the corresponding two subsets in DRIVE. For each set, there is a fundus photograph, a vessel reference standard. The fundus photograph is inherited from DRIVE. For the training set, the vessel reference standard is a modified version of 1st_manual from DRIVE. For the test set, the vessel reference standard is 2nd_manual from DRIVE.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------ | ----------------- | ------------ | ------------------ | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Rite](https://opendatalab.com/RITE) | head_and_neck | segmentation | fundus_photography | 2 | 20/-/20 | yes/-/yes | 2013 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 20 | 91.61 | - | - | 20 | 91.58 |
+| vessel | 20 | 8.39 | - | - | 20 | 8.42 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@InProceedings{10.1007/978-3-642-40763-5_54,
+ author={Hu, Qiao and Abr{\`a}moff, Michael D. and Garvin, Mona K.},
+ title={Automated Separation of Binary Overlapping Trees in Low-Contrast Color Retinal Images},
+ booktitle={Medical Image Computing and Computer-Assisted Intervention -- MICCAI 2013},
+ year={2013},
+ pages={436--443},
+}
+
+
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `rite/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.com/RITE) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── rite
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_rite-512x512.py
new file mode 100644
index 00000000000..27dd4363b16
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_rite-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_rite-512x512.py
new file mode 100644
index 00000000000..48f6f973a1a
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_rite-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_rite-512x512.py
new file mode 100644
index 00000000000..5f5b24ba6a4
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_rite-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_rite-512x512.py
new file mode 100644
index 00000000000..bf66b6f320c
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_rite-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/rite_512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/rite_512x512.py
new file mode 100644
index 00000000000..02f620c665f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/rite_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'RITEDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='test.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/rite/datasets/rite_dataset.py b/projects/medical/2d_image/fundus_photography/rite/datasets/rite_dataset.py
new file mode 100644
index 00000000000..99f688de949
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/datasets/rite_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class RITEDataset(BaseSegDataset):
+ """RITEDataset dataset.
+
+ In segmentation map annotation for RITEDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/rite/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/rite/tools/prepare_dataset.py
new file mode 100644
index 00000000000..ca7e996961f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/tools/prepare_dataset.py
@@ -0,0 +1,98 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.tif'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+src_img_train_dir = os.path.join(root_path, 'AV_groundTruth/training/images/')
+src_img_test_dir = os.path.join(root_path, 'AV_groundTruth/test/images/')
+src_mask_train_dir = os.path.join(root_path, 'AV_groundTruth/training/vessel/')
+src_mask_test_dir = os.path.join(root_path, 'AV_groundTruth/test/vessel/')
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+tgt_mask_test_dir = os.path.join(root_path, 'masks/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+os.system('mkdir -p ' + tgt_mask_test_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ src_img_train_dir, tgt_img_train_dir, suffix=img_suffix)
+
+ convert_pics_into_pngs(
+ src_img_test_dir, tgt_img_test_dir, suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_train_dir, tgt_mask_train_dir, suffix=seg_map_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_test_dir, tgt_mask_test_dir, suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/README.md b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/README.md
new file mode 100644
index 00000000000..97c4a0f0e55
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/README.md
@@ -0,0 +1,123 @@
+# breastCancerCellSegmentation
+
+## Description
+
+This project supports **`breastCancerCellSegmentation`**, which can be downloaded from [here](https://www.heywhale.com/mw/dataset/5e9e9b35ebb37f002c625423).
+
+### Dataset Overview
+
+This dataset, with 58 H&E-stained histopathology images was used for breast cancer cell detection and associated real-world data.
+Conventional histology uses a combination of hematoxylin and eosin stains, commonly referred to as H&E. These images are stained because most cells are inherently transparent with little or no intrinsic pigment.
+Certain special stains selectively bind to specific components and can be used to identify biological structures such as cells.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| -------------------------------------------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [breastCancerCellSegmentation](https://www.heywhale.com/mw/dataset/5e9e9b35ebb37f002c625423) | cell | segmentation | histopathology | 2 | 58/-/- | yes/-/- | 2020 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 58 | 98.37 | - | - | - | - |
+| breastCancerCell | 58 | 1.63 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Usage
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow (PIL) v9.3.0
+- scikit-learn (sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `breastCancerCellSegmentation/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- Download dataset from [here](https://www.heywhale.com/mw/dataset/5e9e9b35ebb37f002c625423) and save it to the `data/` directory .
+- Decompress data to path `data/`. This will create a new folder named `data/breastCancerCellSegmentation/`, which contains the original image data.
+- run script `python tools/prepare_dataset.py` to format data and change folder structure as below.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── breastCancerCellSegmentation
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── breastCancerCellSegmentation
+ | │ │ │ │ │ │ ├── train.txt
+ | │ │ │ │ │ │ ├── val.txt
+ | │ │ │ │ │ │ ├── images
+ | │ │ │ │ │ │ | ├── xxx.tif
+ | │ │ │ │ │ │ ├── masks
+ | │ │ │ │ │ │ | ├── xxx.TIF
+
+```
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/breastCancerCellSegmentation_512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/breastCancerCellSegmentation_512x512.py
new file mode 100644
index 00000000000..1cf0fccf5be
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/breastCancerCellSegmentation_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'breastCancerCellSegmentationDataset'
+data_root = 'data/breastCancerCellSegmentation'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile', imdecode_backend='tifffile'),
+ dict(type='LoadAnnotations', imdecode_backend='tifffile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile', imdecode_backend='tifffile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations', imdecode_backend='tifffile'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images', seg_map_path='masks'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images', seg_map_path='masks'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breastCancerCellSegmentation-512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breastCancerCellSegmentation-512x512.py
new file mode 100644
index 00000000000..55d17089686
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breastCancerCellSegmentation-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './breastCancerCellSegmentation_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breastCancerCellSegmentation_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breastCancerCellSegmentation-512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breastCancerCellSegmentation-512x512.py
new file mode 100644
index 00000000000..cf28aad739f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breastCancerCellSegmentation-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './breastCancerCellSegmentation_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breastCancerCellSegmentation_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breastCancerCellSegmentation-512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breastCancerCellSegmentation-512x512.py
new file mode 100644
index 00000000000..29aaff38941
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breastCancerCellSegmentation-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './breastCancerCellSegmentation_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breastCancerCellSegmentation_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/datasets/breastCancerCellSegmentation_dataset.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/datasets/breastCancerCellSegmentation_dataset.py
new file mode 100644
index 00000000000..eeceb6318c0
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/datasets/breastCancerCellSegmentation_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class breastCancerCellSegmentationDataset(BaseSegDataset):
+ """breastCancerCellSegmentationDataset dataset.
+
+ In segmentation map annotation for breastCancerCellSegmentationDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'breastCancerCell'))
+
+ def __init__(self,
+ img_suffix='_ccd.tif',
+ seg_map_suffix='.TIF',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/tools/prepare_dataset.py
new file mode 100644
index 00000000000..09cc689c862
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/tools/prepare_dataset.py
@@ -0,0 +1,36 @@
+import argparse
+import glob
+import os
+
+from sklearn.model_selection import train_test_split
+
+
+def save_anno(img_list, file_path, suffix):
+ # 只保留文件名,不保留后缀
+ img_list = [x.split('/')[-1][:-len(suffix)] for x in img_list]
+
+ with open(file_path, 'w') as file_:
+ for x in list(img_list):
+ file_.write(x + '\n')
+
+
+if __name__ == '__main__':
+ parser = argparse.ArgumentParser()
+ parser.add_argument(
+ '--data_root', default='data/breastCancerCellSegmentation/')
+ args = parser.parse_args()
+ data_root = args.data_root
+
+ # 1. 划分训练集、验证集
+ # 1.1 获取所有图片路径
+ img_list = glob.glob(os.path.join(data_root, 'images', '*.tif'))
+ img_list.sort()
+ mask_list = glob.glob(os.path.join(data_root, 'masks', '*.TIF'))
+ mask_list.sort()
+ assert len(img_list) == len(mask_list)
+ # 1.2 划分训练集、验证集、测试集
+ train_img_list, val_img_list, train_mask_list, val_mask_list = train_test_split( # noqa
+ img_list, mask_list, test_size=0.2, random_state=42)
+ # 1.3 保存划分结果
+ save_anno(train_img_list, os.path.join(data_root, 'train.txt'), '_ccd.tif')
+ save_anno(val_img_list, os.path.join(data_root, 'val.txt'), '_ccd.tif')
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/README.md b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/README.md
new file mode 100644
index 00000000000..b6f1ca63419
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/README.md
@@ -0,0 +1,147 @@
+# Breast Cancer Cell Segmentation
+
+## Description
+
+This project support **`Breast Cancer Cell Segmentation`**, and the dataset used in this project can be downloaded from [here](https://tianchi.aliyun.com/dataset/dataDetail?dataId=90152).
+
+### Dataset Overview
+
+In this dataset, there are 58 H&E stained histopathology images used in breast cancer cell detection with associated ground truth data available. Routine histology uses the stain combination of hematoxylin and eosin, commonly referred to as H&E. These images are stained since most cells are essentially transparent, with little or no intrinsic pigment. Certain special stains, which bind selectively to particular components, are be used to identify biological structures such as cells. In those images, the challenging problem is cell segmentation for subsequent classification in benign and malignant cells.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------------------------------------------------ |
+| [Breast Cancer Cell Segmentation](https://tianchi.aliyun.com/dataset/dataDetail?dataId=90152) | thorax | segmentation | histopathology | 2 | 58/-/- | yes/-/- | 2021 | [CC-BY-SA-NC 4.0](http://creativecommons.org/licenses/by-sa/4.0/?spm=5176.12282016.0.0.3f5b5291ypBxb2) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 58 | 98.37 | - | - | - | - |
+| breast cancer cell | 58 | 1.63 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```
+@inproceedings{gelasca2008evaluation,
+ title={Evaluation and benchmark for biological image segmentation},
+ author={Gelasca, Elisa Drelie and Byun, Jiyun and Obara, Boguslaw and Manjunath, BS},
+ booktitle={2008 15th IEEE international conference on image processing},
+ pages={1816--1819},
+ year={2008},
+ organization={IEEE}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `breast_cancer_cell_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://tianchi.aliyun.com/dataset/dataDetail?dataId=90152) and decompression data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── breast_cancer_cell_seg
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 46 | 98.36 | 12 | 98.41 | - | - |
+| erythrocytes | 46 | 1.64 | 12 | 1.59 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/breast-cancer-cell-seg_512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/breast-cancer-cell-seg_512x512.py
new file mode 100644
index 00000000000..ead40e4345c
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/breast-cancer-cell-seg_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'BreastCancerCellSegDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breast-cancer-cell-seg-512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breast-cancer-cell-seg-512x512.py
new file mode 100644
index 00000000000..691a0ff613d
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breast-cancer-cell-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './breast-cancer-cell-seg_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breast-cancer-cell-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breast-cancer-cell-seg-512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breast-cancer-cell-seg-512x512.py
new file mode 100644
index 00000000000..719b767ab1f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breast-cancer-cell-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './breast-cancer-cell-seg_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breast-cancer-cell-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breast-cancer-cell-seg-512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breast-cancer-cell-seg-512x512.py
new file mode 100644
index 00000000000..9dfe70f761f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breast-cancer-cell-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './breast-cancer-cell-seg_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breast-cancer-cell-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/datasets/breast-cancer-cell-seg_dataset.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/datasets/breast-cancer-cell-seg_dataset.py
new file mode 100644
index 00000000000..6f27029d396
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/datasets/breast-cancer-cell-seg_dataset.py
@@ -0,0 +1,29 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class BreastCancerCellSegDataset(BaseSegDataset):
+ """BreastCancerCellSegDataset dataset.
+
+ In segmentation map annotation for BreastCancerCellSegDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('normal', 'breast cancer cell'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/tools/prepare_dataset.py
new file mode 100755
index 00000000000..775f2eed18a
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/tools/prepare_dataset.py
@@ -0,0 +1,47 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.tif'
+seg_map_suffix = '.TIF'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(
+ os.path.join('data/Breast Cancer Cell Segmentation_datasets/Images/*' +
+ img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+D2_255_convert_dict = {0: 0, 255: 1}
+
+
+def convert_2d(img, convert_dict=D2_255_convert_dict):
+ arr_2d = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr_2d[img == c] = i
+ return arr_2d
+
+
+part_dir_dict = {0: 'train/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_save_path = root_path + 'images/' + part_dir + basename.split(
+ '.')[0] + save_img_suffix
+ Image.open(img).save(img_save_path)
+ mask_path = root_path + 'Breast Cancer Cell Segmentation_datasets/Masks/' + '_'.join( # noqa
+ basename.split('_')[:-1]) + seg_map_suffix
+ label = np.array(Image.open(mask_path))
+
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix
+ assert len(label.shape) == 2 and 255 in label and 1 not in label
+ mask = convert_2d(label)
+ mask = Image.fromarray(mask.astype(np.uint8))
+ mask.save(save_mask_path)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/README.md b/projects/medical/2d_image/histopathology/conic2022_seg/README.md
new file mode 100644
index 00000000000..1f55b44ed6d
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/README.md
@@ -0,0 +1,207 @@
+# CoNIC: Colon Nuclei Identification and Counting Challenge
+
+## Description
+
+This project supports **`CoNIC: Colon Nuclei Identification and Counting Challenge`**, which can be downloaded from [here](https://drive.google.com/drive/folders/1il9jG7uA4-ebQ_lNmXbbF2eOK9uNwheb).
+
+### Dataset Overview
+
+Nuclear segmentation, classification and quantification within Haematoxylin & Eosin stained histology images enables the extraction of interpretable cell-based features that can be used in downstream explainable models in computational pathology (CPath). To help drive forward research and innovation for automatic nuclei recognition in CPath, we organise the Colon Nuclei Identification and Counting (CoNIC) Challenge. The challenge requires researchers to develop algorithms that perform segmentation, classification and counting of 6 different types of nuclei within the current largest known publicly available nuclei-level dataset in CPath, containing around half a million labelled nuclei.
+
+### Task Information
+
+The CONIC challenge has 2 tasks:
+
+- Task 1: Nuclear segmentation and classification.
+
+The first task requires participants to segment nuclei within the tissue, while also classifying each nucleus into one of the following categories: epithelial, lymphocyte, plasma, eosinophil, neutrophil or connective tissue.
+
+- Task 2: Prediction of cellular composition.
+
+For the second task, we ask participants to predict how many nuclei of each class are present in each input image.
+
+The output of Task 1 can be directly used to perform Task 2, but these can be treated as independent tasks. Therefore, if it is preferred, prediction of cellular composition can be treated as a stand alone regression task.
+
+***NOTE:We only consider `Task 1` in the following sections.***
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| -------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------------------------------------------------------ |
+| [CoNIC202](https://conic-challenge.grand-challenge.org/) | abdomen | segmentation | histopathology | 7 | 4981/-/- | yes/-/- | 2022 | [Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 4981 | 83.97 | - | - | - | - |
+| neutrophil | 1218 | 0.13 | - | - | - | - |
+| epithelial | 4256 | 10.31 | - | - | - | - |
+| lymphocyte | 4473 | 1.85 | - | - | - | - |
+| plasma | 3316 | 0.55 | - | - | - | - |
+| eosinophil | 1456 | 0.1 | - | - | - | - |
+| connective | 4613 | 3.08 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `conic2022_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://drive.google.com/drive/folders/1il9jG7uA4-ebQ_lNmXbbF2eOK9uNwheb/) and move data to path `'data/CoNIC_Challenge'`. The directory should be like:
+ ```shell
+ data/CoNIC_Challenge
+ ├── README.txt
+ ├── by-nc-sa.md
+ ├── counts.csv
+ ├── images.npy
+ ├── labels.npy
+ └── patch_info.csv
+ ```
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── conic2022_seg
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 3984 | 84.06 | 997 | 83.65 | - | - |
+| neutrophil | 956 | 0.12 | 262 | 0.13 | - | - |
+| epithelial | 3400 | 10.26 | 856 | 10.52 | - | - |
+| lymphocyte | 3567 | 1.83 | 906 | 1.96 | - | - |
+| plasma | 2645 | 0.55 | 671 | 0.56 | - | - |
+| eosinophil | 1154 | 0.1 | 302 | 0.1 | - | - |
+| connective | 3680 | 3.08 | 933 | 3.08 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Organizers
+
+- Simon Graham (TIA, PathLAKE)
+- Mostafa Jahanifar (TIA, PathLAKE)
+- Dang Vu (TIA)
+- Giorgos Hadjigeorghiou (TIA, PathLAKE)
+- Thomas Leech (TIA, PathLAKE)
+- David Snead (UHCW, PathLAKE)
+- Shan Raza (TIA, PathLAKE)
+- Fayyaz Minhas (TIA, PathLAKE)
+- Nasir Rajpoot (TIA, PathLAKE)
+
+TIA: Tissue Image Analytics Centre, Department of Computer Science, University of Warwick, United Kingdom
+
+UHCW: Department of Pathology, University Hospitals Coventry and Warwickshire, United Kingdom
+
+PathLAKE: Pathology Image Data Lake for Analytics Knowledge & Education, , University Hospitals Coventry and Warwickshire, United Kingdom
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@inproceedings{graham2021lizard,
+ title={Lizard: A large-scale dataset for colonic nuclear instance segmentation and classification},
+ author={Graham, Simon and Jahanifar, Mostafa and Azam, Ayesha and Nimir, Mohammed and Tsang, Yee-Wah and Dodd, Katherine and Hero, Emily and Sahota, Harvir and Tank, Atisha and Benes, Ksenija and others},
+ booktitle={Proceedings of the IEEE/CVF International Conference on Computer Vision},
+ pages={684--693},
+ year={2021}
+}
+@article{graham2021conic,
+ title={Conic: Colon nuclei identification and counting challenge 2022},
+ author={Graham, Simon and Jahanifar, Mostafa and Vu, Quoc Dang and Hadjigeorghiou, Giorgos and Leech, Thomas and Snead, David and Raza, Shan E Ahmed and Minhas, Fayyaz and Rajpoot, Nasir},
+ journal={arXiv preprint arXiv:2111.14485},
+ year={2021}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/conic2022-seg_512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/conic2022-seg_512x512.py
new file mode 100644
index 00000000000..51b4e5782ab
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/conic2022-seg_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'Conic2022SegDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_conic2022-512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_conic2022-512x512.py
new file mode 100644
index 00000000000..3e0248c78ce
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_conic2022-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './conic2022-seg_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.conic2022-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=7),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_conic2022-512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_conic2022-512x512.py
new file mode 100644
index 00000000000..fd0e9d8d28b
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_conic2022-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './conic2022-seg_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.conic2022-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=7),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_conic2022-512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_conic2022-512x512.py
new file mode 100644
index 00000000000..bb667f14fd4
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_conic2022-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './conic2022-seg_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.conic2022-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=7),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/conic2022_seg_dataset.png b/projects/medical/2d_image/histopathology/conic2022_seg/conic2022_seg_dataset.png
new file mode 100644
index 00000000000..65bb0bbe0a5
Binary files /dev/null and b/projects/medical/2d_image/histopathology/conic2022_seg/conic2022_seg_dataset.png differ
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/datasets/conic2022-seg_dataset.py b/projects/medical/2d_image/histopathology/conic2022_seg/datasets/conic2022-seg_dataset.py
new file mode 100644
index 00000000000..9af0958ab34
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/datasets/conic2022-seg_dataset.py
@@ -0,0 +1,29 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class Conic2022SegDataset(BaseSegDataset):
+ """Conic2022SegDataset dataset.
+
+ In segmentation map annotation for Conic2022SegDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(
+ classes=('background', 'neutrophil', 'epithelial', 'lymphocyte',
+ 'plasma', 'eosinophil', 'connective'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/conic2022_seg/tools/prepare_dataset.py
new file mode 100755
index 00000000000..89cfb4aae24
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/tools/prepare_dataset.py
@@ -0,0 +1,65 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+
+img_save_root = 'data/'
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+label_set = set()
+
+
+def save_masks_from_npz(data, save_root, part='masks/'):
+ global label_set
+ num = data.shape[0]
+ for i in range(num):
+ # np_img = data[i, :, :, :]
+ np_mask = data[i, :, :, 1]
+ label_set = set.union(label_set, set(np.unique(np_mask)))
+ img = Image.fromarray(np_mask)
+ save_path = os.path.join(save_root, part, str(i) + save_seg_map_suffix)
+ img.save(save_path)
+
+
+def save_images_from_npz(data, save_root, part='images/'):
+ num = data.shape[0]
+ for i in range(num):
+ np_img = data[i, :, :, :]
+ img = Image.fromarray(np_img)
+ save_path = os.path.join(save_root, part, str(i) + save_img_suffix)
+ img.save(save_path)
+
+
+images_npy = np.load('data/CoNIC_Challenge/images.npy')
+labels_npy = np.load('data/CoNIC_Challenge/labels.npy')
+
+os.system('mkdir -p ' + img_save_root + 'images_ori')
+os.system('mkdir -p ' + img_save_root + 'labels')
+save_images_from_npz(images_npy, img_save_root, 'images_ori')
+save_masks_from_npz(labels_npy, img_save_root, 'labels')
+print(label_set)
+
+x_train = glob.glob(os.path.join('data/images_ori/*' + img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+part_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ shutil.copy(
+ img, root_path + 'images/' + part_dir + basename.split('.')[0] +
+ save_img_suffix)
+ mask_path = root_path + 'labels/' + basename.split(
+ '.')[0] + seg_map_suffix
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix
+ shutil.copy(mask_path, save_mask_path)
diff --git a/projects/medical/2d_image/histopathology/consep/README.md b/projects/medical/2d_image/histopathology/consep/README.md
new file mode 100644
index 00000000000..ca3d7aa1089
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/README.md
@@ -0,0 +1,147 @@
+# Colorectal Nuclear Segmentation and Phenotypes (CoNSeP) Dataset
+
+## Description
+
+This project supports **`Colorectal Nuclear Segmentation and Phenotypes (CoNSeP) Dataset`**, which can be downloaded from [here](https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/).
+
+### Dataset Overview
+
+The CoNSeP (Colon Segmentation and Phenotyping) dataset consists of 41 H&E stained image tiles, each with a size of 1,000×1,000 pixels and a magnification of 40x. These images were extracted from 16 colorectal adenocarcinoma (CRA) whole slide images (WSI), each of which belonged to a separate patient and was scanned using an Omnyx VL120 scanner at the Pathology Department of the University Hospitals Coventry and Warwickshire NHS Trust, UK. This dataset was first used in paper named, "HoVer-Net: Simultaneous Segmentation and Classification of Nuclei in Multi-Tissue Histology Images".
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| -------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------- |
+| [CoNIC202](https://conic-challenge.grand-challenge.org/) | abdomen | segmentation | histopathology | 7 | 4981/-/- | yes/-/- | 2022 | - |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :-----------------------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 27 | 83.61 | 14 | 80.4 | - | - |
+| other | 17 | 0.17 | 9 | 0.52 | - | - |
+| inflammatory | 25 | 2.66 | 14 | 2.14 | - | - |
+| healthy epithelial | 3 | 1.47 | 2 | 1.58 | - | - |
+| dysplastic/malignant epithelial | 10 | 7.17 | 8 | 9.16 | - | - |
+| fibroblast | 23 | 3.84 | 14 | 4.63 | - | - |
+| muscle | 8 | 1.05 | 3 | 1.42 | - | - |
+| endothelial | 7 | 0.02 | 4 | 0.15 | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `conic2022_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://opendatalab.com/CoNSeP) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── consep
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@article{graham2019hover,
+ title={Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images},
+ author={Graham, Simon and Vu, Quoc Dang and Raza, Shan E Ahmed and Azam, Ayesha and Tsang, Yee Wah and Kwak, Jin Tae and Rajpoot, Nasir},
+ journal={Medical Image Analysis},
+ volume={58},
+ pages={101563},
+ year={2019},
+ publisher={Elsevier}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/consep/configs/consep_512x512.py b/projects/medical/2d_image/histopathology/consep/configs/consep_512x512.py
new file mode 100644
index 00000000000..0d9b8948b01
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/consep_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ConsepDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_consep-512x512.py b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_consep-512x512.py
new file mode 100644
index 00000000000..cbcf5db775b
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_consep-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './consep_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.consep_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=8),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_consep-512x512.py b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_consep-512x512.py
new file mode 100644
index 00000000000..b374566e6ef
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_consep-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './consep_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.consep_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=8),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_consep-512x512.py b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_consep-512x512.py
new file mode 100644
index 00000000000..35bdaa34c84
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_consep-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './consep_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.consep_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=8),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/consep/datasets/consep_dataset.py b/projects/medical/2d_image/histopathology/consep/datasets/consep_dataset.py
new file mode 100644
index 00000000000..ceb2b3ab25b
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/datasets/consep_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ConsepDataset(BaseSegDataset):
+ """ConsepDataset dataset.
+
+ In segmentation map annotation for ConsepDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(
+ classes=('background', 'other', 'inflammatory', 'healthy epithelial',
+ 'dysplastic/malignant epithelial', 'fibroblast', 'muscle',
+ 'endothelial'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/consep/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/consep/tools/prepare_dataset.py
new file mode 100755
index 00000000000..83a2e18ce10
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/tools/prepare_dataset.py
@@ -0,0 +1,54 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+from scipy.io import loadmat
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.mat'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(os.path.join('data/CoNSeP/Train/Images/*' + img_suffix))
+x_test = glob.glob(os.path.join('data/CoNSeP/Test/Images/*' + img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/val/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+os.system('mkdir -p ' + root_path + 'masks/val/')
+D2_255_convert_dict = {0: 0, 255: 1}
+
+
+def convert_2d(img, convert_dict=D2_255_convert_dict):
+ arr_2d = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr_2d[img == c] = i
+ return arr_2d
+
+
+part_dir_dict = {0: 'CoNSeP/Train/', 1: 'CoNSeP/Test/'}
+save_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ shutil.copy(
+ img, root_path + 'images/' + save_dir_dict[ith] +
+ basename.split('.')[0] + save_img_suffix)
+
+ mask_path = root_path + part_dir + 'Labels/' + basename.split(
+ '.')[0] + seg_map_suffix
+ label_ = loadmat(mask_path)
+ label = label_['inst_map']
+ label_type = label_['inst_type']
+ label_dict = {i + 1: int(val) for i, val in enumerate(label_type)}
+
+ save_mask_path = root_path + 'masks/' + save_dir_dict[
+ ith] + basename.split('.')[0] + save_seg_map_suffix
+
+ res = convert_2d(label, convert_dict=label_dict)
+ res = Image.fromarray(res.astype(np.uint8))
+ res.save(save_mask_path)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/README.md b/projects/medical/2d_image/histopathology/fusc2021/README.md
new file mode 100644
index 00000000000..8130d593503
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/README.md
@@ -0,0 +1,136 @@
+# Foot Ulcer Segmentation Challenge 2021 (FUSC 2021)
+
+## Description
+
+This project supports **`Foot Ulcer Segmentation Challenge 2021 (FUSC 2021) `**, which can be downloaded from [here](https://fusc.grand-challenge.org/).
+
+### Dataset Overview
+
+This chronic wound dataset was collected over 2 years from October 2019 to April 2021 at the center and contains 1,210 foot ulcer images taken from 889 patients during multiple clinical visits. The raw images were taken by Canon SX 620 HS digital camera and iPad Pro under uncontrolled illumination conditions,
+with various backgrounds. The images (shown in Figure 1) are randomly split into 3 subsets: a training set with 810 images, a validation set with 200 images, and a testing set with 200 images. Of course, the annotations of the testing set are kept private. The data collected were de-identified and in accordance with relevant guidelines and regulations and the patient’s informed consent is waived by the institutional review board of the University of Wisconsin-Milwaukee.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [fusc2021](https://fusc.grand-challenge.org/) | lower limb | segmentation | histopathology | 2 | 810/200/200 | yes/yes/no | 2021 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 810 | 98.71 | 200 | 98.78 | - | - |
+| wound | 791 | 1.29 | 195 | 1.22 | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{s41598-020-78799-w,
+ title={Fully automatic wound segmentation with deep convolutional neural networks},
+ author={Chuanbo Wang and D. M. Anisuzzaman and Victor Williamson and Mrinal Kanti Dhar and Behrouz Rostami and Jeffrey Niezgoda and Sandeep Gopalakrishnan and Zeyun Yu},
+ journal={Scientific Reports},
+ volume={10},
+ number={1},
+ pages={21897},
+ year={2020}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `fusc2021/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://fusc.grand-challenge.org/) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── fusc2021
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..c3f42751124
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_fusc2021-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..ed870303fff
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_fusc2021-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..cbc09ae6cdd
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_fusc2021-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..f1477ee725e
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_fusc2021-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fusc2021_512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fusc2021_512x512.py
new file mode 100644
index 00000000000..e650474cea8
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fusc2021_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'FUSC2021Dataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/fusc2021/datasets/fusc2021_dataset.py b/projects/medical/2d_image/histopathology/fusc2021/datasets/fusc2021_dataset.py
new file mode 100644
index 00000000000..d331ac8c3a2
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/datasets/fusc2021_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class FUSC2021Dataset(BaseSegDataset):
+ """FUSC2021Dataset dataset.
+
+ In segmentation map annotation for FUSC2021Dataset, 0 stands for background
+ , which is included in 2 categories. ``reduce_zero_label``
+ is fixed to False. The ``img_suffix`` is fixed to '.png' and
+ ``seg_map_suffix`` is fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'wound'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/fusc2021/tools/prepare_dataset.py
new file mode 100644
index 00000000000..8f2de3daa95
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/tools/prepare_dataset.py
@@ -0,0 +1,114 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+src_img_train_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/train/images')
+src_img_val_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/validation/images')
+src_img_test_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/test/images')
+src_mask_train_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/train/labels')
+src_mask_val_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/validation/labels')
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_val_dir = os.path.join(root_path, 'images/val/')
+tgt_mask_val_dir = os.path.join(root_path, 'masks/val/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_img_val_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_mask_val_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path).convert('L'))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ src_img_train_dir, tgt_img_train_dir, suffix=img_suffix)
+
+ convert_pics_into_pngs(src_img_val_dir, tgt_img_val_dir, suffix=img_suffix)
+
+ convert_pics_into_pngs(
+ src_img_test_dir, tgt_img_test_dir, suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_train_dir, tgt_mask_train_dir, suffix=seg_map_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_val_dir, tgt_mask_val_dir, suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/histopathology/pannuke/README.md b/projects/medical/2d_image/histopathology/pannuke/README.md
new file mode 100644
index 00000000000..e0cade7536d
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/README.md
@@ -0,0 +1,146 @@
+# Pan-Cancer Histology Dataset for Nuclei Instance Segmentation and Classification (PanNuke)
+
+## Description
+
+This project supports **`Pan-Cancer Histology Dataset for Nuclei Instance Segmentation and Classification (PanNuke)`**, which can be downloaded from [here](https://academictorrents.com/details/99f2c7b57b95500711e33f2ee4d14c9fd7c7366c).
+
+### Dataset Overview
+
+Semi automatically generated nuclei instance segmentation and classification dataset with exhaustive nuclei labels across 19 different tissue types. The dataset consists of 481 visual fields, of which 312 are randomly sampled from more than 20K whole slide images at different magnifications, from multiple data sources. In total the dataset contains 205,343 labeled nuclei, each with an instance segmentation mask. Models trained on pannuke can aid in whole slide image tissue type segmentation, and generalise to new tissues. PanNuke demonstrates one of the first successfully semi-automatically generated datasets.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Pannuke](https://academictorrents.com/details/99f2c7b57b95500711e33f2ee4d14c9fd7c7366c) | full_body | segmentation | histopathology | 6 | 7901/-/- | yes/-/- | 2019 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :-----------------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 7901 | 83.32 | - | - | - | - |
+| neoplastic | 4190 | 8.64 | - | - | - | - |
+| non-neoplastic epithelial | 4126 | 1.77 | - | - | - | - |
+| inflammatory | 6137 | 3.73 | - | - | - | - |
+| connective | 232 | 0.07 | - | - | - | - |
+| dead | 1528 | 2.47 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{gamper2019pannuke,
+ title={PanNuke: an open pan-cancer histology dataset for nuclei instance segmentation and classification},
+ author={Gamper, Jevgenij and Koohbanani, Navid Alemi and Benet, Ksenija and Khuram, Ali and Rajpoot, Nasir},
+ booktitle={European Congress on Digital Pathology},
+ pages={11--19},
+ year={2019},
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `pannuke/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://academictorrents.com/details/99f2c7b57b95500711e33f2ee4d14c9fd7c7366c) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── pannuke
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :-----------------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 6320 | 83.38 | 1581 | 83.1 | - | - |
+| neoplastic | 3339 | 8.55 | 851 | 9.0 | - | - |
+| non-neoplastic epithelial | 3293 | 1.77 | 833 | 1.76 | - | - |
+| inflammatory | 4914 | 3.72 | 1223 | 3.76 | - | - |
+| connective | 170 | 0.06 | 62 | 0.09 | - | - |
+| dead | 1235 | 2.51 | 293 | 2.29 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..92584e9a684
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './bactteria-detection_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pannuke-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pannuke-512x512.py
new file mode 100644
index 00000000000..042a08ce008
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pannuke-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pannuke_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pannuke_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=6),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pannuke-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pannuke-512x512.py
new file mode 100644
index 00000000000..e92514c9132
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pannuke-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pannuke_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pannuke_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=6),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pannuke-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pannuke-512x512.py
new file mode 100644
index 00000000000..a9403c849fa
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pannuke-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pannuke_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pannuke_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=6),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/pannuke_512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/pannuke_512x512.py
new file mode 100644
index 00000000000..316ac1ac443
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/pannuke_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'PanNukeDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/pannuke/datasets/pannuke_dataset.py b/projects/medical/2d_image/histopathology/pannuke/datasets/pannuke_dataset.py
new file mode 100644
index 00000000000..4d3c687ff31
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/datasets/pannuke_dataset.py
@@ -0,0 +1,33 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class PanNukeDataset(BaseSegDataset):
+ """PanNukeDataset dataset.
+
+ In segmentation map annotation for PanNukeDataset,
+ 0 stands for background, which is included in 6 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(
+ classes=('background', 'neoplastic', 'non-neoplastic epithelial',
+ 'inflammatory', 'connective', 'dead'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/pannuke/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/pannuke/tools/prepare_dataset.py
new file mode 100644
index 00000000000..7213b181f40
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/tools/prepare_dataset.py
@@ -0,0 +1,49 @@
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+
+tgt_img_dir = os.path.join(root_path, 'images/train')
+tgt_mask_dir = os.path.join(root_path, 'masks/train')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+fold_img_paths = sorted([
+ os.path.join(root_path, 'pannuke/Fold 1/images/fold1/images.npy'),
+ os.path.join(root_path, 'pannuke/Fold 2/images/fold2/images.npy'),
+ os.path.join(root_path, 'pannuke/Fold 3/images/fold3/images.npy')
+])
+
+fold_mask_paths = sorted([
+ os.path.join(root_path, 'pannuke/Fold 1/masks/fold1/masks.npy'),
+ os.path.join(root_path, 'pannuke/Fold 2/masks/fold2/masks.npy'),
+ os.path.join(root_path, 'pannuke/Fold 3/masks/fold3/masks.npy')
+])
+
+for n, (img_path,
+ mask_path) in enumerate(zip(fold_img_paths, fold_mask_paths)):
+ fold_name = str(n + 1)
+ imgs = np.load(img_path)
+ masks = np.load(mask_path)
+
+ for i in range(imgs.shape[0]):
+ img = np.uint8(imgs[i])
+ mask_multichannel = np.minimum(np.uint8(masks[i]), 1)
+ mask = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
+ for j in range(mask_multichannel.shape[-1]):
+ factor = (j + 1) % mask_multichannel.shape[-1]
+ # convert [0,1,2,3,4,5] to [1,2,3,4,5,0],
+ # with the last label being background
+ mask[mask_multichannel[..., j] == 1] = factor
+
+ file_name = 'fold' + fold_name + '_' + str(i).rjust(4, '0') + '.png'
+ print('Processing: ', file_name)
+ tgt_img_path = os.path.join(tgt_img_dir, file_name)
+ tgt_mask_path = os.path.join(tgt_mask_dir, file_name)
+ Image.fromarray(img).save(tgt_img_path)
+ Image.fromarray(mask).save(tgt_mask_path)
+
+ del imgs
+ del masks
diff --git a/projects/medical/2d_image/histopathology/pcam/README.md b/projects/medical/2d_image/histopathology/pcam/README.md
new file mode 100644
index 00000000000..5a8094950c5
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/README.md
@@ -0,0 +1,153 @@
+# PCam (PatchCamelyon)
+
+## Description
+
+This project supports **`Patch Camelyon (PCam) `**, which can be downloaded from [here](https://opendatalab.com/PCam).
+
+### Dataset Overview
+
+PatchCamelyon is an image classification dataset. It consists of 327680 color images (96 x 96px) extracted from histopathologic scans of lymph node sections. Each image is annotated with a binary label indicating presence of metastatic tissue. PCam provides a new benchmark for machine learning models: bigger than CIFAR10, smaller than ImageNet, trainable on a single GPU.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------ | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [Pcam](https://opendatalab.com/PCam) | throax | segmentation | histopathology | 2 | 327680/-/- | yes/-/- | 2018 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 214849 | 63.77 | - | - | - | - |
+| metastatic tissue | 131832 | 36.22 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{veeling2018rotation,
+ title={Rotation equivariant CNNs for digital pathology},
+ author={Veeling, Bastiaan S and Linmans, Jasper and Winkens, Jim and Cohen, Taco and Welling, Max},
+ booktitle={International Conference on Medical image computing and computer-assisted intervention},
+ pages={210--218},
+ year={2018},
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `pcam/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.com/PCam) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```shell
+mkdir data & cd data
+pip install opendatalab
+odl get PCam
+mv ./PCam/raw/pcamv1 ./
+rm -rf PCam
+cd ..
+python tools/prepare_dataset.py
+python ../../tools/split_seg_dataset.py
+```
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── pcam
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 171948 | 63.82 | 42901 | 63.6 | - | - |
+| metastatic tissue | 105371 | 36.18 | 26461 | 36.4 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pcam-512x512.py
new file mode 100644
index 00000000000..20601f1ea56
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pcam-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pcam-512x512.py
new file mode 100644
index 00000000000..c057535409f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pcam-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pcam-512x512.py
new file mode 100644
index 00000000000..4c1d5fe421c
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pcam-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_pcam-512x512.py
new file mode 100644
index 00000000000..25e3734795c
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_pcam-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/pcam_512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/pcam_512x512.py
new file mode 100644
index 00000000000..04efc23eb51
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/pcam_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'PCamDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/pcam/datasets/pcam_dataset.py b/projects/medical/2d_image/histopathology/pcam/datasets/pcam_dataset.py
new file mode 100644
index 00000000000..1c27de543ab
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/datasets/pcam_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class PCamDataset(BaseSegDataset):
+ """PCamDataset dataset.
+
+ In segmentation map annotation for PCamDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'metastatic tissue'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/pcam/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/pcam/tools/prepare_dataset.py
new file mode 100644
index 00000000000..75038e6fb49
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/tools/prepare_dataset.py
@@ -0,0 +1,49 @@
+import os
+
+import h5py
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_val_dir = os.path.join(root_path, 'images/val/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_val_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def extract_pics_from_h5(h5_path, h5_key, save_dir):
+ f = h5py.File(h5_path, 'r')
+ for i, img in enumerate(f[h5_key]):
+ img = img.astype(np.uint8).squeeze()
+ img = Image.fromarray(img)
+ save_image_path = os.path.join(save_dir, str(i).zfill(8) + '.png')
+ img.save(save_image_path)
+
+
+if __name__ == '__main__':
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_train_x.h5',
+ h5_key='x',
+ save_dir=tgt_img_train_dir)
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_valid_x.h5',
+ h5_key='x',
+ save_dir=tgt_img_val_dir)
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_test_x.h5',
+ h5_key='x',
+ save_dir=tgt_img_test_dir)
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_train_mask.h5',
+ h5_key='mask',
+ save_dir=tgt_mask_train_dir)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/README.md b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/README.md
new file mode 100644
index 00000000000..ca95921ba36
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/README.md
@@ -0,0 +1,167 @@
+# RAVIR: A Dataset and Methodology for the Semantic Segmentation and Quantitative Analysis of Retinal Arteries and Veins in Infrared Reflectance Imaging
+
+## Description
+
+This project support **`RAVIR: A Dataset and Methodology for the Semantic Segmentation and Quantitative Analysis of Retinal Arteries and Veins in Infrared Reflectance Imaging`**, and the dataset used in this project can be downloaded from [here](https://ravir.grand-challenge.org/).
+
+### Dataset Overview
+
+The retinal vasculature provides important clues in the diagnosis and monitoring of systemic diseases including hypertension and diabetes. The microvascular system is of primary involvement in such conditions, and the retina is the only anatomical site where the microvasculature can be directly observed. The objective assessment of retinal vessels has long been considered a surrogate biomarker for systemic vascular diseases, and with recent advancements in retinal imaging and computer vision technologies, this topic has become the subject of renewed attention. In this paper, we present a novel dataset, dubbed RAVIR, for the semantic segmentation of Retinal Arteries and Veins in Infrared Reflectance (IR) imaging. It enables the creation of deep learning-based models that distinguish extracted vessel type without extensive post-processing.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------- | ----------------- | ------------ | ---------------------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Ravir](https://ravir.grand-challenge.org/) | eye | segmentation | infrared reflectance imaging | 3 | 23/-/19 | yes/-/- | 2022 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 23 | 87.22 | - | - | - | - |
+| artery | 23 | 5.45 | - | - | - | - |
+| vein | 23 | 7.33 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```bibtex
+@article{hatamizadeh2022ravir,
+ title={RAVIR: A dataset and methodology for the semantic segmentation and quantitative analysis of retinal arteries and veins in infrared reflectance imaging},
+ author={Hatamizadeh, Ali and Hosseini, Hamid and Patel, Niraj and Choi, Jinseo and Pole, Cameron C and Hoeferlin, Cory M and Schwartz, Steven D and Terzopoulos, Demetri},
+ journal={IEEE Journal of Biomedical and Health Informatics},
+ volume={26},
+ number={7},
+ pages={3272--3283},
+ year={2022},
+ publisher={IEEE}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `ravir/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://ravir.grand-challenge.org/) and decompression data to path `'data/ravir/'`.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py --data_root data/ravir"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── infrared_reflectance_imaging
+ │ │ │ │ ├── ravir
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 18 | 87.41 | 5 | 86.53 | - | - |
+| artery | 18 | 5.44 | 5 | 5.50 | - | - |
+| vein | 18 | 7.15 | 5 | 7.97 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+### Testing commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### Ravir
+
+| Method | Backbone | Crop Size | lr | config |
+| :-------------: | :------: | :-------: | :----: | :------------------------------------------------------------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py
new file mode 100755
index 00000000000..375ad5abf27
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './ravir_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.ravir_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py
new file mode 100755
index 00000000000..a7ecf6dd453
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './ravir_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.ravir_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=dict(size=img_scale),
+ pretrained=None,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py
new file mode 100755
index 00000000000..28556df53d1
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './ravir_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.ravir_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/ravir_512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/ravir_512x512.py
new file mode 100755
index 00000000000..cb4c292d1f7
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/ravir_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'RAVIRDataset'
+data_root = 'data/ravir'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/__init__.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/__init__.py
new file mode 100755
index 00000000000..6f1d051bcff
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/__init__.py
@@ -0,0 +1,3 @@
+from .ravir_dataset import RAVIRDataset
+
+__all__ = ['RAVIRDataset']
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/ravir_dataset.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/ravir_dataset.py
new file mode 100755
index 00000000000..c9e0a8ed21e
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/ravir_dataset.py
@@ -0,0 +1,28 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class RAVIRDataset(BaseSegDataset):
+ """RAVIRDataset dataset.
+
+ In segmentation map annotation for RAVIRDataset, 0 stands for background,
+ which is included in 3 categories. ``reduce_zero_label`` is fixed to
+ False. The ``img_suffix`` is fixed to '.png' and ``seg_map_suffix`` is
+ fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'artery', 'vein'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/tools/prepare_dataset.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/tools/prepare_dataset.py
new file mode 100644
index 00000000000..068dcad8145
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/tools/prepare_dataset.py
@@ -0,0 +1,33 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+from tqdm import tqdm
+
+# map = {255:2, 128:1, 0:0}
+
+os.makedirs('data/ravir/images/train', exist_ok=True)
+os.makedirs('data/ravir/images/test', exist_ok=True)
+os.makedirs('data/ravir/masks/train', exist_ok=True)
+
+os.system(
+ r'cp data/ravir/RAVIR\ Dataset/train/training_images/* data/ravir/images/train' # noqa
+)
+os.system(
+ r'cp data/ravir/RAVIR\ Dataset/train/training_masks/* data/ravir/masks/train' # noqa
+)
+os.system(r'cp data/ravir/RAVIR\ Dataset/test/* data/ravir/images/test')
+
+os.system(r'rm -rf data/ravir/RAVIR\ Dataset')
+
+imgs = glob.glob(os.path.join('data/ravir/masks/train', '*.png'))
+
+for im_path in tqdm(imgs):
+ im = Image.open(im_path)
+ imn = np.array(im)
+ imn[imn == 255] = 2
+ imn[imn == 128] = 1
+ imn[imn == 0] = 0
+ new_im = Image.fromarray(imn)
+ new_im.save(im_path)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/README.md b/projects/medical/2d_image/microscopy_images/2pm_vessel/README.md
new file mode 100644
index 00000000000..1feb433a319
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/README.md
@@ -0,0 +1,153 @@
+# 2-PM Vessel Dataset
+
+## Description
+
+This project supports **`2-PM Vessel Dataset`**, which can be downloaded from [here](https://opendatalab.org.cn/2-PM_Vessel_Dataset).
+
+### Dataset Overview
+
+An open-source volumetric brain vasculature dataset obtained with two-photon microscopy at Focused Ultrasound Lab, at Sunnybrook Research Institute (affiliated with University of Toronto by Dr. Alison Burgess, Charissa Poon and Marc Santos).
+
+The dataset contains a total of 12 volumetric stacks consisting images of mouse brain vasculature and tumor vasculature.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------------ | ----------------- | ------------ | ----------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [2pm_vessel](https://opendatalab.org.cn/2-PM_Vessel_Dataset) | vessel | segmentation | microscopy_images | 2 | 216/-/- | yes/-/- | 2021 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 216 | 85.78 | - | - | - | - |
+| vessel | 180 | 14.22 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{teikari2016deep,
+ title={Deep learning convolutional networks for multiphoton microscopy vasculature segmentation},
+ author={Teikari, Petteri and Santos, Marc and Poon, Charissa and Hynynen, Kullervo},
+ journal={arXiv preprint arXiv:1606.02382},
+ year={2016}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `2pm_vessel/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.org.cn/2-PM_Vessel_Dataset) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```shell
+mkdir data & cd data
+pip install opendatalab
+odl get 2-PM_Vessel_Dataset
+cd ..
+python tools/prepare_dataset.py
+python ../../tools/split_seg_dataset.py
+```
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── microscopy_images
+ │ │ │ │ ├── 2pm_vessel
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 172 | 85.88 | 44 | 85.4 | - | - |
+| vessel | 142 | 14.12 | 38 | 14.6 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/2pm-vessel_512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/2pm-vessel_512x512.py
new file mode 100644
index 00000000000..124403fa973
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/2pm-vessel_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'TwoPMVesselDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_2pm-vessel-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_2pm-vessel-512x512.py
new file mode 100644
index 00000000000..2a429e9068e
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_2pm-vessel-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_2pm-vessel-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_2pm-vessel-512x512.py
new file mode 100644
index 00000000000..10d9bb82f25
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_2pm-vessel-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_2pm-vessel-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_2pm-vessel-512x512.py
new file mode 100644
index 00000000000..65c1579ec71
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_2pm-vessel-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..91ed6ada3f2
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/datasets/2pm-vessel_dataset.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/datasets/2pm-vessel_dataset.py
new file mode 100644
index 00000000000..984b5a1361d
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/datasets/2pm-vessel_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class TwoPMVesselDataset(BaseSegDataset):
+ """TwoPMVesselDataset dataset.
+
+ In segmentation map annotation for TwoPMVesselDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/tools/prepare_dataset.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/tools/prepare_dataset.py
new file mode 100644
index 00000000000..1b46af2cad0
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/tools/prepare_dataset.py
@@ -0,0 +1,46 @@
+import os
+
+import tifffile as tiff
+from PIL import Image
+
+root_path = 'data/'
+
+image_dir = os.path.join(root_path,
+ '2-PM_Vessel_Dataset/raw/vesselNN_dataset/denoised')
+label_dir = os.path.join(root_path,
+ '2-PM_Vessel_Dataset/raw/vesselNN_dataset/labels')
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+
+
+def filter_suffix(src_dir, suffix):
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_names = [_ for _ in os.listdir(src_dir) if _.endswith(suffix)]
+ file_paths = [os.path.join(src_dir, _) for _ in file_names]
+ return sorted(file_paths), sorted(file_names)
+
+
+if __name__ == '__main__':
+
+ image_path_list, _ = filter_suffix(image_dir, suffix='tif')
+ label_path_list, _ = filter_suffix(label_dir, suffix='.tif')
+
+ for img_path, label_path in zip(image_path_list, label_path_list):
+ labels = tiff.imread(label_path)
+ images = tiff.imread(img_path)
+ assert labels.ndim == 3
+ assert images.shape == labels.shape
+ name = img_path.split('/')[-1].replace('.tif', '')
+ # a single .tif file contains multiple slices
+ # as long as it is read by tifffile package.
+ for i in range(labels.shape[0]):
+ slice_name = name + '_' + str(i).rjust(3, '0') + '.png'
+ image = images[i]
+ label = labels[i] // 255
+
+ save_path_label = os.path.join(tgt_mask_train_dir, slice_name)
+ Image.fromarray(label).save(save_path_label)
+ save_path_image = os.path.join(tgt_img_train_dir, slice_name)
+ Image.fromarray(image).convert('RGB').save(save_path_image)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/README.md b/projects/medical/2d_image/microscopy_images/bactteria_detection/README.md
new file mode 100644
index 00000000000..1cedda715a7
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/README.md
@@ -0,0 +1,160 @@
+# Bactteria detection with darkfield microscopy
+
+## Description
+
+This project supports **`Bactteria detection with darkfield microscopy`**, which can be downloaded from [here](https://tianchi.aliyun.com/dataset/94411).
+
+### Dataset Overview
+
+Spirochaeta is a genus of bacteria classified within the phylum Spirochaetes. Included in this dataset are 366 darkfield microscopy images and manually annotated masks which can be used for classification and segmentation purposes. Detecting bacteria in blood could have a huge significance for research in both the medical and computer science field.
+
+It was gathered and annotated by students (hand-on experience)
+It has more than just one targeted class (blood cell and bacteria were annotated)
+It is highly imbalanced, so naive loss functions would work less properly
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------------------------- | ----------------- | ------------ | ---------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Bactteria detection](https://tianchi.aliyun.com/dataset/94411) | bacteria | segmentation | microscopy | 3 | 366/-/- | yes/-/- | 2017 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 366 | 85.9 | - | - | - | - |
+| erythrocytes | 345 | 13.03 | - | - | - | - |
+| spirochaete | 288 | 1.07 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Usage
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow (PIL) v9.3.0
+- scikit-learn (sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `bactteria_detection/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- Download dataset from [here](https://tianchi.aliyun.com/dataset/94411) and save it to the `data/` directory .
+- Decompress data to path `data/`. This will create a new folder named `data/Bacteria_detection_with_darkfield_microscopy_datasets/`, which contains the original image data.
+- run script `python tools/prepare_dataset.py` to format data and change folder structure as below.
+- run script `python ../../tools/split_seg_dataset.py` to split dataset. For the Bacteria_detection dataset, as there is no test or validation dataset, we sample 20% samples from the whole dataset as the validation dataset and 80% samples for training data and make two filename lists `train.txt` and `val.txt`. As we set the random seed as the hard code, we eliminated the randomness, the dataset split actually can be reproducible.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── microscopy_images
+ │ │ │ │ ├── bactteria_detection
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── Bacteria_detection_with_darkfield_microscopy_datasets
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 292 | 85.66 | 74 | 86.7 | - | - |
+| erythrocytes | 274 | 13.25 | 71 | 12.29 | - | - |
+| spirochaete | 231 | 1.09 | 57 | 1.01 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### Bactteria detection with darkfield microscopy
+
+***Note: The following experimental results are based on the data randomly partitioned according to the above method described in the dataset preparing section.***
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config | download |
+| :-------------: | :------: | :-------: | :----: | :---: | :---: | :--------------------------------------------------------------------------------------: | :----------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | 76.48 | 84.68 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | 61.06 | 63.69 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | 58.87 | 62.42 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py) | [model](<>) \| [log](<>) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/bactteria-detection_512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/bactteria-detection_512x512.py
new file mode 100644
index 00000000000..e3eab4e3868
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/bactteria-detection_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'BactteriaDetectionDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..ede58d785c1
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './bactteria-detection_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..bde3fa14acd
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './bactteria-detection_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..08e204f3809
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './bactteria-detection_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/datasets/bactteria-detection_dataset.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/datasets/bactteria-detection_dataset.py
new file mode 100644
index 00000000000..c95097b1acd
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/datasets/bactteria-detection_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class BactteriaDetectionDataset(BaseSegDataset):
+ """BactteriaDetectionDataset dataset.
+
+ In segmentation map annotation for BactteriaDetectionDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'erythrocytes', 'spirochaete'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/tools/prepare_dataset.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/tools/prepare_dataset.py
new file mode 100755
index 00000000000..8dcc719e262
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/tools/prepare_dataset.py
@@ -0,0 +1,33 @@
+import glob
+import os
+import shutil
+
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(
+ 'data/Bacteria_detection_with_darkfield_microscopy_datasets/images/*' +
+ img_suffix) # noqa
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+part_dir_dict = {0: 'train/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ basename.split('.')[0] + save_img_suffix)
+ shutil.copy(img, img_save_path)
+ mask_path = 'data/Bacteria_detection_with_darkfield_microscopy_datasets/masks/' + basename # noqa
+ mask = Image.open(mask_path).convert('L')
+ mask_save_path = os.path.join(
+ root_path, 'masks', part_dir,
+ basename.split('.')[0] + save_seg_map_suffix)
+ mask.save(mask_save_path)
diff --git a/projects/medical/2d_image/tools/split_seg_dataset.py b/projects/medical/2d_image/tools/split_seg_dataset.py
new file mode 100644
index 00000000000..9ab2e9282fa
--- /dev/null
+++ b/projects/medical/2d_image/tools/split_seg_dataset.py
@@ -0,0 +1,42 @@
+import argparse
+import glob
+import os
+
+from sklearn.model_selection import train_test_split
+
+
+def save_anno(img_list, file_path, remove_suffix=True):
+ if remove_suffix:
+ img_list = [
+ '/'.join(img_path.split('/')[-2:]) for img_path in img_list
+ ]
+ img_list = [
+ '.'.join(img_path.split('.')[:-1]) for img_path in img_list
+ ]
+ with open(file_path, 'w') as file_:
+ for x in list(img_list):
+ file_.write(x + '\n')
+
+
+if __name__ == '__main__':
+ parser = argparse.ArgumentParser()
+ parser.add_argument('--data_root', default='data/')
+ args = parser.parse_args()
+ data_root = args.data_root
+ if os.path.exists(os.path.join(data_root, 'masks/val')):
+ x_val = sorted(glob.glob(data_root + '/images/val/*.png'))
+ save_anno(x_val, data_root + '/val.txt')
+ if os.path.exists(os.path.join(data_root, 'masks/test')):
+ x_test = sorted(glob.glob(data_root + '/images/test/*.png'))
+ save_anno(x_test, data_root + '/test.txt')
+ if not os.path.exists(os.path.join(
+ data_root, 'masks/val')) and not os.path.exists(
+ os.path.join(data_root, 'masks/test')):
+ all_imgs = sorted(glob.glob(data_root + '/images/train/*.png'))
+ x_train, x_val = train_test_split(
+ all_imgs, test_size=0.2, random_state=0)
+ save_anno(x_train, data_root + '/train.txt')
+ save_anno(x_val, data_root + '/val.txt')
+ else:
+ x_train = sorted(glob.glob(data_root + '/images/train/*.png'))
+ save_anno(x_train, data_root + '/train.txt')
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/README.md b/projects/medical/2d_image/x_ray/chest_image_pneum/README.md
new file mode 100644
index 00000000000..a1cd27ba455
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/README.md
@@ -0,0 +1,147 @@
+# Chest Image Dataset for Pneumothorax Segmentation
+
+## Description
+
+This project supports **`Chest Image Dataset for Pneumothorax Segmentation`**, which can be downloaded from [here](https://tianchi.aliyun.com/dataset/83075).
+
+### Dataset Overview
+
+Pneumothorax can be caused by a blunt chest injury, damage from underlying lung disease, or most horrifying—it may occur for no obvious reason at all. On some occasions, a collapsed lung can be a life-threatening event.
+Pneumothorax is usually diagnosed by a radiologist on a chest x-ray, and can sometimes be very difficult to confirm. An accurate AI algorithm to detect pneumothorax would be useful in a lot of clinical scenarios. AI could be used to triage chest radiographs for priority interpretation, or to provide a more confident diagnosis for non-radiologists.
+
+The dataset is provided by the Society for Imaging Informatics in Medicine(SIIM), American College of Radiology (ACR),Society of Thoracic Radiology (STR) and MD.ai. You can develop a model to classify (and if present, segment) pneumothorax from a set of chest radiographic images. If successful, you could aid in the early recognition of pneumothoraces and save lives.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------------ |
+| [pneumothorax segmentation](https://tianchi.aliyun.com/dataset/83075) | thorax | segmentation | x_ray | 2 | 12089/-/3205 | yes/-/no | - | [CC-BY-SA-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 12089 | 99.75 | - | - | - | - |
+| pneumothorax area | 2669 | 0.25 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `chest_image_pneum/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://tianchi.aliyun.com/dataset/83075) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── chest_image_pneum
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 9637 | 99.75 | 2410 | 99.74 | - | - |
+| pneumothorax area | 2137 | 0.25 | 532 | 0.26 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### Bactteria detection with darkfield microscopy
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config | download |
+| :-------------: | :------: | :-------: | :----: | :--: | :---: | :------------------------------------------------------------------------------------: | :----------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | - | - | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | - | - | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | - | - | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py) | [model](<>) \| [log](<>) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/chest-image-pneum_512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/chest-image-pneum_512x512.py
new file mode 100644
index 00000000000..411229bd413
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/chest-image-pneum_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ChestImagePneumDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py
new file mode 100644
index 00000000000..0f26459467c
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './chest-image-pneum_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.chest-image-pneum_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py
new file mode 100644
index 00000000000..37b91889d8a
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './chest-image-pneum_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.chest-image-pneum_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py
new file mode 100644
index 00000000000..379e8181f3d
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './chest-image-pneum_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.chest-image-pneum_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/datasets/chest-image-pneum_dataset.py b/projects/medical/2d_image/x_ray/chest_image_pneum/datasets/chest-image-pneum_dataset.py
new file mode 100644
index 00000000000..aeee60ae92e
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/datasets/chest-image-pneum_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ChestImagePneumDataset(BaseSegDataset):
+ """ChestImagePneumDataset dataset.
+
+ In segmentation map annotation for ChestImagePneumDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('normal', 'pneumothorax area'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/chest_image_pneum/tools/prepare_dataset.py
new file mode 100755
index 00000000000..47eddc96dc5
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/tools/prepare_dataset.py
@@ -0,0 +1,73 @@
+import os
+
+import numpy as np
+import pandas as pd
+import pydicom
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.dcm'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = []
+for fpath, dirname, fnames in os.walk('data/chestimage_train_datasets'):
+ for fname in fnames:
+ if fname.endswith('.dcm'):
+ x_train.append(os.path.join(fpath, fname))
+x_test = []
+for fpath, dirname, fnames in os.walk('data/chestimage_test_datasets/'):
+ for fname in fnames:
+ if fname.endswith('.dcm'):
+ x_test.append(os.path.join(fpath, fname))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/test/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+
+def rle_decode(rle, width, height):
+ mask = np.zeros(width * height, dtype=np.uint8)
+ array = np.asarray([int(x) for x in rle.split()])
+ starts = array[0::2]
+ lengths = array[1::2]
+
+ current_position = 0
+ for index, start in enumerate(starts):
+ current_position += start
+ mask[current_position:current_position + lengths[index]] = 1
+ current_position += lengths[index]
+
+ return mask.reshape(width, height, order='F')
+
+
+part_dir_dict = {0: 'train/', 1: 'test/'}
+dict_from_csv = pd.read_csv(
+ root_path + 'chestimage_train-rle_datasets.csv', sep=',',
+ index_col=0).to_dict()[' EncodedPixels']
+
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_id = '.'.join(basename.split('.')[:-1])
+ if ith == 0 and (img_id not in dict_from_csv.keys()):
+ continue
+ image = pydicom.read_file(img).pixel_array
+ save_img_path = root_path + 'images/' + part_dir + '.'.join(
+ basename.split('.')[:-1]) + save_img_suffix
+ print(save_img_path)
+ img_h, img_w = image.shape[:2]
+ image = Image.fromarray(image)
+ image.save(save_img_path)
+ if ith == 1:
+ continue
+ if dict_from_csv[img_id] == '-1':
+ mask = np.zeros((img_h, img_w), dtype=np.uint8)
+ else:
+ mask = rle_decode(dict_from_csv[img_id], img_h, img_w)
+ save_mask_path = root_path + 'masks/' + part_dir + '.'.join(
+ basename.split('.')[:-1]) + save_seg_map_suffix
+ mask = Image.fromarray(mask)
+ mask.save(save_mask_path)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/README.md b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/README.md
new file mode 100644
index 00000000000..7cb099c8a43
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/README.md
@@ -0,0 +1,119 @@
+# Chest X-ray Images with Pneumothorax Masks
+
+## Description
+
+This project support **`Chest X-ray Images with Pneumothorax Masks `**, and the dataset used in this project can be downloaded from [here](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks).
+
+### Dataset Overview
+
+A pneumothorax (noo-moe-THOR-aks) is a collapsed lung. A pneumothorax occurs when air leaks into the space between your lung and chest wall. This air pushes on the outside of your lung and makes it collapse. Pneumothorax can be a complete lung collapse or a collapse of only a portion of the lung.
+
+A pneumothorax can be caused by a blunt or penetrating chest injury, certain medical procedures, or damage from underlying lung disease. Or it may occur for no obvious reason. Symptoms usually include sudden chest pain and shortness of breath. On some occasions, a collapsed lung can be a life-threatening event.
+
+Treatment for a pneumothorax usually involves inserting a needle or chest tube between the ribs to remove the excess air. However, a small pneumothorax may heal on its own.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release date | License |
+| --------------------------------------------------------------------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Chest-x-ray-images-with-pneumothorax-masks](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks) | throax | segmentation | x_ray | 2 | 10675/-/1372 | yes/-/yes | 2020 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 10675 | 99.7 | - | - | 1372 | 99.71 |
+| pneumothroax | 2379 | 0.3 | - | - | 290 | 0.29 |
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python 3.8
+- PyTorch 1.10.0
+- pillow(PIL) 9.3.0
+- scikit-learn(sklearn) 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of PYTHONPATH, which should point to the project's directory so that Python can locate the module files. In chest_x_ray_images_with_pneumothorax_masks/ root directory, run the following line to add the current directory to PYTHONPATH:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks) and decompression data to path 'data/'.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── chest_x_ray_images_with_pneumothorax_masks
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+To train on multiple GPUs, e.g. 8 GPUs, run the following command:
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH} --launcher pytorch --gpus 8
+```
+
+### Testing commands
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [x] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/chest-x-ray-images-with-pneumothorax-masks_512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/chest-x-ray-images-with-pneumothorax-masks_512x512.py
new file mode 100644
index 00000000000..96676de8616
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/chest-x-ray-images-with-pneumothorax-masks_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ChestPenumoMaskDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..76c214d04cb
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,20 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..066996dda94
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..a7065b82315
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..e5682ee76b7
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/datasets/chest-x-ray-images-with-pneumothorax-masks_dataset.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/datasets/chest-x-ray-images-with-pneumothorax-masks_dataset.py
new file mode 100644
index 00000000000..d32f597a5a0
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/datasets/chest-x-ray-images-with-pneumothorax-masks_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ChestPenumoMaskDataset(BaseSegDataset):
+ """ChestPenumoMaskDataset dataset.
+
+ In segmentation map annotation for ChestPenumoMaskDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'penumothroax'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/tools/prepare_dataset.py
new file mode 100644
index 00000000000..c7de1f19042
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/tools/prepare_dataset.py
@@ -0,0 +1,36 @@
+import glob
+import os
+import shutil
+
+from PIL import Image
+from sklearn.model_selection import train_test_split
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+all_imgs = glob.glob('data/siim-acr-pneumothorax/png_images/*' + img_suffix)
+x_train, x_test = train_test_split(all_imgs, test_size=0.2, random_state=0)
+
+print(len(x_train), len(x_test))
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/val/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+os.system('mkdir -p ' + root_path + 'masks/val/')
+
+part_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ basename.split('.')[0] + save_img_suffix)
+ shutil.copy(img, img_save_path)
+ mask_path = 'data/siim-acr-pneumothorax/png_masks/' + basename
+ mask = Image.open(mask_path).convert('L')
+ mask_save_path = os.path.join(
+ root_path, 'masks', part_dir,
+ basename.split('.')[0] + save_seg_map_suffix)
+ mask.save(mask_save_path)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/README.md b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/README.md
new file mode 100644
index 00000000000..8469219effa
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/README.md
@@ -0,0 +1,158 @@
+# Covid-19 CT Chest X-ray Dataset
+
+## Description
+
+This project supports **`Covid-19 CT Chest X-ray Dataset`**, which can be downloaded from [here](https://github.com/ieee8023/covid-chestxray-dataset).
+
+### Dataset Overview
+
+In the context of a COVID-19 pandemic, we want to improve prognostic predictions to triage and manage patient care. Data is the first step to developing any diagnostic/prognostic tool. While there exist large public datasets of more typical chest X-rays from the NIH \[Wang 2017\], Spain \[Bustos 2019\], Stanford \[Irvin 2019\], MIT \[Johnson 2019\] and Indiana University \[Demner-Fushman 2016\], there is no collection of COVID-19 chest X-rays or CT scans designed to be used for computational analysis.
+
+The 2019 novel coronavirus (COVID-19) presents several unique features [Fang, 2020](https://pubs.rsna.org/doi/10.1148/radiol.2020200432) and [Ai 2020](https://pubs.rsna.org/doi/10.1148/radiol.2020200642). While the diagnosis is confirmed using polymerase chain reaction (PCR), infected patients with pneumonia may present on chest X-ray and computed tomography (CT) images with a pattern that is only moderately characteristic for the human eye [Ng, 2020](https://pubs.rsna.org/doi/10.1148/ryct.2020200034). In late January, a Chinese team published a paper detailing the clinical and paraclinical features of COVID-19. They reported that patients present abnormalities in chest CT images with most having bilateral involvement [Huang 2020](). Bilateral multiple lobular and subsegmental areas of consolidation constitute the typical findings in chest CT images of intensive care unit (ICU) patients on admission [Huang 2020](). In comparison, non-ICU patients show bilateral ground-glass opacity and subsegmental areas of consolidation in their chest CT images [Huang 2020](). In these patients, later chest CT images display bilateral ground-glass opacity with resolved consolidation [Huang 2020]().
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Nnum. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release date | License |
+| ---------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------- | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------------- |
+| [Covid-19-ct-cxr](https://github.com/ieee8023/covid-chestxray-dataset) | thorax | segmentation | x_ray | 2 | 205/-/714 | yes/-/no | 2021 | [CC-BY-NC-SA 4.0](https://creativecommons.org/licenses/by-nc-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 205 | 72.84 | - | - | - | - |
+| lung | 205 | 27.16 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{cohen2020covidProspective,
+ title={{COVID-19} Image Data Collection: Prospective Predictions Are the Future},
+ author={Joseph Paul Cohen and Paul Morrison and Lan Dao and Karsten Roth and Tim Q Duong and Marzyeh Ghassemi},
+ journal={arXiv 2006.11988},
+ year={2020}
+}
+
+@article{cohen2020covid,
+ title={COVID-19 image data collection},
+ author={Joseph Paul Cohen and Paul Morrison and Lan Dao},
+ journal={arXiv 2003.11597},
+ year={2020}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `covid_19_ct_cxr/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://github.com/ieee8023/covid-chestxray-dataset) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```shell
+mkdir data && cd data
+git clone git@github.com:ieee8023/covid-chestxray-dataset.git
+cd ..
+python tools/prepare_dataset.py
+python ../../tools/split_seg_dataset.py
+```
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── covid_19_ct_cxr
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 164 | 72.88 | 41 | 72.69 | - | - |
+| lung | 164 | 27.12 | 41 | 27.31 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [x] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/covid-19-ct-cxr_512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/covid-19-ct-cxr_512x512.py
new file mode 100644
index 00000000000..5242d06c374
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/covid-19-ct-cxr_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'Covid19CXRDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..59a7bedaa06
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..83b8527d46d
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..10cfcbda6ec
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..aaccc8fd8dd
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/datasets/covid-19-ct-cxr_dataset.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/datasets/covid-19-ct-cxr_dataset.py
new file mode 100644
index 00000000000..68a1bb331f7
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/datasets/covid-19-ct-cxr_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class Covid19CXRDataset(BaseSegDataset):
+ """Covid19CXRDataset dataset.
+
+ In segmentation map annotation for Covid19CXRDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'lung'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/tools/prepare_dataset.py
new file mode 100644
index 00000000000..72f6435389d
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/tools/prepare_dataset.py
@@ -0,0 +1,52 @@
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+src_img_dir = os.path.join(root_path, 'covid-chestxray-dataset', 'images')
+src_mask_dir = os.path.join(root_path, 'covid-chestxray-dataset',
+ 'annotations/lungVAE-masks')
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+if __name__ == '__main__':
+
+ all_img_names = os.listdir(src_img_dir)
+ all_mask_names = os.listdir(src_mask_dir)
+
+ for img_name in all_img_names:
+ base_name = img_name.replace('.png', '')
+ base_name = base_name.replace('.jpg', '')
+ base_name = base_name.replace('.jpeg', '')
+ mask_name_orig = base_name + '_mask.png'
+ if mask_name_orig in all_mask_names:
+ mask_name = base_name + '.png'
+ src_img_path = os.path.join(src_img_dir, img_name)
+ src_mask_path = os.path.join(src_mask_dir, mask_name_orig)
+ tgt_img_path = os.path.join(tgt_img_train_dir, img_name)
+ tgt_mask_path = os.path.join(tgt_mask_train_dir, mask_name)
+
+ img = Image.open(src_img_path).convert('RGB')
+ img.save(tgt_img_path)
+ mask = np.array(Image.open(src_mask_path))
+ mask = convert_label(mask, {0: 0, 255: 1})
+ mask = Image.fromarray(mask)
+ mask.save(tgt_mask_path)
+ else:
+ src_img_path = os.path.join(src_img_dir, img_name)
+ tgt_img_path = os.path.join(tgt_img_test_dir, img_name)
+ img = Image.open(src_img_path).convert('RGB')
+ img.save(tgt_img_path)
diff --git a/projects/medical/2d_image/x_ray/crass/README.md b/projects/medical/2d_image/x_ray/crass/README.md
new file mode 100644
index 00000000000..0621205be81
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/README.md
@@ -0,0 +1,144 @@
+# Chest Radiograph Anatomical Structure Segmentation (CRASS)
+
+## Description
+
+This project supports **`Chest Radiograph Anatomical Structure Segmentation (CRASS) `**, which can be downloaded from [here](https://crass.grand-challenge.org/).
+
+### Dataset Overview
+
+A set of consecutively obtained posterior-anterior chest radiograph were selected from a database containing images acquired at two sites in sub Saharan Africa with a high tuberculosis incidence. All subjects were 15 years or older. Images from digital chest radiography units were used (Delft Imaging Systems, The Netherlands) of varying resolutions, with a typical resolution of 1800--2000 pixels, the pixel size was 250 lm isotropic. From the total set of images, 225 were considered to be normal by an expert radiologist, while 333 of the images contained abnormalities. Of the abnormal images, 220 contained abnormalities in the upper area of the lung where the clavicle is located. The data was divided into a training and a test set. The training set consisted of 299 images, the test set of 249 images.
+The current data is still incomplete and to be added later.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [crass](https://crass.grand-challenge.org/) | pulmonary | segmentation | x_ray | 2 | 299/-/234 | yes/-/no | 2021 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 299 | 98.38 | - | - | - | - |
+| clavicles | 299 | 1.62 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{HOGEWEG20121490,
+ title={Clavicle segmentation in chest radiographs},
+ journal={Medical Image Analysis},
+ volume={16},
+ number={8},
+ pages={1490-1502},
+ year={2012}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `crass/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://crass.grand-challenge.org/) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── crass
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 227 | 98.38 | 57 | 98.39 | - | - |
+| clavicles | 227 | 1.62 | 57 | 1.61 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/crass/configs/crass_512x512.py b/projects/medical/2d_image/x_ray/crass/configs/crass_512x512.py
new file mode 100644
index 00000000000..1425f50cc46
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/crass_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'CRASSDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='tval.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_crass-512x512.py
new file mode 100644
index 00000000000..b52bc78f790
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_crass-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_crass-512x512.py
new file mode 100644
index 00000000000..45242c65b48
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_crass-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_crass-512x512.py
new file mode 100644
index 00000000000..bcf9d0a5ca5
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_crass-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-lr0.01-sigmoid-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-lr0.01-sigmoid-20k_crass-512x512.py
new file mode 100644
index 00000000000..0dde736bf76
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-lr0.01-sigmoid-20k_crass-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/datasets/crass_dataset.py b/projects/medical/2d_image/x_ray/crass/datasets/crass_dataset.py
new file mode 100644
index 00000000000..f6b5c5228bc
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/datasets/crass_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class CRASSDataset(BaseSegDataset):
+ """CRASSDataset dataset.
+
+ In segmentation map annotation for CRASSDataset, 0 stands for background,
+ which is included in 2 categories. ``reduce_zero_label`` is fixed to
+ False. The ``img_suffix`` is fixed to '.png' and ``seg_map_suffix`` is
+ fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'clavicles'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/crass/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/crass/tools/prepare_dataset.py
new file mode 100644
index 00000000000..bbd5d8891d4
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/tools/prepare_dataset.py
@@ -0,0 +1,84 @@
+import glob
+import os
+
+import cv2
+import SimpleITK as sitk
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.tif'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+src_img_train_dir = os.path.join(root_path, 'CRASS/data_train')
+src_mask_train_dir = os.path.join(root_path, 'CRASS/mask_mhd')
+src_img_test_dir = os.path.join(root_path, 'CRASS/data_test')
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def read_single_array_from_med(path):
+ return sitk.GetArrayFromImage(sitk.ReadImage(path)).squeeze()
+
+
+def convert_meds_into_pngs(src_dir,
+ tgt_dir,
+ suffix='.dcm',
+ norm_min=0,
+ norm_max=255,
+ convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, '.png')
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = read_single_array_from_med(src_path)
+ if norm_min is not None and norm_max is not None:
+ img = cv2.normalize(img, None, norm_min, norm_max, cv2.NORM_MINMAX,
+ cv2.CV_8U)
+ pil = Image.fromarray(img).convert(convert)
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+convert_meds_into_pngs(
+ src_img_train_dir,
+ tgt_img_train_dir,
+ suffix='.mhd',
+ norm_min=0,
+ norm_max=255,
+ convert='RGB')
+
+convert_meds_into_pngs(
+ src_img_test_dir,
+ tgt_img_test_dir,
+ suffix='.mhd',
+ norm_min=0,
+ norm_max=255,
+ convert='RGB')
+
+convert_meds_into_pngs(
+ src_mask_train_dir,
+ tgt_mask_train_dir,
+ suffix='.mhd',
+ norm_min=0,
+ norm_max=1,
+ convert='L')
diff --git a/projects/pp_mobileseg/README.md b/projects/pp_mobileseg/README.md
new file mode 100644
index 00000000000..c9f9c128e74
--- /dev/null
+++ b/projects/pp_mobileseg/README.md
@@ -0,0 +1,123 @@
+# PP-MobileSeg: Exploring Transformer Blocks for Efficient Mobile Segmentation.
+
+## Reference
+
+> [PP-MobileSeg: Explore the Fast and Accurate Semantic Segmentation Model on Mobile Devices. ](https://arxiv.org/abs/2304.05152)
+
+## Introduction
+
+Official Repo
+
+Code Snippet
+
+##
diff --git a/projects/isnet/README.md b/projects/isnet/README.md
index 3a3172a9d99..0a79ad6a4fa 100644
--- a/projects/isnet/README.md
+++ b/projects/isnet/README.md
@@ -96,11 +96,11 @@ A project does not necessarily have to be finished in a single PR, but it's esse
- [ ] Type hints and docstrings
-
+
- [ ] Unit tests
-
+
- [ ] Code polishing
@@ -108,10 +108,10 @@ A project does not necessarily have to be finished in a single PR, but it's esse
- [ ] Metafile.yml
-
+
- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
-
+
- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/mapillary_dataset/README.md b/projects/mapillary_dataset/README.md
index cdc61d53a93..44a1e33ef93 100644
--- a/projects/mapillary_dataset/README.md
+++ b/projects/mapillary_dataset/README.md
@@ -10,7 +10,7 @@ This project implements **`Mapillary Vistas Dataset`**
### Dataset preparing
-Preparing `Mapillary Vistas Dataset` dataset following [Mapillary Vistas Dataset Preparing Guide](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md)
+Preparing `Mapillary Vistas Dataset` dataset following [Mapillary Vistas Dataset Preparing Guide](https://github.com/open-mmlab/mmsegmentation/tree/main/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md)
```none
mmsegmentation
diff --git a/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md b/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md
index fa074543300..c5cbc0f9b80 100644
--- a/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md
+++ b/projects/mapillary_dataset/docs/en/user_guides/2_dataset_prepare.md
@@ -47,7 +47,7 @@
```
- You could set Datasets version with `MapillaryDataset_v1` and `MapillaryDataset_v2` in your configs.
- View the Mapillary Vistas Datasets config file here [V1.2](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/datasets/mapillary_v1.py) and [V2.0](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/datasets/mapillary_v2.py)
+ View the Mapillary Vistas Datasets config file here [V1.2](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/datasets/mapillary_v1.py) and [V2.0](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/datasets/mapillary_v2.py)
- **View datasets labels index and palette**
diff --git a/projects/medical/2d_image/ct/cranium/README.md b/projects/medical/2d_image/ct/cranium/README.md
new file mode 100644
index 00000000000..d3fa64ea406
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/README.md
@@ -0,0 +1,142 @@
+# Brain CT Images with Intracranial Hemorrhage Masks (Cranium)
+
+## Description
+
+This project supports **`Brain CT Images with Intracranial Hemorrhage Masks (Cranium)`**, which can be downloaded from [here](https://www.kaggle.com/datasets/vbookshelf/computed-tomography-ct-images).
+
+### Dataset Overview
+
+This dataset consists of head CT (Computed Thomography) images in jpg format. There are 2500 brain window images and 2500 bone window images, for 82 patients. There are approximately 30 image slices per patient. 318 images have associated intracranial image masks. Also included are csv files containing hemorrhage diagnosis data and patient data.
+This is version 1.0.0 of this dataset. A full description of this dataset as well as updated versions can be found here:
+https://physionet.org/content/ct-ich/1.0.0/
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ----------------------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------- |
+| [Cranium](https://www.kaggle.com/datasets/vbookshelf/computed-tomography-ct-images) | head_and_neck | segmentation | ct | 2 | 2501/-/- | yes/-/- | 2020 | [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 2501 | 99.93 | - | - | - | - |
+| hemorrhage | 318 | 0.07 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```
+@article{hssayeni2020computed,
+ title={Computed tomography images for intracranial hemorrhage detection and segmentation},
+ author={Hssayeni, Murtadha and Croock, MS and Salman, AD and Al-khafaji, HF and Yahya, ZA and Ghoraani, B},
+ journal={Intracranial Hemorrhage Segmentation Using A Deep Convolutional Model. Data},
+ volume={5},
+ number={1},
+ pages={179},
+ year={2020}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `cranium/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://www.kaggle.com/datasets/vbookshelf/computed-tomography-ct-images) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── ct
+ │ │ │ │ ├── cranium
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 2000 | 99.93 | 501 | 99.92 | - | - |
+| hemorrhage | 260 | 0.07 | 260 | 0.08 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/ct/cranium/configs/cranium_512x512.py b/projects/medical/2d_image/ct/cranium/configs/cranium_512x512.py
new file mode 100644
index 00000000000..d9b44362a5c
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/cranium_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'CraniumDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_cranium-512x512.py
new file mode 100644
index 00000000000..ac013a215ae
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_cranium-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_cranium-512x512.py
new file mode 100644
index 00000000000..c71110a21f7
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_cranium-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_cranium-512x512.py
new file mode 100644
index 00000000000..abbdac285b2
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_cranium-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_cranium-512x512.py b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_cranium-512x512.py
new file mode 100644
index 00000000000..418595268f9
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_cranium-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './cranium_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.cranium_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/ct/cranium/datasets/cranium_dataset.py b/projects/medical/2d_image/ct/cranium/datasets/cranium_dataset.py
new file mode 100644
index 00000000000..d65f1cbfc68
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/datasets/cranium_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class CraniumDataset(BaseSegDataset):
+ """CraniumDataset dataset.
+
+ In segmentation map annotation for CraniumDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'hemorrhage'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/ct/cranium/tools/prepare_dataset.py b/projects/medical/2d_image/ct/cranium/tools/prepare_dataset.py
new file mode 100644
index 00000000000..1aa4e435614
--- /dev/null
+++ b/projects/medical/2d_image/ct/cranium/tools/prepare_dataset.py
@@ -0,0 +1,66 @@
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+
+def read_single_array_from_pil(path):
+ return np.asarray(Image.open(path))
+
+
+def save_png_from_array(arr, save_path, mode=None):
+ Image.fromarray(arr, mode=mode).save(save_path)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+patients_dir = os.path.join(
+ root_path, 'Cranium/computed-tomography-images-for-' +
+ 'intracranial-hemorrhage-detection-and-segmentation-1.0.0' +
+ '/Patients_CT')
+
+patients = sorted(os.listdir(patients_dir))
+for p in patients:
+ data_dir = os.path.join(patients_dir, p, 'brain')
+ file_names = os.listdir(data_dir)
+ img_w_mask_names = [
+ _.replace('_HGE_Seg', '') for _ in file_names if 'Seg' in _
+ ]
+ img_wo_mask_names = [
+ _ for _ in file_names if _ not in img_w_mask_names and 'Seg' not in _
+ ]
+
+ for file_name in file_names:
+ path = os.path.join(data_dir, file_name)
+ img = read_single_array_from_pil(path)
+ tgt_name = file_name.replace('.jpg', img_suffix)
+ tgt_name = p + '_' + tgt_name
+ if 'Seg' in file_name: # is a mask
+ tgt_name = tgt_name.replace('_HGE_Seg', '')
+ mask_path = os.path.join(tgt_mask_dir, tgt_name)
+ mask = convert_label(img, convert_dict={0: 0, 255: 1})
+ save_png_from_array(mask, mask_path)
+ else:
+ img_path = os.path.join(tgt_img_dir, tgt_name)
+ pil = Image.fromarray(img).convert('RGB')
+ pil.save(img_path)
+
+ if file_name in img_wo_mask_names:
+ mask = np.zeros_like(img, dtype=np.uint8)
+ mask_path = os.path.join(tgt_mask_dir, tgt_name)
+ save_png_from_array(mask, mask_path)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/README.md b/projects/medical/2d_image/dermoscopy/isic2016_task1/README.md
new file mode 100644
index 00000000000..6e44e415ed6
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/README.md
@@ -0,0 +1,149 @@
+# ISIC-2016 Task1
+
+## Description
+
+This project support **`ISIC-2016 Task1 `**, and the dataset used in this project can be downloaded from [here](https://challenge.isic-archive.com/data/#2016).
+
+### Dataset Overview
+
+The overarching goal of the challenge is to develop image analysis tools to enable the automated diagnosis of melanoma from dermoscopic images.
+
+This challenge provides training data (~900 images) for participants to engage in all 3 components of lesion image analysis. A separate test dataset (~350 images) will be provided for participants to generate and submit automated results.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------- | ----------------- | ------------ | ---------- | ------------ | --------------------- | ---------------------- | ------------ | ---------------------------------------------------------------------- |
+| [ISIC-2016 Task1](https://challenge.isic-archive.com/data/#2016) | full body | segmentation | dermoscopy | 2 | 900/-/379- | yes/-/yes | 2016 | [CC-0](https://creativecommons.org/share-your-work/public-domain/cc0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 900 | 82.08 | - | - | 379 | 81.98 |
+| skin lesion | 900 | 17.92 | - | - | 379 | 18.02 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python 3.8
+- PyTorch 1.10.0
+- pillow(PIL) 9.3.0
+- scikit-learn(sklearn) 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of PYTHONPATH, which should point to the project's directory so that Python can locate the module files. In isic2016_task1/ root directory, run the following line to add the current directory to PYTHONPATH:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://challenge.isic-archive.com/data/#2016) and decompression data to path 'data/'.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── dermoscopy
+ │ │ │ │ ├── isic2016_task1
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+```
+
+### Training commands
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+To train on multiple GPUs, e.g. 8 GPUs, run the following command:
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH} --launcher pytorch --gpus 8
+```
+
+### Testing commands
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### ISIC-2016 Task1
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config |
+| :-------------: | :------: | :-------: | :----: | :--: | :---: | :---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py
new file mode 100644
index 00000000000..5638de4d560
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2016-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2016-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2016-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py
new file mode 100644
index 00000000000..bf17faa5380
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2016-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2016-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2016-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py
new file mode 100644
index 00000000000..f7bfcf6158f
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2016-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2016-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2016-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/isic2016-task1_512x512.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/isic2016-task1_512x512.py
new file mode 100644
index 00000000000..029f5d4d7ec
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/configs/isic2016-task1_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ISIC2017Task1'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='test.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/datasets/isic2016-task1_dataset.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/datasets/isic2016-task1_dataset.py
new file mode 100644
index 00000000000..8f11bdd0ba9
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/datasets/isic2016-task1_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ISIC2017Task1(BaseSegDataset):
+ """ISIC2017Task1 dataset.
+
+ In segmentation map annotation for ISIC2017Task1,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('normal', 'skin lesion'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/dermoscopy/isic2016_task1/tools/prepare_dataset.py b/projects/medical/2d_image/dermoscopy/isic2016_task1/tools/prepare_dataset.py
new file mode 100755
index 00000000000..ef4dad54086
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2016_task1/tools/prepare_dataset.py
@@ -0,0 +1,120 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+
+
+def check_maskid(train_imgs):
+ for i in train_masks:
+ img = Image.open(i)
+ print(np.unique(np.array(img)))
+
+
+def reformulate_file(image_list, mask_list):
+ file_list = []
+ for idx, (imgp,
+ maskp) in enumerate(zip(sorted(image_list), sorted(mask_list))):
+ item = {'image': imgp, 'label': maskp}
+ file_list.append(item)
+ return file_list
+
+
+def check_file_exist(pair_list):
+ rel_path = os.getcwd()
+ for idx, sample in enumerate(pair_list):
+ image_path = sample['image']
+ assert os.path.exists(os.path.join(rel_path, image_path))
+ if 'label' in sample:
+ mask_path = sample['label']
+ assert os.path.exists(os.path.join(rel_path, mask_path))
+ print('all file path ok!')
+
+
+def convert_maskid(mask):
+ # add mask id conversion
+ arr_mask = np.array(mask).astype(np.uint8)
+ arr_mask[arr_mask == 255] = 1
+ return Image.fromarray(arr_mask)
+
+
+def process_dataset(file_lists, part_dir_dict):
+ for ith, part in enumerate(file_lists):
+ part_dir = part_dir_dict[ith]
+ for sample in part:
+ # read image and mask
+ image_path = sample['image']
+ if 'label' in sample:
+ mask_path = sample['label']
+
+ basename = os.path.basename(image_path)
+ targetname = basename.split('.')[0] # from image name
+
+ # check image file
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ targetname + save_img_suffix)
+ if not os.path.exists(img_save_path):
+ if not image_path.endswith('.png'):
+ src = Image.open(image_path)
+ src.save(img_save_path)
+ else:
+ shutil.copy(image_path, img_save_path)
+
+ if mask_path is not None:
+ mask_save_path = os.path.join(root_path, 'masks', part_dir,
+ targetname + save_seg_map_suffix)
+ if not os.path.exists(mask_save_path):
+ # check mask file
+ mask = Image.open(mask_path).convert('L')
+ # convert mask id
+ mask = convert_maskid(mask)
+ if not mask_path.endswith('.png'):
+ mask.save(mask_save_path)
+ else:
+ mask.save(mask_save_path)
+
+ # print image num
+ part_dir_folder = os.path.join(root_path, 'images', part_dir)
+ print(
+ f'{part_dir} has {len(os.listdir(part_dir_folder))} images completed!' # noqa
+ )
+
+
+if __name__ == '__main__':
+
+ root_path = 'data/' # original file
+ img_suffix = '.jpg'
+ seg_map_suffix = '.png'
+ save_img_suffix = '.png'
+ save_seg_map_suffix = '.png'
+
+ train_imgs = glob.glob('data/ISBI2016_ISIC_Part1_Training_Data/*' # noqa
+ + img_suffix)
+ train_masks = glob.glob(
+ 'data/ISBI2016_ISIC_Part1_Training_GroundTruth/*' # noqa
+ + seg_map_suffix)
+
+ test_imgs = glob.glob('data/ISBI2016_ISIC_Part1_Test_Data/*' + img_suffix)
+ test_masks = glob.glob(
+ 'data/ISBI2016_ISIC_Part1_Test_GroundTruth/*' # noqa
+ + seg_map_suffix)
+
+ assert len(train_imgs) == len(train_masks)
+ assert len(test_imgs) == len(test_masks)
+
+ print(f'training images: {len(train_imgs)}, test images: {len(test_imgs)}')
+
+ os.system('mkdir -p ' + root_path + 'images/train/')
+ os.system('mkdir -p ' + root_path + 'images/test/')
+ os.system('mkdir -p ' + root_path + 'masks/train/')
+ os.system('mkdir -p ' + root_path + 'masks/test/')
+
+ train_pair_list = reformulate_file(train_imgs, train_masks)
+ test_pair_list = reformulate_file(test_imgs, test_masks)
+
+ check_file_exist(train_pair_list)
+ check_file_exist(test_pair_list)
+
+ part_dir_dict = {0: 'train/', 1: 'test/'}
+ process_dataset([train_pair_list, test_pair_list], part_dir_dict)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/README.md b/projects/medical/2d_image/dermoscopy/isic2017_task1/README.md
new file mode 100644
index 00000000000..c7cc27096be
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/README.md
@@ -0,0 +1,158 @@
+# ISIC-2017 Task1
+
+## Description
+
+This project support **`ISIC-2017 Task1 `**, and the dataset used in this project can be downloaded from [here](https://challenge.isic-archive.com/data/#2017).
+
+### Dataset Overview
+
+The goal of the challenge is to help participants develop image analysis tools to enable the automated diagnosis of melanoma from dermoscopic images.
+
+This challenge provides training data (~2000 images) for participants to engage in all 3 components of lesion image analysis. A separate public validation dataset (~150 images) and blind held-out test dataset (~600 images) will be provided for participants to generate and submit automated results.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------- | ----------------- | ------------ | ---------- | ------------ | --------------------- | ---------------------- | ------------ | ---------------------------------------------------------------------- |
+| [ISIC-2017 Task1](https://challenge.isic-archive.com/data/#2017) | full body | segmentation | dermoscopy | 2 | 2000/150/600 | yes/yes/yes | 2017 | [CC-0](https://creativecommons.org/share-your-work/public-domain/cc0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 2000 | 82.86 | 150 | 73.88 | 600 | 70.62 |
+| skin lesion | 2000 | 17.14 | 150 | 26.12 | 600 | 29.38 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python 3.8
+- PyTorch 1.10.0
+- pillow(PIL) 9.3.0
+- scikit-learn(sklearn) 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of PYTHONPATH, which should point to the project's directory so that Python can locate the module files. In isic2017_task1/ root directory, run the following line to add the current directory to PYTHONPATH:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://challenge.isic-archive.com/data/#2017) and decompression data to path 'data/'.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── dermoscopy
+ │ │ │ │ ├── isic2017_task1
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── val
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── val
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+```
+
+### Training commands
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+To train on multiple GPUs, e.g. 8 GPUs, run the following command:
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH} --launcher pytorch --gpus 8
+```
+
+### Testing commands
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### ISIC-2017 Task1
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config |
+| :-------------: | :------: | :-------: | :----: | :--: | :---: | :---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | - | - | [config](https://github.com/open-mmlab/mmsegmentation/tree/dev-1.x/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py
new file mode 100644
index 00000000000..58d0a125d33
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_isic2017-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2017-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2017-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py
new file mode 100644
index 00000000000..3becacf64fb
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_isic2017-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2017-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2017-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py
new file mode 100644
index 00000000000..654ef4dc3d5
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_isic2017-task1-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './isic2017-task1_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.isic2017-task1_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/isic2017-task1_512x512.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/isic2017-task1_512x512.py
new file mode 100644
index 00000000000..95997a10997
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/configs/isic2017-task1_512x512.py
@@ -0,0 +1,41 @@
+dataset_type = 'ISIC2017Task1'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/train/', seg_map_path='masks/train/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(img_path='images/val/', seg_map_path='masks/val/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/datasets/isic2017-task1_dataset.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/datasets/isic2017-task1_dataset.py
new file mode 100644
index 00000000000..8f11bdd0ba9
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/datasets/isic2017-task1_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ISIC2017Task1(BaseSegDataset):
+ """ISIC2017Task1 dataset.
+
+ In segmentation map annotation for ISIC2017Task1,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('normal', 'skin lesion'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/dermoscopy/isic2017_task1/tools/prepare_dataset.py b/projects/medical/2d_image/dermoscopy/isic2017_task1/tools/prepare_dataset.py
new file mode 100755
index 00000000000..b3643c93590
--- /dev/null
+++ b/projects/medical/2d_image/dermoscopy/isic2017_task1/tools/prepare_dataset.py
@@ -0,0 +1,127 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+
+
+def check_maskid(train_imgs):
+ for i in train_masks:
+ img = Image.open(i)
+ print(np.unique(np.array(img)))
+
+
+def reformulate_file(image_list, mask_list):
+ file_list = []
+ for idx, (imgp,
+ maskp) in enumerate(zip(sorted(image_list), sorted(mask_list))):
+ item = {'image': imgp, 'label': maskp}
+ file_list.append(item)
+ return file_list
+
+
+def convert_maskid(mask):
+ # add mask id conversion
+ arr_mask = np.array(mask).astype(np.uint8)
+ arr_mask[arr_mask == 255] = 1
+ return Image.fromarray(arr_mask)
+
+
+def check_file_exist(pair_list):
+ rel_path = os.getcwd()
+ for idx, sample in enumerate(pair_list):
+ image_path = sample['image']
+ assert os.path.exists(os.path.join(rel_path, image_path))
+ if 'label' in sample:
+ mask_path = sample['label']
+ assert os.path.exists(os.path.join(rel_path, mask_path))
+ print('all file path ok!')
+
+
+def process_dataset(file_lists, part_dir_dict):
+ for ith, part in enumerate(file_lists):
+ part_dir = part_dir_dict[ith]
+ for sample in part:
+ # read image and mask
+ image_path = sample['image']
+ if 'label' in sample:
+ mask_path = sample['label']
+
+ basename = os.path.basename(image_path)
+ targetname = basename.split('.')[0] # from image name
+
+ # check image file
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ targetname + save_img_suffix)
+ if not os.path.exists(img_save_path):
+ if not image_path.endswith('.png'):
+ src = Image.open(image_path)
+ src.save(img_save_path)
+ else:
+ shutil.copy(image_path, img_save_path)
+
+ if mask_path is not None:
+ mask_save_path = os.path.join(root_path, 'masks', part_dir,
+ targetname + save_seg_map_suffix)
+ if not os.path.exists(mask_save_path):
+ # check mask file
+ mask = Image.open(mask_path).convert('L')
+ # convert mask id
+ mask = convert_maskid(mask)
+ if not mask_path.endswith('.png'):
+ mask.save(mask_save_path)
+ else:
+ mask.save(mask_save_path)
+
+ # print image num
+ part_dir_folder = os.path.join(root_path, 'images', part_dir)
+ print(
+ f'{part_dir} has {len(os.listdir(part_dir_folder))} images completed!' # noqa
+ )
+
+
+if __name__ == '__main__':
+
+ root_path = 'data/' # original file
+ img_suffix = '.jpg'
+ seg_map_suffix = '.png'
+ save_img_suffix = '.png'
+ save_seg_map_suffix = '.png'
+
+ train_imgs = glob.glob('data/ISIC-2017_Training_Data/*' + img_suffix)
+ train_masks = glob.glob('data/ISIC-2017_Training_Part1_GroundTruth/*' +
+ seg_map_suffix)
+
+ val_imgs = glob.glob('data/ISIC-2017_Validation_Data/*' + img_suffix)
+ val_masks = glob.glob('data/ISIC-2017_Validation_Part1_GroundTruth/*' +
+ seg_map_suffix)
+
+ test_imgs = glob.glob('data/ISIC-2017_Test_v2_Data/*' + img_suffix)
+ test_masks = glob.glob('data/ISIC-2017_Test_v2_Part1_GroundTruth/*' +
+ seg_map_suffix)
+
+ assert len(train_imgs) == len(train_masks)
+ assert len(val_imgs) == len(val_masks)
+ assert len(test_imgs) == len(test_masks)
+
+ os.system('mkdir -p ' + root_path + 'images/train/')
+ os.system('mkdir -p ' + root_path + 'images/val/')
+ os.system('mkdir -p ' + root_path + 'images/test/')
+ os.system('mkdir -p ' + root_path + 'masks/train/')
+ os.system('mkdir -p ' + root_path + 'masks/val/')
+ os.system('mkdir -p ' + root_path + 'masks/test/')
+
+ part_dir_dict = {0: 'train/', 1: 'val/', 2: 'test/'}
+
+ train_pair_list = reformulate_file(train_imgs, train_masks)
+ val_pair_list = reformulate_file(val_imgs, val_masks)
+ test_pair_list = reformulate_file(test_imgs, test_masks)
+
+ check_file_exist(train_pair_list)
+ check_file_exist(val_pair_list)
+ check_file_exist(test_pair_list)
+
+ part_dir_dict = {0: 'train/', 1: 'val/', 2: 'test/'}
+ process_dataset([train_pair_list, val_pair_list, test_pair_list],
+ part_dir_dict)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/README.md b/projects/medical/2d_image/endoscopy/kvasir_seg/README.md
new file mode 100644
index 00000000000..ea597bc4401
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/README.md
@@ -0,0 +1,145 @@
+# Kvasir-Sessile Dataset (Kvasir SEG)
+
+## Description
+
+This project supports **`Kvasir-Sessile Dataset (Kvasir SEG) `**, which can be downloaded from [here](https://opendatalab.com/Kvasir-Sessile_dataset).
+
+## Dataset Overview
+
+The Kvasir-SEG dataset contains polyp images and their corresponding ground truth from the Kvasir Dataset v2. The resolution of the images contained in Kvasir-SEG varies from 332x487 to 1920x1072 pixels.
+
+
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------------- | ----------------- | ------------ | --------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------- |
+| [Kvarsir-SEG](https://opendatalab.com/Kvasir-Sessile_dataset) | abdomen | segmentation | endoscopy | 2 | 196/-/- | yes/-/- | 2020 | [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 196 | 92.31 | - | - | - | - |
+| polyp | 196 | 7.69 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{jha2020kvasir,
+ title={Kvasir-seg: A segmented polyp dataset},
+ author={Jha, Debesh and Smedsrud, Pia H and Riegler, Michael A and Halvorsen, P{\aa}l and Lange, Thomas de and Johansen, Dag and Johansen, H{\aa}vard D},
+ booktitle={International Conference on Multimedia Modeling},
+ pages={451--462},
+ year={2020},
+ organization={Springer}
+ }
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `kvasir_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.com/Kvasir-Sessile_dataset) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── endoscopy
+ │ │ │ │ ├── kvasir_seg
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 156 | 92.28 | 40 | 92.41 | - | - |
+| polyp | 156 | 7.72 | 40 | 7.59 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg .configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..145d5a7a172
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_kvasir-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..3ea05c51090
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..7e064a716aa
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-512x512.py
new file mode 100644
index 00000000000..0fc1d6e99d7
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './kvasir-seg_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/configs/kvasir-seg_512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/kvasir-seg_512x512.py
new file mode 100644
index 00000000000..e8b2467f8cf
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/configs/kvasir-seg_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'KvasirSEGDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/datasets/kvasir-seg_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg/datasets/kvasir-seg_dataset.py
new file mode 100644
index 00000000000..9d601328eb6
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/datasets/kvasir-seg_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class KvasirSEGDataset(BaseSegDataset):
+ """KvasirSEGDataset dataset.
+
+ In segmentation map annotation for KvasirSEGDataset, 0 stands for
+ background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix`` is
+ fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'polyp'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg/tools/prepare_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg/tools/prepare_dataset.py
new file mode 100644
index 00000000000..74c43e96351
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg/tools/prepare_dataset.py
@@ -0,0 +1,87 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '.jpg'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ os.path.join(root_path, 'sessile-main-Kvasir-SEG/images'),
+ tgt_img_dir,
+ suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ os.path.join(root_path, 'sessile-main-Kvasir-SEG/masks'),
+ tgt_mask_dir,
+ suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/README.md b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/README.md
new file mode 100644
index 00000000000..80eb00f51bd
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/README.md
@@ -0,0 +1,145 @@
+# Kvasir-SEG Segmented Polyp Dataset from Aliyun (Kvasir SEG Aliyun)
+
+## Description
+
+This project supports **`Kvasir-SEG Segmented Polyp Dataset from Aliyun (Kvasir SEG Aliyun) `**, which can be downloaded from [here](https://tianchi.aliyun.com/dataset/84385).
+
+### Dataset Overview
+
+Colorectal cancer is the second most common cancer type among women and third most common among men. Polyps are precursors to colorectal cancer and therefore important to detect and remove at an early stage. Polyps are found in nearly half of the individuals at age 50 that undergo a colonoscopy screening, and their frequency increase with age.Polyps are abnormal tissue growth from the mucous membrane, which is lining the inside of the GI tract, and can sometimes be cancerous. Colonoscopy is the gold standard for detection and assessment of these polyps with subsequent biopsy and removal of the polyps. Early disease detection has a huge impact on survival from colorectal cancer. Increasing the detection of polyps has been shown to decrease risk of colorectal cancer. Thus, automatic detection of more polyps at an early stage can play a crucial role in prevention and survival from colorectal cancer.
+
+The Kvasir-SEG dataset is based on the previous Kvasir dataset, which is the first multi-class dataset for gastrointestinal (GI) tract disease detection and classification. It contains annotated polyp images and their corresponding masks. The pixels depicting polyp tissue, the ROI, are represented by the foreground (white mask), while the background (in black) does not contain positive pixels. These images were collected and verified by experienced gastroenterologists from Vestre Viken Health Trust in Norway. The classes include anatomical landmarks, pathological findings and endoscopic procedures.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------ | ----------------- | ------------ | --------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------- |
+| [kvasir-seg](https://tianchi.aliyun.com/dataset/84385) | abdomen | segmentation | endoscopy | 2 | 1000/-/- | yes/-/- | 2020 | [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 1000 | 84.72 | - | - | - | - |
+| polyp | 1000 | 15.28 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{jha2020kvasir,
+ title={Kvasir-seg: A segmented polyp dataset},
+ author={Jha, Debesh and Smedsrud, Pia H and Riegler, Michael A and Halvorsen, P{\aa}l and Lange, Thomas de and Johansen, Dag and Johansen, H{\aa}vard D},
+ booktitle={International Conference on Multimedia Modeling},
+ pages={451--462},
+ year={2020},
+ organization={Springer}
+ }
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `kvasir_seg_aliyun/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://tianchi.aliyun.com/dataset/84385) and decompression data to path 'data/.'.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── endoscopy
+ │ │ │ │ ├── kvasir_seg_aliyun
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 800 | 84.66 | 200 | 84.94 | - | - |
+| polyp | 800 | 15.34 | 200 | 15.06 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..b59db95232b
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..6c526680cd4
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..a192a5bd240
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_kvasir-seg-aliyun-512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_kvasir-seg-aliyun-512x512.py
new file mode 100644
index 00000000000..5325e1f0807
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_kvasir-seg-aliyun-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './kvasir-seg-aliyun_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.kvasir-seg-aliyun_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/kvasir-seg-aliyun_512x512.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/kvasir-seg-aliyun_512x512.py
new file mode 100644
index 00000000000..5f868804679
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/configs/kvasir-seg-aliyun_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'KvasirSEGAliyunDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/datasets/kvasir-seg-aliyun_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/datasets/kvasir-seg-aliyun_dataset.py
new file mode 100644
index 00000000000..198caf07bcd
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/datasets/kvasir-seg-aliyun_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class KvasirSEGAliyunDataset(BaseSegDataset):
+ """KvasirSEGAliyunDataset dataset.
+
+ In segmentation map annotation for KvasirSEGAliyunDataset,
+ 0 stands for background,which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'polyp'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/tools/prepare_dataset.py b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/tools/prepare_dataset.py
new file mode 100644
index 00000000000..b230e7fef58
--- /dev/null
+++ b/projects/medical/2d_image/endoscopy/kvasir_seg_aliyun/tools/prepare_dataset.py
@@ -0,0 +1,86 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '.jpg'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path).convert('L'))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+ convert_pics_into_pngs(
+ os.path.join(root_path, 'Kvasir-SEG/images'),
+ tgt_img_dir,
+ suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ os.path.join(root_path, 'Kvasir-SEG/masks'),
+ tgt_mask_dir,
+ suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/README.md b/projects/medical/2d_image/fluorescein_angriogram/vampire/README.md
new file mode 100644
index 00000000000..c2c61c46a0b
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/README.md
@@ -0,0 +1,158 @@
+# Vessel Assessment and Measurement Platform for Images of the REtina
+
+## Description
+
+This project support **`Vessel Assessment and Measurement Platform for Images of the REtina`**, and the dataset used in this project can be downloaded from [here](https://vampire.computing.dundee.ac.uk/vesselseg.html).
+
+### Dataset Overview
+
+In order to promote evaluation of vessel segmentation on ultra-wide field-of-view (UWFV) fluorescein angriogram (FA) frames, we make public 8 frames from two different sequences, the manually annotated images and the result of our automatic vessel segmentation algorithm.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------- | ----------------- | ------------ | ---------------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Vampire](https://vampire.computing.dundee.ac.uk/vesselseg.html) | vessel | segmentation | fluorescein angriogram | 2 | 8/-/- | yes/-/- | 2017 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 8 | 96.75 | - | - | - | - |
+| vessel | 8 | 3.25 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```bibtex
+
+@inproceedings{perez2011improving,
+ title={Improving vessel segmentation in ultra-wide field-of-view retinal fluorescein angiograms},
+ author={Perez-Rovira, Adria and Zutis, K and Hubschman, Jean Pierre and Trucco, Emanuele},
+ booktitle={2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society},
+ pages={2614--2617},
+ year={2011},
+ organization={IEEE}
+}
+
+@article{perez2011rerbee,
+ title={RERBEE: robust efficient registration via bifurcations and elongated elements applied to retinal fluorescein angiogram sequences},
+ author={Perez-Rovira, Adria and Cabido, Raul and Trucco, Emanuele and McKenna, Stephen J and Hubschman, Jean Pierre},
+ journal={IEEE Transactions on Medical Imaging},
+ volume={31},
+ number={1},
+ pages={140--150},
+ year={2011},
+ publisher={IEEE}
+}
+
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `vampire/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://vampire.computing.dundee.ac.uk/vesselseg.html) and decompression data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `python ../../tools/split_seg_dataset.py` to split dataset. For the Bacteria_detection dataset, as there is no test or validation dataset, we sample 20% samples from the whole dataset as the validation dataset and 80% samples for training data and make two filename lists `train.txt` and `val.txt`. As we set the random seed as the hard code, we eliminated the randomness, the dataset split actually can be reproducible.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fluorescein_angriogram
+ │ │ │ │ ├── vampire
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 6 | 97.48 | 2 | 94.54 | - | - |
+| vessel | 6 | 2.52 | 2 | 5.46 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_vampire-512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_vampire-512x512.py
new file mode 100755
index 00000000000..7f5273aaff9
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_vampire-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './vampire_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.vampire_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_vampire-512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_vampire-512x512.py
new file mode 100755
index 00000000000..4382229989b
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_vampire-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './vampire_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.vampire_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=dict(size=img_scale),
+ pretrained=None,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_vampire-512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_vampire-512x512.py
new file mode 100755
index 00000000000..8d93e17627f
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_vampire-512x512.py
@@ -0,0 +1,22 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './vampire_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.vampire_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(
+ num_classes=2,
+ loss_decode=dict(type='CrossEntropyLoss', use_sigmoid=True),
+ out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/vampire_512x512.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/vampire_512x512.py
new file mode 100755
index 00000000000..4eda92f9f22
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/configs/vampire_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'VampireDataset'
+data_root = 'data'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/__init__.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/__init__.py
new file mode 100755
index 00000000000..93f9cbf0506
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/__init__.py
@@ -0,0 +1,3 @@
+from .vampire_dataset import VampireDataset
+
+__all__ = ['VampireDataset']
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/vampire_dataset.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/vampire_dataset.py
new file mode 100755
index 00000000000..4d38040f7f1
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/datasets/vampire_dataset.py
@@ -0,0 +1,28 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class VampireDataset(BaseSegDataset):
+ """VampireDataset dataset.
+
+ In segmentation map annotation for VampireDataset, 0 stands for background,
+ which is included in 2 categories. ``reduce_zero_label`` is fixed to
+ False. The ``img_suffix`` is fixed to '.png' and ``seg_map_suffix`` is
+ fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/fluorescein_angriogram/vampire/tools/prepare_dataset.py b/projects/medical/2d_image/fluorescein_angriogram/vampire/tools/prepare_dataset.py
new file mode 100644
index 00000000000..2755b5d28bd
--- /dev/null
+++ b/projects/medical/2d_image/fluorescein_angriogram/vampire/tools/prepare_dataset.py
@@ -0,0 +1,44 @@
+import os
+import shutil
+
+from PIL import Image
+
+path = 'data'
+
+if not os.path.exists(os.path.join(path, 'images', 'train')):
+ os.system(f'mkdir -p {os.path.join(path, "images", "train")}')
+
+if not os.path.exists(os.path.join(path, 'masks', 'train')):
+ os.system(f'mkdir -p {os.path.join(path, "masks", "train")}')
+
+origin_data_path = os.path.join(path, 'vesselSegmentation')
+
+imgs_amd14 = os.listdir(os.path.join(origin_data_path, 'AMD14'))
+imgs_ger7 = os.listdir(os.path.join(origin_data_path, 'GER7'))
+
+for img in imgs_amd14:
+ shutil.copy(
+ os.path.join(origin_data_path, 'AMD14', img),
+ os.path.join(path, 'images', 'train', img))
+ # copy GT
+ img_gt = img.replace('.png', '-GT.png')
+ shutil.copy(
+ os.path.join(origin_data_path, 'AMD14-GT', f'{img_gt}'),
+ os.path.join(path, 'masks', 'train', img))
+
+for img in imgs_ger7:
+ shutil.copy(
+ os.path.join(origin_data_path, 'GER7', img),
+ os.path.join(path, 'images', 'train', img))
+ # copy GT
+ img_gt = img.replace('.bmp', '-GT.png')
+ img = img.replace('bmp', 'png')
+ shutil.copy(
+ os.path.join(origin_data_path, 'GER7-GT', img_gt),
+ os.path.join(path, 'masks', 'train', img))
+
+imgs = os.listdir(os.path.join(path, 'images', 'train'))
+for img in imgs:
+ if not img.endswith('.png'):
+ im = Image.open(os.path.join(path, 'images', 'train', img))
+ im.save(os.path.join(path, 'images', 'train', img[:-4] + '.png'))
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/README.md b/projects/medical/2d_image/fundus_photography/dr_hagis/README.md
new file mode 100644
index 00000000000..85d8a3e271c
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/README.md
@@ -0,0 +1,155 @@
+# DR HAGIS: Diabetic Retinopathy, Hypertension, Age-related macular degeneration and Glacuoma ImageS
+
+## Description
+
+This project supports **`DR HAGIS: Diabetic Retinopathy, Hypertension, Age-related macular degeneration and Glacuoma ImageS`**, which can be downloaded from [here](https://paperswithcode.com/dataset/dr-hagis).
+
+### Dataset Overview
+
+The DR HAGIS database has been created to aid the development of vessel extraction algorithms suitable for retinal screening programmes. Researchers are encouraged to test their segmentation algorithms using this database. All thirty-nine fundus images were obtained from a diabetic retinopathy screening programme in the UK. Hence, all images were taken from diabetic patients.
+
+Besides the fundus images, the manual segmentation of the retinal surface vessels is provided by an expert grader. These manually segmented images can be used as the ground truth to compare and assess the automatic vessel extraction algorithms. Masks of the FOV are provided as well to quantify the accuracy of vessel extraction within the FOV only. The images were acquired in different screening centers, therefore reflecting the range of image resolutions, digital cameras and fundus cameras used in the clinic. The fundus images were captured using a Topcon TRC-NW6s, Topcon TRC-NW8 or a Canon CR DGi fundus camera with a horizontal 45 degree field-of-view (FOV). The images are 4752x3168 pixels, 3456x2304 pixels, 3126x2136 pixels, 2896x1944 pixels or 2816x1880 pixels in size.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------- | ----------------- | ------------ | ------------------ | ------------ | --------------------- | ---------------------- | ------------ | ------- |
+| [DR HAGIS](https://paperswithcode.com/dataset/dr-hagis) | head and neck | segmentation | fundus photography | 2 | 40/-/- | yes/-/- | 2017 | - |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 40 | 96.38 | - | - | - | - |
+| vessel | 40 | 3.62 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Usage
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `dr_hagis/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://paperswithcode.com/dataset/dr-hagis) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── dr_hagis
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 32 | 96.21 | 8 | 97.12 | - | - |
+| vessel | 32 | 3.79 | 8 | 2.88 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@article{holm2017dr,
+ title={DR HAGIS—a fundus image database for the automatic extraction of retinal surface vessels from diabetic patients},
+ author={Holm, Sven and Russell, Greg and Nourrit, Vincent and McLoughlin, Niall},
+ journal={Journal of Medical Imaging},
+ volume={4},
+ number={1},
+ pages={014503--014503},
+ year={2017},
+ publisher={Society of Photo-Optical Instrumentation Engineers}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/dr-hagis_512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/dr-hagis_512x512.py
new file mode 100644
index 00000000000..93b96384109
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/dr-hagis_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'DRHAGISDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_dr-hagis-512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_dr-hagis-512x512.py
new file mode 100644
index 00000000000..9d14427c45f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_dr-hagis-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './dr-hagis_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.dr-hagis_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_dr-hagis-512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_dr-hagis-512x512.py
new file mode 100644
index 00000000000..507ec748bf5
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_dr-hagis-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './dr-hagis_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.dr-hagis_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_dr-hagis-512x512.py b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_dr-hagis-512x512.py
new file mode 100644
index 00000000000..092ae00a7d3
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_dr-hagis-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './dr-hagis_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.dr-hagis_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/datasets/dr-hagis_dataset.py b/projects/medical/2d_image/fundus_photography/dr_hagis/datasets/dr-hagis_dataset.py
new file mode 100644
index 00000000000..9659f0b8d77
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/datasets/dr-hagis_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class DRHAGISDataset(BaseSegDataset):
+ """DRHAGISDataset dataset.
+
+ In segmentation map annotation for DRHAGISDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/dr_hagis/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/dr_hagis/tools/prepare_dataset.py
new file mode 100755
index 00000000000..51f4df7dac2
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/dr_hagis/tools/prepare_dataset.py
@@ -0,0 +1,41 @@
+import glob
+import os
+import shutil
+
+import mmengine
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '_manual_orig.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(os.path.join('data/DRHAGIS/**/*' + img_suffix))
+
+mmengine.mkdir_or_exist(root_path + 'images/train/')
+mmengine.mkdir_or_exist(root_path + 'masks/train/')
+
+D3_palette = {0: (0, 0, 0), 1: (1, 1, 1)}
+D3_invert_palette = {v: k for k, v in D3_palette.items()}
+D2_255_convert_dict = {0: 0, 255: 1}
+
+part_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ shutil.copy(
+ img, root_path + 'images/' + part_dir + basename.split('.')[0] +
+ save_img_suffix)
+ mask_path = root_path + 'DRHAGIS/Manual_Segmentations/' + basename.split( # noqa
+ '.')[0] + seg_map_suffix
+ label = np.array(Image.open(mask_path))
+
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix # noqa
+ mask = np.array(Image.open(mask_path)).astype(np.uint8)
+ mask[mask == 255] = 1
+ mask = Image.fromarray(mask)
+ mask.save(save_mask_path)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/README.md b/projects/medical/2d_image/fundus_photography/gamma3/README.md
new file mode 100644
index 00000000000..e834508fcb5
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/README.md
@@ -0,0 +1,167 @@
+# Glaucoma grAding from Multi-Modality imAges Task3
+
+## Description
+
+This project support **`Glaucoma grAding from Multi-Modality imAges Task3`**, and the dataset used in this project can be downloaded from [here](https://aistudio.baidu.com/aistudio/competition/detail/121/0/datasets).
+
+### Dataset Overview
+
+This regular-challenge dataset was provided by Sun Yat-sen Ophthalmic Center, Sun Yat-sen University, Guangzhou, China. The dataset contains 200 fundus color images: 100 pairs in the training set and 100 pairs in the test set.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ----------------------------------------------------------------------------------- | ----------------- | ------------ | --------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [GammaTask3](https://aistudio.baidu.com/aistudio/competition/detail/121/0/datasets) | eye | segmentation | fundus photophy | 3 | 100/-/100 | yes/-/- | 2021 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 100 | 99.02 | - | - | - | - |
+| optic disc | 100 | 0.67 | - | - | - | - |
+| optic cup | 100 | 0.31 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```bibtex
+@article{fu2018joint,
+ title={Joint optic disc and cup segmentation based on multi-label deep network and polar transformation},
+ author={Fu, Huazhu and Cheng, Jun and Xu, Yanwu and Wong, Damon Wing Kee and Liu, Jiang and Cao, Xiaochun},
+ journal={IEEE transactions on medical imaging},
+ volume={37},
+ number={7},
+ pages={1597--1605},
+ year={2018},
+ publisher={IEEE}
+}
+
+@article{sevastopolsky2017optic,
+ title={Optic disc and cup segmentation methods for glaucoma detection with modification of U-Net convolutional neural network},
+ author={Sevastopolsky, Artem},
+ journal={Pattern Recognition and Image Analysis},
+ volume={27},
+ pages={618--624},
+ year={2017},
+ publisher={Springer}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `gammm3/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://aistudio.baidu.com/aistudio/competition/detail/121/0/datasets) and decompression data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── gamma3
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 80 | 99.01 | 20 | 99.07 | - | - |
+| optic disc | 80 | 0.68 | 20 | 0.63 | - | - |
+| optic cup | 80 | 0.32 | 20 | 0.31 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_gamma3-512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_gamma3-512x512.py
new file mode 100644
index 00000000000..0daac51e10f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_gamma3-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './gamma3_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.gamma3_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_gamma3-512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_gamma3-512x512.py
new file mode 100644
index 00000000000..8a25cd0d266
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_gamma3-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './gamma3_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.gamma3_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_gamma3-512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_gamma3-512x512.py
new file mode 100644
index 00000000000..ea648438672
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_gamma3-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './gamma3_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.gamma3_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/configs/gamma3_512x512.py b/projects/medical/2d_image/fundus_photography/gamma3/configs/gamma3_512x512.py
new file mode 100644
index 00000000000..d23ab55ca71
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/configs/gamma3_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'Gamma3Dataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/datasets/gamma3_dataset.py b/projects/medical/2d_image/fundus_photography/gamma3/datasets/gamma3_dataset.py
new file mode 100644
index 00000000000..56cbdd63e61
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/datasets/gamma3_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class Gamma3Dataset(BaseSegDataset):
+ """Gamma3Dataset dataset.
+
+ In segmentation map annotation for Gamma3Dataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'disc', 'cup'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/gamma3/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/gamma3/tools/prepare_dataset.py
new file mode 100644
index 00000000000..eb820b6b740
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/gamma3/tools/prepare_dataset.py
@@ -0,0 +1,107 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**/*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 2,
+ 128: 1,
+ 255: 0
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ os.path.join(
+ root_path,
+ 'task3_disc_cup_segmentation/training/fundus color images/'),
+ tgt_img_train_dir,
+ suffix=img_suffix)
+
+ convert_pics_into_pngs(
+ os.path.join(
+ root_path,
+ 'task3_disc_cup_segmentation/testing/fundus color images/'),
+ tgt_img_test_dir,
+ suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ os.path.join(root_path,
+ 'task3_disc_cup_segmentation/training/Disc_Cup_Mask/'),
+ tgt_mask_train_dir,
+ suffix=seg_map_suffix,
+ convert_dict={
+ 0: 2,
+ 128: 1,
+ 255: 0
+ })
+ # original: [0, 128, 255] for ['optic cup', 'optic disc', 'background']
+ # converted: [0, 1, 2] for ['background', 'optic disc', 'optic cup']
diff --git a/projects/medical/2d_image/fundus_photography/orvs/README.md b/projects/medical/2d_image/fundus_photography/orvs/README.md
new file mode 100644
index 00000000000..6f09203ac4d
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/README.md
@@ -0,0 +1,140 @@
+# ORVS (Online Retinal image for Vessel Segmentation (ORVS))
+
+## Description
+
+This project supports **`ORVS (Online Retinal image for Vessel Segmentation (ORVS))`**, which can be downloaded from [here](https://opendatalab.org.cn/ORVS).
+
+### Dataset Overview
+
+The ORVS dataset is a newly established collaboration between the Department of Computer Science and the Department of Vision Science at the University of Calgary. The dataset contains 49 images collected from a clinic in Calgary, Canada, consisting of 42 training images and 7 testing images. All images were obtained using a Zeiss Visucam 200 with a 30-degree field of view (FOV). The image size is 1444×1444 pixels with 24 bits per pixel. The images are stored in JPEG format with low compression, which is common in ophthalmic practice. All images were manually traced by an expert who has been working in the field of retinal image analysis and has been trained to mark all pixels belonging to retinal vessels. The Windows Paint 3D tool was used for manual image annotation.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------ | ----------------- | ------------ | ------------------ | ------------ | --------------------- | ---------------------- | ------------ | ------- |
+| [Bactteria detection](https://opendatalab.org.cn/ORVS) | bacteria | segmentation | fundus photography | 2 | 130/-/72 | yes/-/yes | 2020 | - |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 130 | 94.83 | - | - | 72 | 94.25 |
+| vessel | 130 | 5.17 | - | - | 72 | 5.75 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `orvs/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- Clone this [repository](https://github.com/AbdullahSarhan/ICPRVessels), then move `Vessels-Datasets` to `data/`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── orvs
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@inproceedings{sarhan2021transfer,
+ title={Transfer learning through weighted loss function and group normalization for vessel segmentation from retinal images},
+ author={Sarhan, Abdullah and Rokne, Jon and Alhajj, Reda and Crichton, Andrew},
+ booktitle={2020 25th International Conference on Pattern Recognition (ICPR)},
+ pages={9211--9218},
+ year={2021},
+ organization={IEEE}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [ ] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_orvs-512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_orvs-512x512.py
new file mode 100644
index 00000000000..662f837158a
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_orvs-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './orvs_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.orvs_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_orvs-512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_orvs-512x512.py
new file mode 100644
index 00000000000..c47cdb6b24d
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_orvs-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './orvs_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.orvs_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_orvs-512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_orvs-512x512.py
new file mode 100644
index 00000000000..1097aade286
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_orvs-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './orvs_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.orvs_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/configs/orvs_512x512.py b/projects/medical/2d_image/fundus_photography/orvs/configs/orvs_512x512.py
new file mode 100644
index 00000000000..a5594dec388
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/configs/orvs_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ORVSDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='test.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/orvs/datasets/orvs_dataset.py b/projects/medical/2d_image/fundus_photography/orvs/datasets/orvs_dataset.py
new file mode 100644
index 00000000000..e915ae4cd2b
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/datasets/orvs_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ORVSDataset(BaseSegDataset):
+ """ORVSDataset dataset.
+
+ In segmentation map annotation for ORVSDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/orvs/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/orvs/tools/prepare_dataset.py
new file mode 100755
index 00000000000..f902d871010
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/orvs/tools/prepare_dataset.py
@@ -0,0 +1,55 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.jpg'
+seg_map_suffix_list = ['.jpg', '.png', '.tif']
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(
+ os.path.join('data/Vessels-Datasets/*/Train/Original/Images/*' +
+ img_suffix))
+x_test = glob.glob(
+ os.path.join('data/Vessels-Datasets/*/Test/Original/Images/*' +
+ img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/test/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+os.system('mkdir -p ' + root_path + 'masks/test/')
+
+part_dir_dict = {0: 'train/', 1: 'test/'}
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ type_name = img.split('/')[-5]
+ basename = type_name + '_' + os.path.basename(img)
+ save_img_path = root_path + 'images/' + part_dir + basename.split(
+ '.')[0] + save_img_suffix
+ Image.open(img).save(save_img_path)
+
+ for seg_map_suffix in seg_map_suffix_list:
+ if os.path.exists('/'.join(img.split('/')[:-1]).replace(
+ 'Images', 'Labels')):
+ mask_path = img.replace('Images', 'Labels').replace(
+ img_suffix, seg_map_suffix)
+ else:
+ mask_path = img.replace('Images', 'labels').replace(
+ img_suffix, seg_map_suffix)
+ if os.path.exists(mask_path):
+ break
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix
+ masks = np.array(Image.open(mask_path).convert('L')).astype(np.uint8)
+ if len(np.unique(masks)) == 2 and 1 in np.unique(masks):
+ print(np.unique(masks))
+ pass
+ else:
+ masks[masks < 128] = 0
+ masks[masks >= 128] = 1
+ masks = Image.fromarray(masks)
+ masks.save(save_mask_path)
diff --git a/projects/medical/2d_image/fundus_photography/rite/README.md b/projects/medical/2d_image/fundus_photography/rite/README.md
new file mode 100644
index 00000000000..0aea9b00d17
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/README.md
@@ -0,0 +1,135 @@
+# Retinal Images vessel Tree Extraction (RITE)
+
+## Description
+
+This project supports **`Retinal Images vessel Tree Extraction (RITE) `**, which can be downloaded from [here](https://opendatalab.com/RITE).
+
+### Dataset Overview
+
+The RITE (Retinal Images vessel Tree Extraction) is a database that enables comparative studies on segmentation or classification of arteries and veins on retinal fundus images, which is established based on the public available DRIVE database (Digital Retinal Images for Vessel Extraction). RITE contains 40 sets of images, equally separated into a training subset and a test subset, the same as DRIVE. The two subsets are built from the corresponding two subsets in DRIVE. For each set, there is a fundus photograph, a vessel reference standard. The fundus photograph is inherited from DRIVE. For the training set, the vessel reference standard is a modified version of 1st_manual from DRIVE. For the test set, the vessel reference standard is 2nd_manual from DRIVE.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------ | ----------------- | ------------ | ------------------ | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Rite](https://opendatalab.com/RITE) | head_and_neck | segmentation | fundus_photography | 2 | 20/-/20 | yes/-/yes | 2013 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 20 | 91.61 | - | - | 20 | 91.58 |
+| vessel | 20 | 8.39 | - | - | 20 | 8.42 |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@InProceedings{10.1007/978-3-642-40763-5_54,
+ author={Hu, Qiao and Abr{\`a}moff, Michael D. and Garvin, Mona K.},
+ title={Automated Separation of Binary Overlapping Trees in Low-Contrast Color Retinal Images},
+ booktitle={Medical Image Computing and Computer-Assisted Intervention -- MICCAI 2013},
+ year={2013},
+ pages={436--443},
+}
+
+
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `rite/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.com/RITE) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── fundus_photography
+ │ │ │ │ ├── rite
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_rite-512x512.py
new file mode 100644
index 00000000000..27dd4363b16
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_rite-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_rite-512x512.py
new file mode 100644
index 00000000000..48f6f973a1a
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_rite-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_rite-512x512.py
new file mode 100644
index 00000000000..5f5b24ba6a4
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_rite-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_rite-512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_rite-512x512.py
new file mode 100644
index 00000000000..bf66b6f320c
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_rite-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './rite_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.rite_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/fundus_photography/rite/configs/rite_512x512.py b/projects/medical/2d_image/fundus_photography/rite/configs/rite_512x512.py
new file mode 100644
index 00000000000..02f620c665f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/configs/rite_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'RITEDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='test.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/fundus_photography/rite/datasets/rite_dataset.py b/projects/medical/2d_image/fundus_photography/rite/datasets/rite_dataset.py
new file mode 100644
index 00000000000..99f688de949
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/datasets/rite_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class RITEDataset(BaseSegDataset):
+ """RITEDataset dataset.
+
+ In segmentation map annotation for RITEDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/fundus_photography/rite/tools/prepare_dataset.py b/projects/medical/2d_image/fundus_photography/rite/tools/prepare_dataset.py
new file mode 100644
index 00000000000..ca7e996961f
--- /dev/null
+++ b/projects/medical/2d_image/fundus_photography/rite/tools/prepare_dataset.py
@@ -0,0 +1,98 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.tif'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+src_img_train_dir = os.path.join(root_path, 'AV_groundTruth/training/images/')
+src_img_test_dir = os.path.join(root_path, 'AV_groundTruth/test/images/')
+src_mask_train_dir = os.path.join(root_path, 'AV_groundTruth/training/vessel/')
+src_mask_test_dir = os.path.join(root_path, 'AV_groundTruth/test/vessel/')
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+tgt_mask_test_dir = os.path.join(root_path, 'masks/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+os.system('mkdir -p ' + tgt_mask_test_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ src_img_train_dir, tgt_img_train_dir, suffix=img_suffix)
+
+ convert_pics_into_pngs(
+ src_img_test_dir, tgt_img_test_dir, suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_train_dir, tgt_mask_train_dir, suffix=seg_map_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_test_dir, tgt_mask_test_dir, suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/README.md b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/README.md
new file mode 100644
index 00000000000..97c4a0f0e55
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/README.md
@@ -0,0 +1,123 @@
+# breastCancerCellSegmentation
+
+## Description
+
+This project supports **`breastCancerCellSegmentation`**, which can be downloaded from [here](https://www.heywhale.com/mw/dataset/5e9e9b35ebb37f002c625423).
+
+### Dataset Overview
+
+This dataset, with 58 H&E-stained histopathology images was used for breast cancer cell detection and associated real-world data.
+Conventional histology uses a combination of hematoxylin and eosin stains, commonly referred to as H&E. These images are stained because most cells are inherently transparent with little or no intrinsic pigment.
+Certain special stains selectively bind to specific components and can be used to identify biological structures such as cells.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| -------------------------------------------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [breastCancerCellSegmentation](https://www.heywhale.com/mw/dataset/5e9e9b35ebb37f002c625423) | cell | segmentation | histopathology | 2 | 58/-/- | yes/-/- | 2020 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 58 | 98.37 | - | - | - | - |
+| breastCancerCell | 58 | 1.63 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Usage
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow (PIL) v9.3.0
+- scikit-learn (sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `breastCancerCellSegmentation/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- Download dataset from [here](https://www.heywhale.com/mw/dataset/5e9e9b35ebb37f002c625423) and save it to the `data/` directory .
+- Decompress data to path `data/`. This will create a new folder named `data/breastCancerCellSegmentation/`, which contains the original image data.
+- run script `python tools/prepare_dataset.py` to format data and change folder structure as below.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── breastCancerCellSegmentation
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── breastCancerCellSegmentation
+ | │ │ │ │ │ │ ├── train.txt
+ | │ │ │ │ │ │ ├── val.txt
+ | │ │ │ │ │ │ ├── images
+ | │ │ │ │ │ │ | ├── xxx.tif
+ | │ │ │ │ │ │ ├── masks
+ | │ │ │ │ │ │ | ├── xxx.TIF
+
+```
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/breastCancerCellSegmentation_512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/breastCancerCellSegmentation_512x512.py
new file mode 100644
index 00000000000..1cf0fccf5be
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/breastCancerCellSegmentation_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'breastCancerCellSegmentationDataset'
+data_root = 'data/breastCancerCellSegmentation'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile', imdecode_backend='tifffile'),
+ dict(type='LoadAnnotations', imdecode_backend='tifffile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile', imdecode_backend='tifffile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations', imdecode_backend='tifffile'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images', seg_map_path='masks'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images', seg_map_path='masks'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breastCancerCellSegmentation-512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breastCancerCellSegmentation-512x512.py
new file mode 100644
index 00000000000..55d17089686
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breastCancerCellSegmentation-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './breastCancerCellSegmentation_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breastCancerCellSegmentation_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breastCancerCellSegmentation-512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breastCancerCellSegmentation-512x512.py
new file mode 100644
index 00000000000..cf28aad739f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breastCancerCellSegmentation-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './breastCancerCellSegmentation_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breastCancerCellSegmentation_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breastCancerCellSegmentation-512x512.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breastCancerCellSegmentation-512x512.py
new file mode 100644
index 00000000000..29aaff38941
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breastCancerCellSegmentation-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './breastCancerCellSegmentation_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breastCancerCellSegmentation_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/datasets/breastCancerCellSegmentation_dataset.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/datasets/breastCancerCellSegmentation_dataset.py
new file mode 100644
index 00000000000..eeceb6318c0
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/datasets/breastCancerCellSegmentation_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class breastCancerCellSegmentationDataset(BaseSegDataset):
+ """breastCancerCellSegmentationDataset dataset.
+
+ In segmentation map annotation for breastCancerCellSegmentationDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'breastCancerCell'))
+
+ def __init__(self,
+ img_suffix='_ccd.tif',
+ seg_map_suffix='.TIF',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/tools/prepare_dataset.py
new file mode 100644
index 00000000000..09cc689c862
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breastCancerCellSegmentation/tools/prepare_dataset.py
@@ -0,0 +1,36 @@
+import argparse
+import glob
+import os
+
+from sklearn.model_selection import train_test_split
+
+
+def save_anno(img_list, file_path, suffix):
+ # 只保留文件名,不保留后缀
+ img_list = [x.split('/')[-1][:-len(suffix)] for x in img_list]
+
+ with open(file_path, 'w') as file_:
+ for x in list(img_list):
+ file_.write(x + '\n')
+
+
+if __name__ == '__main__':
+ parser = argparse.ArgumentParser()
+ parser.add_argument(
+ '--data_root', default='data/breastCancerCellSegmentation/')
+ args = parser.parse_args()
+ data_root = args.data_root
+
+ # 1. 划分训练集、验证集
+ # 1.1 获取所有图片路径
+ img_list = glob.glob(os.path.join(data_root, 'images', '*.tif'))
+ img_list.sort()
+ mask_list = glob.glob(os.path.join(data_root, 'masks', '*.TIF'))
+ mask_list.sort()
+ assert len(img_list) == len(mask_list)
+ # 1.2 划分训练集、验证集、测试集
+ train_img_list, val_img_list, train_mask_list, val_mask_list = train_test_split( # noqa
+ img_list, mask_list, test_size=0.2, random_state=42)
+ # 1.3 保存划分结果
+ save_anno(train_img_list, os.path.join(data_root, 'train.txt'), '_ccd.tif')
+ save_anno(val_img_list, os.path.join(data_root, 'val.txt'), '_ccd.tif')
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/README.md b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/README.md
new file mode 100644
index 00000000000..b6f1ca63419
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/README.md
@@ -0,0 +1,147 @@
+# Breast Cancer Cell Segmentation
+
+## Description
+
+This project support **`Breast Cancer Cell Segmentation`**, and the dataset used in this project can be downloaded from [here](https://tianchi.aliyun.com/dataset/dataDetail?dataId=90152).
+
+### Dataset Overview
+
+In this dataset, there are 58 H&E stained histopathology images used in breast cancer cell detection with associated ground truth data available. Routine histology uses the stain combination of hematoxylin and eosin, commonly referred to as H&E. These images are stained since most cells are essentially transparent, with little or no intrinsic pigment. Certain special stains, which bind selectively to particular components, are be used to identify biological structures such as cells. In those images, the challenging problem is cell segmentation for subsequent classification in benign and malignant cells.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------------------------------------------------ |
+| [Breast Cancer Cell Segmentation](https://tianchi.aliyun.com/dataset/dataDetail?dataId=90152) | thorax | segmentation | histopathology | 2 | 58/-/- | yes/-/- | 2021 | [CC-BY-SA-NC 4.0](http://creativecommons.org/licenses/by-sa/4.0/?spm=5176.12282016.0.0.3f5b5291ypBxb2) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 58 | 98.37 | - | - | - | - |
+| breast cancer cell | 58 | 1.63 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```
+@inproceedings{gelasca2008evaluation,
+ title={Evaluation and benchmark for biological image segmentation},
+ author={Gelasca, Elisa Drelie and Byun, Jiyun and Obara, Boguslaw and Manjunath, BS},
+ booktitle={2008 15th IEEE international conference on image processing},
+ pages={1816--1819},
+ year={2008},
+ organization={IEEE}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `breast_cancer_cell_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://tianchi.aliyun.com/dataset/dataDetail?dataId=90152) and decompression data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── breast_cancer_cell_seg
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 46 | 98.36 | 12 | 98.41 | - | - |
+| erythrocytes | 46 | 1.64 | 12 | 1.59 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/breast-cancer-cell-seg_512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/breast-cancer-cell-seg_512x512.py
new file mode 100644
index 00000000000..ead40e4345c
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/breast-cancer-cell-seg_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'BreastCancerCellSegDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breast-cancer-cell-seg-512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breast-cancer-cell-seg-512x512.py
new file mode 100644
index 00000000000..691a0ff613d
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_breast-cancer-cell-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './breast-cancer-cell-seg_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breast-cancer-cell-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breast-cancer-cell-seg-512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breast-cancer-cell-seg-512x512.py
new file mode 100644
index 00000000000..719b767ab1f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_breast-cancer-cell-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './breast-cancer-cell-seg_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breast-cancer-cell-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breast-cancer-cell-seg-512x512.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breast-cancer-cell-seg-512x512.py
new file mode 100644
index 00000000000..9dfe70f761f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_breast-cancer-cell-seg-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './breast-cancer-cell-seg_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.breast-cancer-cell-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/datasets/breast-cancer-cell-seg_dataset.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/datasets/breast-cancer-cell-seg_dataset.py
new file mode 100644
index 00000000000..6f27029d396
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/datasets/breast-cancer-cell-seg_dataset.py
@@ -0,0 +1,29 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class BreastCancerCellSegDataset(BaseSegDataset):
+ """BreastCancerCellSegDataset dataset.
+
+ In segmentation map annotation for BreastCancerCellSegDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('normal', 'breast cancer cell'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/tools/prepare_dataset.py
new file mode 100755
index 00000000000..775f2eed18a
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/breast_cancer_cell_seg/tools/prepare_dataset.py
@@ -0,0 +1,47 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.tif'
+seg_map_suffix = '.TIF'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(
+ os.path.join('data/Breast Cancer Cell Segmentation_datasets/Images/*' +
+ img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+D2_255_convert_dict = {0: 0, 255: 1}
+
+
+def convert_2d(img, convert_dict=D2_255_convert_dict):
+ arr_2d = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr_2d[img == c] = i
+ return arr_2d
+
+
+part_dir_dict = {0: 'train/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_save_path = root_path + 'images/' + part_dir + basename.split(
+ '.')[0] + save_img_suffix
+ Image.open(img).save(img_save_path)
+ mask_path = root_path + 'Breast Cancer Cell Segmentation_datasets/Masks/' + '_'.join( # noqa
+ basename.split('_')[:-1]) + seg_map_suffix
+ label = np.array(Image.open(mask_path))
+
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix
+ assert len(label.shape) == 2 and 255 in label and 1 not in label
+ mask = convert_2d(label)
+ mask = Image.fromarray(mask.astype(np.uint8))
+ mask.save(save_mask_path)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/README.md b/projects/medical/2d_image/histopathology/conic2022_seg/README.md
new file mode 100644
index 00000000000..1f55b44ed6d
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/README.md
@@ -0,0 +1,207 @@
+# CoNIC: Colon Nuclei Identification and Counting Challenge
+
+## Description
+
+This project supports **`CoNIC: Colon Nuclei Identification and Counting Challenge`**, which can be downloaded from [here](https://drive.google.com/drive/folders/1il9jG7uA4-ebQ_lNmXbbF2eOK9uNwheb).
+
+### Dataset Overview
+
+Nuclear segmentation, classification and quantification within Haematoxylin & Eosin stained histology images enables the extraction of interpretable cell-based features that can be used in downstream explainable models in computational pathology (CPath). To help drive forward research and innovation for automatic nuclei recognition in CPath, we organise the Colon Nuclei Identification and Counting (CoNIC) Challenge. The challenge requires researchers to develop algorithms that perform segmentation, classification and counting of 6 different types of nuclei within the current largest known publicly available nuclei-level dataset in CPath, containing around half a million labelled nuclei.
+
+### Task Information
+
+The CONIC challenge has 2 tasks:
+
+- Task 1: Nuclear segmentation and classification.
+
+The first task requires participants to segment nuclei within the tissue, while also classifying each nucleus into one of the following categories: epithelial, lymphocyte, plasma, eosinophil, neutrophil or connective tissue.
+
+- Task 2: Prediction of cellular composition.
+
+For the second task, we ask participants to predict how many nuclei of each class are present in each input image.
+
+The output of Task 1 can be directly used to perform Task 2, but these can be treated as independent tasks. Therefore, if it is preferred, prediction of cellular composition can be treated as a stand alone regression task.
+
+***NOTE:We only consider `Task 1` in the following sections.***
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| -------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------------------------------------------------------ |
+| [CoNIC202](https://conic-challenge.grand-challenge.org/) | abdomen | segmentation | histopathology | 7 | 4981/-/- | yes/-/- | 2022 | [Attribution-NonCommercial-ShareAlike 4.0 International](https://creativecommons.org/licenses/by-nc-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 4981 | 83.97 | - | - | - | - |
+| neutrophil | 1218 | 0.13 | - | - | - | - |
+| epithelial | 4256 | 10.31 | - | - | - | - |
+| lymphocyte | 4473 | 1.85 | - | - | - | - |
+| plasma | 3316 | 0.55 | - | - | - | - |
+| eosinophil | 1456 | 0.1 | - | - | - | - |
+| connective | 4613 | 3.08 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `conic2022_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://drive.google.com/drive/folders/1il9jG7uA4-ebQ_lNmXbbF2eOK9uNwheb/) and move data to path `'data/CoNIC_Challenge'`. The directory should be like:
+ ```shell
+ data/CoNIC_Challenge
+ ├── README.txt
+ ├── by-nc-sa.md
+ ├── counts.csv
+ ├── images.npy
+ ├── labels.npy
+ └── patch_info.csv
+ ```
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── conic2022_seg
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 3984 | 84.06 | 997 | 83.65 | - | - |
+| neutrophil | 956 | 0.12 | 262 | 0.13 | - | - |
+| epithelial | 3400 | 10.26 | 856 | 10.52 | - | - |
+| lymphocyte | 3567 | 1.83 | 906 | 1.96 | - | - |
+| plasma | 2645 | 0.55 | 671 | 0.56 | - | - |
+| eosinophil | 1154 | 0.1 | 302 | 0.1 | - | - |
+| connective | 3680 | 3.08 | 933 | 3.08 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Organizers
+
+- Simon Graham (TIA, PathLAKE)
+- Mostafa Jahanifar (TIA, PathLAKE)
+- Dang Vu (TIA)
+- Giorgos Hadjigeorghiou (TIA, PathLAKE)
+- Thomas Leech (TIA, PathLAKE)
+- David Snead (UHCW, PathLAKE)
+- Shan Raza (TIA, PathLAKE)
+- Fayyaz Minhas (TIA, PathLAKE)
+- Nasir Rajpoot (TIA, PathLAKE)
+
+TIA: Tissue Image Analytics Centre, Department of Computer Science, University of Warwick, United Kingdom
+
+UHCW: Department of Pathology, University Hospitals Coventry and Warwickshire, United Kingdom
+
+PathLAKE: Pathology Image Data Lake for Analytics Knowledge & Education, , University Hospitals Coventry and Warwickshire, United Kingdom
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@inproceedings{graham2021lizard,
+ title={Lizard: A large-scale dataset for colonic nuclear instance segmentation and classification},
+ author={Graham, Simon and Jahanifar, Mostafa and Azam, Ayesha and Nimir, Mohammed and Tsang, Yee-Wah and Dodd, Katherine and Hero, Emily and Sahota, Harvir and Tank, Atisha and Benes, Ksenija and others},
+ booktitle={Proceedings of the IEEE/CVF International Conference on Computer Vision},
+ pages={684--693},
+ year={2021}
+}
+@article{graham2021conic,
+ title={Conic: Colon nuclei identification and counting challenge 2022},
+ author={Graham, Simon and Jahanifar, Mostafa and Vu, Quoc Dang and Hadjigeorghiou, Giorgos and Leech, Thomas and Snead, David and Raza, Shan E Ahmed and Minhas, Fayyaz and Rajpoot, Nasir},
+ journal={arXiv preprint arXiv:2111.14485},
+ year={2021}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/conic2022-seg_512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/conic2022-seg_512x512.py
new file mode 100644
index 00000000000..51b4e5782ab
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/conic2022-seg_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'Conic2022SegDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_conic2022-512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_conic2022-512x512.py
new file mode 100644
index 00000000000..3e0248c78ce
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_conic2022-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './conic2022-seg_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.conic2022-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=7),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_conic2022-512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_conic2022-512x512.py
new file mode 100644
index 00000000000..fd0e9d8d28b
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_conic2022-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './conic2022-seg_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.conic2022-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=7),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_conic2022-512x512.py b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_conic2022-512x512.py
new file mode 100644
index 00000000000..bb667f14fd4
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_conic2022-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './conic2022-seg_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.conic2022-seg_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=7),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/conic2022_seg_dataset.png b/projects/medical/2d_image/histopathology/conic2022_seg/conic2022_seg_dataset.png
new file mode 100644
index 00000000000..65bb0bbe0a5
Binary files /dev/null and b/projects/medical/2d_image/histopathology/conic2022_seg/conic2022_seg_dataset.png differ
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/datasets/conic2022-seg_dataset.py b/projects/medical/2d_image/histopathology/conic2022_seg/datasets/conic2022-seg_dataset.py
new file mode 100644
index 00000000000..9af0958ab34
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/datasets/conic2022-seg_dataset.py
@@ -0,0 +1,29 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class Conic2022SegDataset(BaseSegDataset):
+ """Conic2022SegDataset dataset.
+
+ In segmentation map annotation for Conic2022SegDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(
+ classes=('background', 'neutrophil', 'epithelial', 'lymphocyte',
+ 'plasma', 'eosinophil', 'connective'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/conic2022_seg/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/conic2022_seg/tools/prepare_dataset.py
new file mode 100755
index 00000000000..89cfb4aae24
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/conic2022_seg/tools/prepare_dataset.py
@@ -0,0 +1,65 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+
+img_save_root = 'data/'
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+label_set = set()
+
+
+def save_masks_from_npz(data, save_root, part='masks/'):
+ global label_set
+ num = data.shape[0]
+ for i in range(num):
+ # np_img = data[i, :, :, :]
+ np_mask = data[i, :, :, 1]
+ label_set = set.union(label_set, set(np.unique(np_mask)))
+ img = Image.fromarray(np_mask)
+ save_path = os.path.join(save_root, part, str(i) + save_seg_map_suffix)
+ img.save(save_path)
+
+
+def save_images_from_npz(data, save_root, part='images/'):
+ num = data.shape[0]
+ for i in range(num):
+ np_img = data[i, :, :, :]
+ img = Image.fromarray(np_img)
+ save_path = os.path.join(save_root, part, str(i) + save_img_suffix)
+ img.save(save_path)
+
+
+images_npy = np.load('data/CoNIC_Challenge/images.npy')
+labels_npy = np.load('data/CoNIC_Challenge/labels.npy')
+
+os.system('mkdir -p ' + img_save_root + 'images_ori')
+os.system('mkdir -p ' + img_save_root + 'labels')
+save_images_from_npz(images_npy, img_save_root, 'images_ori')
+save_masks_from_npz(labels_npy, img_save_root, 'labels')
+print(label_set)
+
+x_train = glob.glob(os.path.join('data/images_ori/*' + img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+part_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ shutil.copy(
+ img, root_path + 'images/' + part_dir + basename.split('.')[0] +
+ save_img_suffix)
+ mask_path = root_path + 'labels/' + basename.split(
+ '.')[0] + seg_map_suffix
+ save_mask_path = root_path + 'masks/' + part_dir + basename.split(
+ '.')[0] + save_seg_map_suffix
+ shutil.copy(mask_path, save_mask_path)
diff --git a/projects/medical/2d_image/histopathology/consep/README.md b/projects/medical/2d_image/histopathology/consep/README.md
new file mode 100644
index 00000000000..ca3d7aa1089
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/README.md
@@ -0,0 +1,147 @@
+# Colorectal Nuclear Segmentation and Phenotypes (CoNSeP) Dataset
+
+## Description
+
+This project supports **`Colorectal Nuclear Segmentation and Phenotypes (CoNSeP) Dataset`**, which can be downloaded from [here](https://warwick.ac.uk/fac/cross_fac/tia/data/hovernet/).
+
+### Dataset Overview
+
+The CoNSeP (Colon Segmentation and Phenotyping) dataset consists of 41 H&E stained image tiles, each with a size of 1,000×1,000 pixels and a magnification of 40x. These images were extracted from 16 colorectal adenocarcinoma (CRA) whole slide images (WSI), each of which belonged to a separate patient and was scanned using an Omnyx VL120 scanner at the Pathology Department of the University Hospitals Coventry and Warwickshire NHS Trust, UK. This dataset was first used in paper named, "HoVer-Net: Simultaneous Segmentation and Classification of Nuclei in Multi-Tissue Histology Images".
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| -------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------- |
+| [CoNIC202](https://conic-challenge.grand-challenge.org/) | abdomen | segmentation | histopathology | 7 | 4981/-/- | yes/-/- | 2022 | - |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :-----------------------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 27 | 83.61 | 14 | 80.4 | - | - |
+| other | 17 | 0.17 | 9 | 0.52 | - | - |
+| inflammatory | 25 | 2.66 | 14 | 2.14 | - | - |
+| healthy epithelial | 3 | 1.47 | 2 | 1.58 | - | - |
+| dysplastic/malignant epithelial | 10 | 7.17 | 8 | 9.16 | - | - |
+| fibroblast | 23 | 3.84 | 14 | 4.63 | - | - |
+| muscle | 8 | 1.05 | 3 | 1.42 | - | - |
+| endothelial | 7 | 0.02 | 4 | 0.15 | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `conic2022_seg/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://opendatalab.com/CoNSeP) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── consep
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Dataset Citation
+
+If this work is helpful for your research, please consider citing the below paper.
+
+```
+@article{graham2019hover,
+ title={Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images},
+ author={Graham, Simon and Vu, Quoc Dang and Raza, Shan E Ahmed and Azam, Ayesha and Tsang, Yee Wah and Kwak, Jin Tae and Rajpoot, Nasir},
+ journal={Medical Image Analysis},
+ volume={58},
+ pages={101563},
+ year={2019},
+ publisher={Elsevier}
+}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/consep/configs/consep_512x512.py b/projects/medical/2d_image/histopathology/consep/configs/consep_512x512.py
new file mode 100644
index 00000000000..0d9b8948b01
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/consep_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ConsepDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_consep-512x512.py b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_consep-512x512.py
new file mode 100644
index 00000000000..cbcf5db775b
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_consep-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './consep_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.consep_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=8),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_consep-512x512.py b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_consep-512x512.py
new file mode 100644
index 00000000000..b374566e6ef
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_consep-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './consep_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.consep_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=8),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_consep-512x512.py b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_consep-512x512.py
new file mode 100644
index 00000000000..35bdaa34c84
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_consep-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ './consep_512x512.py', 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.consep_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=8),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/consep/datasets/consep_dataset.py b/projects/medical/2d_image/histopathology/consep/datasets/consep_dataset.py
new file mode 100644
index 00000000000..ceb2b3ab25b
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/datasets/consep_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ConsepDataset(BaseSegDataset):
+ """ConsepDataset dataset.
+
+ In segmentation map annotation for ConsepDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(
+ classes=('background', 'other', 'inflammatory', 'healthy epithelial',
+ 'dysplastic/malignant epithelial', 'fibroblast', 'muscle',
+ 'endothelial'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/consep/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/consep/tools/prepare_dataset.py
new file mode 100755
index 00000000000..83a2e18ce10
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/consep/tools/prepare_dataset.py
@@ -0,0 +1,54 @@
+import glob
+import os
+import shutil
+
+import numpy as np
+from PIL import Image
+from scipy.io import loadmat
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.mat'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(os.path.join('data/CoNSeP/Train/Images/*' + img_suffix))
+x_test = glob.glob(os.path.join('data/CoNSeP/Test/Images/*' + img_suffix))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/val/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+os.system('mkdir -p ' + root_path + 'masks/val/')
+D2_255_convert_dict = {0: 0, 255: 1}
+
+
+def convert_2d(img, convert_dict=D2_255_convert_dict):
+ arr_2d = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr_2d[img == c] = i
+ return arr_2d
+
+
+part_dir_dict = {0: 'CoNSeP/Train/', 1: 'CoNSeP/Test/'}
+save_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ shutil.copy(
+ img, root_path + 'images/' + save_dir_dict[ith] +
+ basename.split('.')[0] + save_img_suffix)
+
+ mask_path = root_path + part_dir + 'Labels/' + basename.split(
+ '.')[0] + seg_map_suffix
+ label_ = loadmat(mask_path)
+ label = label_['inst_map']
+ label_type = label_['inst_type']
+ label_dict = {i + 1: int(val) for i, val in enumerate(label_type)}
+
+ save_mask_path = root_path + 'masks/' + save_dir_dict[
+ ith] + basename.split('.')[0] + save_seg_map_suffix
+
+ res = convert_2d(label, convert_dict=label_dict)
+ res = Image.fromarray(res.astype(np.uint8))
+ res.save(save_mask_path)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/README.md b/projects/medical/2d_image/histopathology/fusc2021/README.md
new file mode 100644
index 00000000000..8130d593503
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/README.md
@@ -0,0 +1,136 @@
+# Foot Ulcer Segmentation Challenge 2021 (FUSC 2021)
+
+## Description
+
+This project supports **`Foot Ulcer Segmentation Challenge 2021 (FUSC 2021) `**, which can be downloaded from [here](https://fusc.grand-challenge.org/).
+
+### Dataset Overview
+
+This chronic wound dataset was collected over 2 years from October 2019 to April 2021 at the center and contains 1,210 foot ulcer images taken from 889 patients during multiple clinical visits. The raw images were taken by Canon SX 620 HS digital camera and iPad Pro under uncontrolled illumination conditions,
+with various backgrounds. The images (shown in Figure 1) are randomly split into 3 subsets: a training set with 810 images, a validation set with 200 images, and a testing set with 200 images. Of course, the annotations of the testing set are kept private. The data collected were de-identified and in accordance with relevant guidelines and regulations and the patient’s informed consent is waived by the institutional review board of the University of Wisconsin-Milwaukee.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [fusc2021](https://fusc.grand-challenge.org/) | lower limb | segmentation | histopathology | 2 | 810/200/200 | yes/yes/no | 2021 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 810 | 98.71 | 200 | 98.78 | - | - |
+| wound | 791 | 1.29 | 195 | 1.22 | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{s41598-020-78799-w,
+ title={Fully automatic wound segmentation with deep convolutional neural networks},
+ author={Chuanbo Wang and D. M. Anisuzzaman and Victor Williamson and Mrinal Kanti Dhar and Behrouz Rostami and Jeffrey Niezgoda and Sandeep Gopalakrishnan and Zeyun Yu},
+ journal={Scientific Reports},
+ volume={10},
+ number={1},
+ pages={21897},
+ year={2020}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `fusc2021/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://fusc.grand-challenge.org/) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── fusc2021
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..c3f42751124
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_fusc2021-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..ed870303fff
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_fusc2021-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..cbc09ae6cdd
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_fusc2021-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_fusc2021-512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_fusc2021-512x512.py
new file mode 100644
index 00000000000..f1477ee725e
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_fusc2021-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './fusc2021_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.fusc2021_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/configs/fusc2021_512x512.py b/projects/medical/2d_image/histopathology/fusc2021/configs/fusc2021_512x512.py
new file mode 100644
index 00000000000..e650474cea8
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/configs/fusc2021_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'FUSC2021Dataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/fusc2021/datasets/fusc2021_dataset.py b/projects/medical/2d_image/histopathology/fusc2021/datasets/fusc2021_dataset.py
new file mode 100644
index 00000000000..d331ac8c3a2
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/datasets/fusc2021_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class FUSC2021Dataset(BaseSegDataset):
+ """FUSC2021Dataset dataset.
+
+ In segmentation map annotation for FUSC2021Dataset, 0 stands for background
+ , which is included in 2 categories. ``reduce_zero_label``
+ is fixed to False. The ``img_suffix`` is fixed to '.png' and
+ ``seg_map_suffix`` is fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'wound'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/fusc2021/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/fusc2021/tools/prepare_dataset.py
new file mode 100644
index 00000000000..8f2de3daa95
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/fusc2021/tools/prepare_dataset.py
@@ -0,0 +1,114 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+src_img_train_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/train/images')
+src_img_val_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/validation/images')
+src_img_test_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/test/images')
+src_mask_train_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/train/labels')
+src_mask_val_dir = os.path.join(
+ root_path, 'wound-segmentation/data/' +
+ 'Foot Ulcer Segmentation Challenge/validation/labels')
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_val_dir = os.path.join(root_path, 'images/val/')
+tgt_mask_val_dir = os.path.join(root_path, 'masks/val/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_img_val_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_mask_val_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ # filter out file names and paths in source directory
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob.glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+def convert_pics_into_pngs(src_dir, tgt_dir, suffix, convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_img_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+ num = len(src_paths)
+ img = np.array(Image.open(src_path))
+ if len(img.shape) == 2:
+ pil = Image.fromarray(img).convert(convert)
+ elif len(img.shape) == 3:
+ pil = Image.fromarray(img)
+ else:
+ raise ValueError('Input image not 2D/3D: ', img.shape)
+
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+def convert_label_pics_into_pngs(src_dir,
+ tgt_dir,
+ suffix,
+ convert_dict={
+ 0: 0,
+ 255: 1
+ }):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, save_seg_map_suffix)
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = np.array(Image.open(src_path).convert('L'))
+ img = convert_label(img, convert_dict)
+ Image.fromarray(img).save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+if __name__ == '__main__':
+
+ convert_pics_into_pngs(
+ src_img_train_dir, tgt_img_train_dir, suffix=img_suffix)
+
+ convert_pics_into_pngs(src_img_val_dir, tgt_img_val_dir, suffix=img_suffix)
+
+ convert_pics_into_pngs(
+ src_img_test_dir, tgt_img_test_dir, suffix=img_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_train_dir, tgt_mask_train_dir, suffix=seg_map_suffix)
+
+ convert_label_pics_into_pngs(
+ src_mask_val_dir, tgt_mask_val_dir, suffix=seg_map_suffix)
diff --git a/projects/medical/2d_image/histopathology/pannuke/README.md b/projects/medical/2d_image/histopathology/pannuke/README.md
new file mode 100644
index 00000000000..e0cade7536d
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/README.md
@@ -0,0 +1,146 @@
+# Pan-Cancer Histology Dataset for Nuclei Instance Segmentation and Classification (PanNuke)
+
+## Description
+
+This project supports **`Pan-Cancer Histology Dataset for Nuclei Instance Segmentation and Classification (PanNuke)`**, which can be downloaded from [here](https://academictorrents.com/details/99f2c7b57b95500711e33f2ee4d14c9fd7c7366c).
+
+### Dataset Overview
+
+Semi automatically generated nuclei instance segmentation and classification dataset with exhaustive nuclei labels across 19 different tissue types. The dataset consists of 481 visual fields, of which 312 are randomly sampled from more than 20K whole slide images at different magnifications, from multiple data sources. In total the dataset contains 205,343 labeled nuclei, each with an instance segmentation mask. Models trained on pannuke can aid in whole slide image tissue type segmentation, and generalise to new tissues. PanNuke demonstrates one of the first successfully semi-automatically generated datasets.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ---------------------------------------------------------------------------------------- | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Pannuke](https://academictorrents.com/details/99f2c7b57b95500711e33f2ee4d14c9fd7c7366c) | full_body | segmentation | histopathology | 6 | 7901/-/- | yes/-/- | 2019 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :-----------------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 7901 | 83.32 | - | - | - | - |
+| neoplastic | 4190 | 8.64 | - | - | - | - |
+| non-neoplastic epithelial | 4126 | 1.77 | - | - | - | - |
+| inflammatory | 6137 | 3.73 | - | - | - | - |
+| connective | 232 | 0.07 | - | - | - | - |
+| dead | 1528 | 2.47 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{gamper2019pannuke,
+ title={PanNuke: an open pan-cancer histology dataset for nuclei instance segmentation and classification},
+ author={Gamper, Jevgenij and Koohbanani, Navid Alemi and Benet, Ksenija and Khuram, Ali and Rajpoot, Nasir},
+ booktitle={European Congress on Digital Pathology},
+ pages={11--19},
+ year={2019},
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `pannuke/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://academictorrents.com/details/99f2c7b57b95500711e33f2ee4d14c9fd7c7366c) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── pannuke
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :-----------------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 6320 | 83.38 | 1581 | 83.1 | - | - |
+| neoplastic | 3339 | 8.55 | 851 | 9.0 | - | - |
+| non-neoplastic epithelial | 3293 | 1.77 | 833 | 1.76 | - | - |
+| inflammatory | 4914 | 3.72 | 1223 | 3.76 | - | - |
+| connective | 170 | 0.06 | 62 | 0.09 | - | - |
+| dead | 1235 | 2.51 | 293 | 2.29 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..92584e9a684
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './bactteria-detection_512x512.py', 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pannuke-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pannuke-512x512.py
new file mode 100644
index 00000000000..042a08ce008
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pannuke-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pannuke_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pannuke_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=6),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pannuke-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pannuke-512x512.py
new file mode 100644
index 00000000000..e92514c9132
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pannuke-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pannuke_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pannuke_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=6),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pannuke-512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pannuke-512x512.py
new file mode 100644
index 00000000000..a9403c849fa
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pannuke-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pannuke_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pannuke_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=6),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pannuke/configs/pannuke_512x512.py b/projects/medical/2d_image/histopathology/pannuke/configs/pannuke_512x512.py
new file mode 100644
index 00000000000..316ac1ac443
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/configs/pannuke_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'PanNukeDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/pannuke/datasets/pannuke_dataset.py b/projects/medical/2d_image/histopathology/pannuke/datasets/pannuke_dataset.py
new file mode 100644
index 00000000000..4d3c687ff31
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/datasets/pannuke_dataset.py
@@ -0,0 +1,33 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class PanNukeDataset(BaseSegDataset):
+ """PanNukeDataset dataset.
+
+ In segmentation map annotation for PanNukeDataset,
+ 0 stands for background, which is included in 6 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(
+ classes=('background', 'neoplastic', 'non-neoplastic epithelial',
+ 'inflammatory', 'connective', 'dead'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/pannuke/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/pannuke/tools/prepare_dataset.py
new file mode 100644
index 00000000000..7213b181f40
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pannuke/tools/prepare_dataset.py
@@ -0,0 +1,49 @@
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+
+tgt_img_dir = os.path.join(root_path, 'images/train')
+tgt_mask_dir = os.path.join(root_path, 'masks/train')
+os.system('mkdir -p ' + tgt_img_dir)
+os.system('mkdir -p ' + tgt_mask_dir)
+
+fold_img_paths = sorted([
+ os.path.join(root_path, 'pannuke/Fold 1/images/fold1/images.npy'),
+ os.path.join(root_path, 'pannuke/Fold 2/images/fold2/images.npy'),
+ os.path.join(root_path, 'pannuke/Fold 3/images/fold3/images.npy')
+])
+
+fold_mask_paths = sorted([
+ os.path.join(root_path, 'pannuke/Fold 1/masks/fold1/masks.npy'),
+ os.path.join(root_path, 'pannuke/Fold 2/masks/fold2/masks.npy'),
+ os.path.join(root_path, 'pannuke/Fold 3/masks/fold3/masks.npy')
+])
+
+for n, (img_path,
+ mask_path) in enumerate(zip(fold_img_paths, fold_mask_paths)):
+ fold_name = str(n + 1)
+ imgs = np.load(img_path)
+ masks = np.load(mask_path)
+
+ for i in range(imgs.shape[0]):
+ img = np.uint8(imgs[i])
+ mask_multichannel = np.minimum(np.uint8(masks[i]), 1)
+ mask = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
+ for j in range(mask_multichannel.shape[-1]):
+ factor = (j + 1) % mask_multichannel.shape[-1]
+ # convert [0,1,2,3,4,5] to [1,2,3,4,5,0],
+ # with the last label being background
+ mask[mask_multichannel[..., j] == 1] = factor
+
+ file_name = 'fold' + fold_name + '_' + str(i).rjust(4, '0') + '.png'
+ print('Processing: ', file_name)
+ tgt_img_path = os.path.join(tgt_img_dir, file_name)
+ tgt_mask_path = os.path.join(tgt_mask_dir, file_name)
+ Image.fromarray(img).save(tgt_img_path)
+ Image.fromarray(mask).save(tgt_mask_path)
+
+ del imgs
+ del masks
diff --git a/projects/medical/2d_image/histopathology/pcam/README.md b/projects/medical/2d_image/histopathology/pcam/README.md
new file mode 100644
index 00000000000..5a8094950c5
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/README.md
@@ -0,0 +1,153 @@
+# PCam (PatchCamelyon)
+
+## Description
+
+This project supports **`Patch Camelyon (PCam) `**, which can be downloaded from [here](https://opendatalab.com/PCam).
+
+### Dataset Overview
+
+PatchCamelyon is an image classification dataset. It consists of 327680 color images (96 x 96px) extracted from histopathologic scans of lymph node sections. Each image is annotated with a binary label indicating presence of metastatic tissue. PCam provides a new benchmark for machine learning models: bigger than CIFAR10, smaller than ImageNet, trainable on a single GPU.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------ | ----------------- | ------------ | -------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [Pcam](https://opendatalab.com/PCam) | throax | segmentation | histopathology | 2 | 327680/-/- | yes/-/- | 2018 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 214849 | 63.77 | - | - | - | - |
+| metastatic tissue | 131832 | 36.22 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@inproceedings{veeling2018rotation,
+ title={Rotation equivariant CNNs for digital pathology},
+ author={Veeling, Bastiaan S and Linmans, Jasper and Winkens, Jim and Cohen, Taco and Welling, Max},
+ booktitle={International Conference on Medical image computing and computer-assisted intervention},
+ pages={210--218},
+ year={2018},
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `pcam/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.com/PCam) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```shell
+mkdir data & cd data
+pip install opendatalab
+odl get PCam
+mv ./PCam/raw/pcamv1 ./
+rm -rf PCam
+cd ..
+python tools/prepare_dataset.py
+python ../../tools/split_seg_dataset.py
+```
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── histopathology
+ │ │ │ │ ├── pcam
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 171948 | 63.82 | 42901 | 63.6 | - | - |
+| metastatic tissue | 105371 | 36.18 | 26461 | 36.4 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pcam-512x512.py
new file mode 100644
index 00000000000..20601f1ea56
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_pcam-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pcam-512x512.py
new file mode 100644
index 00000000000..c057535409f
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_pcam-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pcam-512x512.py
new file mode 100644
index 00000000000..4c1d5fe421c
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_pcam-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_pcam-512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_pcam-512x512.py
new file mode 100644
index 00000000000..25e3734795c
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_pcam-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './pcam_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.pcam_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/histopathology/pcam/configs/pcam_512x512.py b/projects/medical/2d_image/histopathology/pcam/configs/pcam_512x512.py
new file mode 100644
index 00000000000..04efc23eb51
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/configs/pcam_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'PCamDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/histopathology/pcam/datasets/pcam_dataset.py b/projects/medical/2d_image/histopathology/pcam/datasets/pcam_dataset.py
new file mode 100644
index 00000000000..1c27de543ab
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/datasets/pcam_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class PCamDataset(BaseSegDataset):
+ """PCamDataset dataset.
+
+ In segmentation map annotation for PCamDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'metastatic tissue'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/histopathology/pcam/tools/prepare_dataset.py b/projects/medical/2d_image/histopathology/pcam/tools/prepare_dataset.py
new file mode 100644
index 00000000000..75038e6fb49
--- /dev/null
+++ b/projects/medical/2d_image/histopathology/pcam/tools/prepare_dataset.py
@@ -0,0 +1,49 @@
+import os
+
+import h5py
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_val_dir = os.path.join(root_path, 'images/val/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_val_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def extract_pics_from_h5(h5_path, h5_key, save_dir):
+ f = h5py.File(h5_path, 'r')
+ for i, img in enumerate(f[h5_key]):
+ img = img.astype(np.uint8).squeeze()
+ img = Image.fromarray(img)
+ save_image_path = os.path.join(save_dir, str(i).zfill(8) + '.png')
+ img.save(save_image_path)
+
+
+if __name__ == '__main__':
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_train_x.h5',
+ h5_key='x',
+ save_dir=tgt_img_train_dir)
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_valid_x.h5',
+ h5_key='x',
+ save_dir=tgt_img_val_dir)
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_test_x.h5',
+ h5_key='x',
+ save_dir=tgt_img_test_dir)
+
+ extract_pics_from_h5(
+ 'data/pcamv1/camelyonpatch_level_2_split_train_mask.h5',
+ h5_key='mask',
+ save_dir=tgt_mask_train_dir)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/README.md b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/README.md
new file mode 100644
index 00000000000..ca95921ba36
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/README.md
@@ -0,0 +1,167 @@
+# RAVIR: A Dataset and Methodology for the Semantic Segmentation and Quantitative Analysis of Retinal Arteries and Veins in Infrared Reflectance Imaging
+
+## Description
+
+This project support **`RAVIR: A Dataset and Methodology for the Semantic Segmentation and Quantitative Analysis of Retinal Arteries and Veins in Infrared Reflectance Imaging`**, and the dataset used in this project can be downloaded from [here](https://ravir.grand-challenge.org/).
+
+### Dataset Overview
+
+The retinal vasculature provides important clues in the diagnosis and monitoring of systemic diseases including hypertension and diabetes. The microvascular system is of primary involvement in such conditions, and the retina is the only anatomical site where the microvasculature can be directly observed. The objective assessment of retinal vessels has long been considered a surrogate biomarker for systemic vascular diseases, and with recent advancements in retinal imaging and computer vision technologies, this topic has become the subject of renewed attention. In this paper, we present a novel dataset, dubbed RAVIR, for the semantic segmentation of Retinal Arteries and Veins in Infrared Reflectance (IR) imaging. It enables the creation of deep learning-based models that distinguish extracted vessel type without extensive post-processing.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------- | ----------------- | ------------ | ---------------------------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Ravir](https://ravir.grand-challenge.org/) | eye | segmentation | infrared reflectance imaging | 3 | 23/-/19 | yes/-/- | 2022 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 23 | 87.22 | - | - | - | - |
+| artery | 23 | 5.45 | - | - | - | - |
+| vein | 23 | 7.33 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Dataset Citation
+
+```bibtex
+@article{hatamizadeh2022ravir,
+ title={RAVIR: A dataset and methodology for the semantic segmentation and quantitative analysis of retinal arteries and veins in infrared reflectance imaging},
+ author={Hatamizadeh, Ali and Hosseini, Hamid and Patel, Niraj and Choi, Jinseo and Pole, Cameron C and Hoeferlin, Cory M and Schwartz, Steven D and Terzopoulos, Demetri},
+ journal={IEEE Journal of Biomedical and Health Informatics},
+ volume={26},
+ number={7},
+ pages={3272--3283},
+ year={2022},
+ publisher={IEEE}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `ravir/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://ravir.grand-challenge.org/) and decompression data to path `'data/ravir/'`.
+- run script `"python tools/prepare_dataset.py"` to split dataset and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py --data_root data/ravir"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── infrared_reflectance_imaging
+ │ │ │ │ ├── ravir
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ │ ├── test
+ │ │ │ │ | │ │ │ ├── yyy.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── yyy.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 18 | 87.41 | 5 | 86.53 | - | - |
+| artery | 18 | 5.44 | 5 | 5.50 | - | - |
+| vein | 18 | 7.15 | 5 | 7.97 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+### Testing commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### Ravir
+
+| Method | Backbone | Crop Size | lr | config |
+| :-------------: | :------: | :-------: | :----: | :------------------------------------------------------------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py
new file mode 100755
index 00000000000..375ad5abf27
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_ravir-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './ravir_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.ravir_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py
new file mode 100755
index 00000000000..a7ecf6dd453
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_ravir-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './ravir_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.ravir_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=dict(size=img_scale),
+ pretrained=None,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py
new file mode 100755
index 00000000000..28556df53d1
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_ravir-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './ravir_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.ravir_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained=None,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/ravir_512x512.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/ravir_512x512.py
new file mode 100755
index 00000000000..cb4c292d1f7
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/configs/ravir_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'RAVIRDataset'
+data_root = 'data/ravir'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/__init__.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/__init__.py
new file mode 100755
index 00000000000..6f1d051bcff
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/__init__.py
@@ -0,0 +1,3 @@
+from .ravir_dataset import RAVIRDataset
+
+__all__ = ['RAVIRDataset']
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/ravir_dataset.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/ravir_dataset.py
new file mode 100755
index 00000000000..c9e0a8ed21e
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/datasets/ravir_dataset.py
@@ -0,0 +1,28 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class RAVIRDataset(BaseSegDataset):
+ """RAVIRDataset dataset.
+
+ In segmentation map annotation for RAVIRDataset, 0 stands for background,
+ which is included in 3 categories. ``reduce_zero_label`` is fixed to
+ False. The ``img_suffix`` is fixed to '.png' and ``seg_map_suffix`` is
+ fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'artery', 'vein'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/infrared_reflectance_imaging/ravir/tools/prepare_dataset.py b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/tools/prepare_dataset.py
new file mode 100644
index 00000000000..068dcad8145
--- /dev/null
+++ b/projects/medical/2d_image/infrared_reflectance_imaging/ravir/tools/prepare_dataset.py
@@ -0,0 +1,33 @@
+import glob
+import os
+
+import numpy as np
+from PIL import Image
+from tqdm import tqdm
+
+# map = {255:2, 128:1, 0:0}
+
+os.makedirs('data/ravir/images/train', exist_ok=True)
+os.makedirs('data/ravir/images/test', exist_ok=True)
+os.makedirs('data/ravir/masks/train', exist_ok=True)
+
+os.system(
+ r'cp data/ravir/RAVIR\ Dataset/train/training_images/* data/ravir/images/train' # noqa
+)
+os.system(
+ r'cp data/ravir/RAVIR\ Dataset/train/training_masks/* data/ravir/masks/train' # noqa
+)
+os.system(r'cp data/ravir/RAVIR\ Dataset/test/* data/ravir/images/test')
+
+os.system(r'rm -rf data/ravir/RAVIR\ Dataset')
+
+imgs = glob.glob(os.path.join('data/ravir/masks/train', '*.png'))
+
+for im_path in tqdm(imgs):
+ im = Image.open(im_path)
+ imn = np.array(im)
+ imn[imn == 255] = 2
+ imn[imn == 128] = 1
+ imn[imn == 0] = 0
+ new_im = Image.fromarray(imn)
+ new_im.save(im_path)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/README.md b/projects/medical/2d_image/microscopy_images/2pm_vessel/README.md
new file mode 100644
index 00000000000..1feb433a319
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/README.md
@@ -0,0 +1,153 @@
+# 2-PM Vessel Dataset
+
+## Description
+
+This project supports **`2-PM Vessel Dataset`**, which can be downloaded from [here](https://opendatalab.org.cn/2-PM_Vessel_Dataset).
+
+### Dataset Overview
+
+An open-source volumetric brain vasculature dataset obtained with two-photon microscopy at Focused Ultrasound Lab, at Sunnybrook Research Institute (affiliated with University of Toronto by Dr. Alison Burgess, Charissa Poon and Marc Santos).
+
+The dataset contains a total of 12 volumetric stacks consisting images of mouse brain vasculature and tumor vasculature.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------------------------ | ----------------- | ------------ | ----------------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [2pm_vessel](https://opendatalab.org.cn/2-PM_Vessel_Dataset) | vessel | segmentation | microscopy_images | 2 | 216/-/- | yes/-/- | 2021 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 216 | 85.78 | - | - | - | - |
+| vessel | 180 | 14.22 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{teikari2016deep,
+ title={Deep learning convolutional networks for multiphoton microscopy vasculature segmentation},
+ author={Teikari, Petteri and Santos, Marc and Poon, Charissa and Hynynen, Kullervo},
+ journal={arXiv preprint arXiv:1606.02382},
+ year={2016}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `2pm_vessel/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://opendatalab.org.cn/2-PM_Vessel_Dataset) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```shell
+mkdir data & cd data
+pip install opendatalab
+odl get 2-PM_Vessel_Dataset
+cd ..
+python tools/prepare_dataset.py
+python ../../tools/split_seg_dataset.py
+```
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── microscopy_images
+ │ │ │ │ ├── 2pm_vessel
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 172 | 85.88 | 44 | 85.4 | - | - |
+| vessel | 142 | 14.12 | 38 | 14.6 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [ ] Milestone 2: Indicates a successful model implementation.
+
+ - [ ] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/2pm-vessel_512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/2pm-vessel_512x512.py
new file mode 100644
index 00000000000..124403fa973
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/2pm-vessel_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'TwoPMVesselDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_2pm-vessel-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_2pm-vessel-512x512.py
new file mode 100644
index 00000000000..2a429e9068e
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_2pm-vessel-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_2pm-vessel-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_2pm-vessel-512x512.py
new file mode 100644
index 00000000000..10d9bb82f25
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_2pm-vessel-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_2pm-vessel-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_2pm-vessel-512x512.py
new file mode 100644
index 00000000000..65c1579ec71
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_2pm-vessel-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..91ed6ada3f2
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/configs/fcn-unet-s5-d16_unet_1xb16-0.01lr-sigmoid-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './2pm-vessel_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.2pm-vessel_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/datasets/2pm-vessel_dataset.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/datasets/2pm-vessel_dataset.py
new file mode 100644
index 00000000000..984b5a1361d
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/datasets/2pm-vessel_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class TwoPMVesselDataset(BaseSegDataset):
+ """TwoPMVesselDataset dataset.
+
+ In segmentation map annotation for TwoPMVesselDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'vessel'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/microscopy_images/2pm_vessel/tools/prepare_dataset.py b/projects/medical/2d_image/microscopy_images/2pm_vessel/tools/prepare_dataset.py
new file mode 100644
index 00000000000..1b46af2cad0
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/2pm_vessel/tools/prepare_dataset.py
@@ -0,0 +1,46 @@
+import os
+
+import tifffile as tiff
+from PIL import Image
+
+root_path = 'data/'
+
+image_dir = os.path.join(root_path,
+ '2-PM_Vessel_Dataset/raw/vesselNN_dataset/denoised')
+label_dir = os.path.join(root_path,
+ '2-PM_Vessel_Dataset/raw/vesselNN_dataset/labels')
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+
+
+def filter_suffix(src_dir, suffix):
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_names = [_ for _ in os.listdir(src_dir) if _.endswith(suffix)]
+ file_paths = [os.path.join(src_dir, _) for _ in file_names]
+ return sorted(file_paths), sorted(file_names)
+
+
+if __name__ == '__main__':
+
+ image_path_list, _ = filter_suffix(image_dir, suffix='tif')
+ label_path_list, _ = filter_suffix(label_dir, suffix='.tif')
+
+ for img_path, label_path in zip(image_path_list, label_path_list):
+ labels = tiff.imread(label_path)
+ images = tiff.imread(img_path)
+ assert labels.ndim == 3
+ assert images.shape == labels.shape
+ name = img_path.split('/')[-1].replace('.tif', '')
+ # a single .tif file contains multiple slices
+ # as long as it is read by tifffile package.
+ for i in range(labels.shape[0]):
+ slice_name = name + '_' + str(i).rjust(3, '0') + '.png'
+ image = images[i]
+ label = labels[i] // 255
+
+ save_path_label = os.path.join(tgt_mask_train_dir, slice_name)
+ Image.fromarray(label).save(save_path_label)
+ save_path_image = os.path.join(tgt_img_train_dir, slice_name)
+ Image.fromarray(image).convert('RGB').save(save_path_image)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/README.md b/projects/medical/2d_image/microscopy_images/bactteria_detection/README.md
new file mode 100644
index 00000000000..1cedda715a7
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/README.md
@@ -0,0 +1,160 @@
+# Bactteria detection with darkfield microscopy
+
+## Description
+
+This project supports **`Bactteria detection with darkfield microscopy`**, which can be downloaded from [here](https://tianchi.aliyun.com/dataset/94411).
+
+### Dataset Overview
+
+Spirochaeta is a genus of bacteria classified within the phylum Spirochaetes. Included in this dataset are 366 darkfield microscopy images and manually annotated masks which can be used for classification and segmentation purposes. Detecting bacteria in blood could have a huge significance for research in both the medical and computer science field.
+
+It was gathered and annotated by students (hand-on experience)
+It has more than just one targeted class (blood cell and bacteria were annotated)
+It is highly imbalanced, so naive loss functions would work less properly
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------------------------- | ----------------- | ------------ | ---------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Bactteria detection](https://tianchi.aliyun.com/dataset/94411) | bacteria | segmentation | microscopy | 3 | 366/-/- | yes/-/- | 2017 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 366 | 85.9 | - | - | - | - |
+| erythrocytes | 345 | 13.03 | - | - | - | - |
+| spirochaete | 288 | 1.07 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+## Usage
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow (PIL) v9.3.0
+- scikit-learn (sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `bactteria_detection/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- Download dataset from [here](https://tianchi.aliyun.com/dataset/94411) and save it to the `data/` directory .
+- Decompress data to path `data/`. This will create a new folder named `data/Bacteria_detection_with_darkfield_microscopy_datasets/`, which contains the original image data.
+- run script `python tools/prepare_dataset.py` to format data and change folder structure as below.
+- run script `python ../../tools/split_seg_dataset.py` to split dataset. For the Bacteria_detection dataset, as there is no test or validation dataset, we sample 20% samples from the whole dataset as the validation dataset and 80% samples for training data and make two filename lists `train.txt` and `val.txt`. As we set the random seed as the hard code, we eliminated the randomness, the dataset split actually can be reproducible.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── microscopy_images
+ │ │ │ │ ├── bactteria_detection
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── Bacteria_detection_with_darkfield_microscopy_datasets
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 292 | 85.66 | 74 | 86.7 | - | - |
+| erythrocytes | 274 | 13.25 | 71 | 12.29 | - | - |
+| spirochaete | 231 | 1.09 | 57 | 1.01 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### Bactteria detection with darkfield microscopy
+
+***Note: The following experimental results are based on the data randomly partitioned according to the above method described in the dataset preparing section.***
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config | download |
+| :-------------: | :------: | :-------: | :----: | :---: | :---: | :--------------------------------------------------------------------------------------: | :----------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | 76.48 | 84.68 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | 61.06 | 63.69 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | 58.87 | 62.42 | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py) | [model](<>) \| [log](<>) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/bactteria-detection_512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/bactteria-detection_512x512.py
new file mode 100644
index 00000000000..e3eab4e3868
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/bactteria-detection_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'BactteriaDetectionDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..ede58d785c1
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './bactteria-detection_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..bde3fa14acd
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './bactteria-detection_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py
new file mode 100644
index 00000000000..08e204f3809
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_bactteria-detection-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './bactteria-detection_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.bactteria-detection_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=3),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/datasets/bactteria-detection_dataset.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/datasets/bactteria-detection_dataset.py
new file mode 100644
index 00000000000..c95097b1acd
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/datasets/bactteria-detection_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class BactteriaDetectionDataset(BaseSegDataset):
+ """BactteriaDetectionDataset dataset.
+
+ In segmentation map annotation for BactteriaDetectionDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('background', 'erythrocytes', 'spirochaete'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/microscopy_images/bactteria_detection/tools/prepare_dataset.py b/projects/medical/2d_image/microscopy_images/bactteria_detection/tools/prepare_dataset.py
new file mode 100755
index 00000000000..8dcc719e262
--- /dev/null
+++ b/projects/medical/2d_image/microscopy_images/bactteria_detection/tools/prepare_dataset.py
@@ -0,0 +1,33 @@
+import glob
+import os
+import shutil
+
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = glob.glob(
+ 'data/Bacteria_detection_with_darkfield_microscopy_datasets/images/*' +
+ img_suffix) # noqa
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+part_dir_dict = {0: 'train/'}
+for ith, part in enumerate([x_train]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ basename.split('.')[0] + save_img_suffix)
+ shutil.copy(img, img_save_path)
+ mask_path = 'data/Bacteria_detection_with_darkfield_microscopy_datasets/masks/' + basename # noqa
+ mask = Image.open(mask_path).convert('L')
+ mask_save_path = os.path.join(
+ root_path, 'masks', part_dir,
+ basename.split('.')[0] + save_seg_map_suffix)
+ mask.save(mask_save_path)
diff --git a/projects/medical/2d_image/tools/split_seg_dataset.py b/projects/medical/2d_image/tools/split_seg_dataset.py
new file mode 100644
index 00000000000..9ab2e9282fa
--- /dev/null
+++ b/projects/medical/2d_image/tools/split_seg_dataset.py
@@ -0,0 +1,42 @@
+import argparse
+import glob
+import os
+
+from sklearn.model_selection import train_test_split
+
+
+def save_anno(img_list, file_path, remove_suffix=True):
+ if remove_suffix:
+ img_list = [
+ '/'.join(img_path.split('/')[-2:]) for img_path in img_list
+ ]
+ img_list = [
+ '.'.join(img_path.split('.')[:-1]) for img_path in img_list
+ ]
+ with open(file_path, 'w') as file_:
+ for x in list(img_list):
+ file_.write(x + '\n')
+
+
+if __name__ == '__main__':
+ parser = argparse.ArgumentParser()
+ parser.add_argument('--data_root', default='data/')
+ args = parser.parse_args()
+ data_root = args.data_root
+ if os.path.exists(os.path.join(data_root, 'masks/val')):
+ x_val = sorted(glob.glob(data_root + '/images/val/*.png'))
+ save_anno(x_val, data_root + '/val.txt')
+ if os.path.exists(os.path.join(data_root, 'masks/test')):
+ x_test = sorted(glob.glob(data_root + '/images/test/*.png'))
+ save_anno(x_test, data_root + '/test.txt')
+ if not os.path.exists(os.path.join(
+ data_root, 'masks/val')) and not os.path.exists(
+ os.path.join(data_root, 'masks/test')):
+ all_imgs = sorted(glob.glob(data_root + '/images/train/*.png'))
+ x_train, x_val = train_test_split(
+ all_imgs, test_size=0.2, random_state=0)
+ save_anno(x_train, data_root + '/train.txt')
+ save_anno(x_val, data_root + '/val.txt')
+ else:
+ x_train = sorted(glob.glob(data_root + '/images/train/*.png'))
+ save_anno(x_train, data_root + '/train.txt')
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/README.md b/projects/medical/2d_image/x_ray/chest_image_pneum/README.md
new file mode 100644
index 00000000000..a1cd27ba455
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/README.md
@@ -0,0 +1,147 @@
+# Chest Image Dataset for Pneumothorax Segmentation
+
+## Description
+
+This project supports **`Chest Image Dataset for Pneumothorax Segmentation`**, which can be downloaded from [here](https://tianchi.aliyun.com/dataset/83075).
+
+### Dataset Overview
+
+Pneumothorax can be caused by a blunt chest injury, damage from underlying lung disease, or most horrifying—it may occur for no obvious reason at all. On some occasions, a collapsed lung can be a life-threatening event.
+Pneumothorax is usually diagnosed by a radiologist on a chest x-ray, and can sometimes be very difficult to confirm. An accurate AI algorithm to detect pneumothorax would be useful in a lot of clinical scenarios. AI could be used to triage chest radiographs for priority interpretation, or to provide a more confident diagnosis for non-radiologists.
+
+The dataset is provided by the Society for Imaging Informatics in Medicine(SIIM), American College of Radiology (ACR),Society of Thoracic Radiology (STR) and MD.ai. You can develop a model to classify (and if present, segment) pneumothorax from a set of chest radiographic images. If successful, you could aid in the early recognition of pneumothoraces and save lives.
+
+### Original Statistic Information
+
+| Dataset name | Anatomical region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| --------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------------ |
+| [pneumothorax segmentation](https://tianchi.aliyun.com/dataset/83075) | thorax | segmentation | x_ray | 2 | 12089/-/3205 | yes/-/no | - | [CC-BY-SA-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 12089 | 99.75 | - | - | - | - |
+| pneumothorax area | 2669 | 0.25 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `chest_image_pneum/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://tianchi.aliyun.com/dataset/83075) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set can't be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── chest_image_pneum
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── test.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :---------------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| normal | 9637 | 99.75 | 2410 | 99.74 | - | - |
+| pneumothorax area | 2137 | 0.25 | 532 | 0.26 | - | - |
+
+### Training commands
+
+Train models on a single server with one GPU.
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+Test models on a single server with one GPU.
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Results
+
+### Bactteria detection with darkfield microscopy
+
+| Method | Backbone | Crop Size | lr | mIoU | mDice | config | download |
+| :-------------: | :------: | :-------: | :----: | :--: | :---: | :------------------------------------------------------------------------------------: | :----------------------: |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.01 | - | - | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.001 | - | - | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py) | [model](<>) \| [log](<>) |
+| fcn_unet_s5-d16 | unet | 512x512 | 0.0001 | - | - | [config](./configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py) | [model](<>) \| [log](<>) |
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+
+ - [x] Basic docstrings & proper citation
+
+ - [x] Test-time correctness
+
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+
+ - [ ] Unit tests
+
+ - [ ] Code polishing
+
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/chest-image-pneum_512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/chest-image-pneum_512x512.py
new file mode 100644
index 00000000000..411229bd413
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/chest-image-pneum_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ChestImagePneumDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py
new file mode 100644
index 00000000000..0f26459467c
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-image-pneum-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './chest-image-pneum_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.chest-image-pneum_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py
new file mode 100644
index 00000000000..37b91889d8a
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-image-pneum-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './chest-image-pneum_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.chest-image-pneum_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py
new file mode 100644
index 00000000000..379e8181f3d
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-image-pneum-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ './chest-image-pneum_512x512.py',
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.chest-image-pneum_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/datasets/chest-image-pneum_dataset.py b/projects/medical/2d_image/x_ray/chest_image_pneum/datasets/chest-image-pneum_dataset.py
new file mode 100644
index 00000000000..aeee60ae92e
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/datasets/chest-image-pneum_dataset.py
@@ -0,0 +1,27 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ChestImagePneumDataset(BaseSegDataset):
+ """ChestImagePneumDataset dataset.
+
+ In segmentation map annotation for ChestImagePneumDataset,
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ """
+ METAINFO = dict(classes=('normal', 'pneumothorax area'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=False,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/chest_image_pneum/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/chest_image_pneum/tools/prepare_dataset.py
new file mode 100755
index 00000000000..47eddc96dc5
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_image_pneum/tools/prepare_dataset.py
@@ -0,0 +1,73 @@
+import os
+
+import numpy as np
+import pandas as pd
+import pydicom
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.dcm'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+x_train = []
+for fpath, dirname, fnames in os.walk('data/chestimage_train_datasets'):
+ for fname in fnames:
+ if fname.endswith('.dcm'):
+ x_train.append(os.path.join(fpath, fname))
+x_test = []
+for fpath, dirname, fnames in os.walk('data/chestimage_test_datasets/'):
+ for fname in fnames:
+ if fname.endswith('.dcm'):
+ x_test.append(os.path.join(fpath, fname))
+
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/test/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+
+
+def rle_decode(rle, width, height):
+ mask = np.zeros(width * height, dtype=np.uint8)
+ array = np.asarray([int(x) for x in rle.split()])
+ starts = array[0::2]
+ lengths = array[1::2]
+
+ current_position = 0
+ for index, start in enumerate(starts):
+ current_position += start
+ mask[current_position:current_position + lengths[index]] = 1
+ current_position += lengths[index]
+
+ return mask.reshape(width, height, order='F')
+
+
+part_dir_dict = {0: 'train/', 1: 'test/'}
+dict_from_csv = pd.read_csv(
+ root_path + 'chestimage_train-rle_datasets.csv', sep=',',
+ index_col=0).to_dict()[' EncodedPixels']
+
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_id = '.'.join(basename.split('.')[:-1])
+ if ith == 0 and (img_id not in dict_from_csv.keys()):
+ continue
+ image = pydicom.read_file(img).pixel_array
+ save_img_path = root_path + 'images/' + part_dir + '.'.join(
+ basename.split('.')[:-1]) + save_img_suffix
+ print(save_img_path)
+ img_h, img_w = image.shape[:2]
+ image = Image.fromarray(image)
+ image.save(save_img_path)
+ if ith == 1:
+ continue
+ if dict_from_csv[img_id] == '-1':
+ mask = np.zeros((img_h, img_w), dtype=np.uint8)
+ else:
+ mask = rle_decode(dict_from_csv[img_id], img_h, img_w)
+ save_mask_path = root_path + 'masks/' + part_dir + '.'.join(
+ basename.split('.')[:-1]) + save_seg_map_suffix
+ mask = Image.fromarray(mask)
+ mask.save(save_mask_path)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/README.md b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/README.md
new file mode 100644
index 00000000000..7cb099c8a43
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/README.md
@@ -0,0 +1,119 @@
+# Chest X-ray Images with Pneumothorax Masks
+
+## Description
+
+This project support **`Chest X-ray Images with Pneumothorax Masks `**, and the dataset used in this project can be downloaded from [here](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks).
+
+### Dataset Overview
+
+A pneumothorax (noo-moe-THOR-aks) is a collapsed lung. A pneumothorax occurs when air leaks into the space between your lung and chest wall. This air pushes on the outside of your lung and makes it collapse. Pneumothorax can be a complete lung collapse or a collapse of only a portion of the lung.
+
+A pneumothorax can be caused by a blunt or penetrating chest injury, certain medical procedures, or damage from underlying lung disease. Or it may occur for no obvious reason. Symptoms usually include sudden chest pain and shortness of breath. On some occasions, a collapsed lung can be a life-threatening event.
+
+Treatment for a pneumothorax usually involves inserting a needle or chest tube between the ribs to remove the excess air. However, a small pneumothorax may heal on its own.
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release date | License |
+| --------------------------------------------------------------------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------- |
+| [Chest-x-ray-images-with-pneumothorax-masks](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks) | throax | segmentation | x_ray | 2 | 10675/-/1372 | yes/-/yes | 2020 | [CC-BY-NC 4.0](https://creativecommons.org/licenses/by-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :----------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 10675 | 99.7 | - | - | 1372 | 99.71 |
+| pneumothroax | 2379 | 0.3 | - | - | 290 | 0.29 |
+
+### Visualization
+
+
+
+### Prerequisites
+
+- Python 3.8
+- PyTorch 1.10.0
+- pillow(PIL) 9.3.0
+- scikit-learn(sklearn) 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of PYTHONPATH, which should point to the project's directory so that Python can locate the module files. In chest_x_ray_images_with_pneumothorax_masks/ root directory, run the following line to add the current directory to PYTHONPATH:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset preparing
+
+- download dataset from [here](https://www.kaggle.com/datasets/vbookshelf/pneumothorax-chest-xray-images-and-masks) and decompression data to path 'data/'.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── chest_x_ray_images_with_pneumothorax_masks
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Training commands
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH}
+```
+
+To train on multiple GPUs, e.g. 8 GPUs, run the following command:
+
+```shell
+mim train mmseg ./configs/${CONFIG_PATH} --launcher pytorch --gpus 8
+```
+
+### Testing commands
+
+```shell
+mim test mmseg ./configs/${CONFIG_PATH} --checkpoint ${CHECKPOINT_PATH}
+```
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [x] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/chest-x-ray-images-with-pneumothorax-masks_512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/chest-x-ray-images-with-pneumothorax-masks_512x512.py
new file mode 100644
index 00000000000..96676de8616
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/chest-x-ray-images-with-pneumothorax-masks_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'ChestPenumoMaskDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..76c214d04cb
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,20 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..066996dda94
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..a7065b82315
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
new file mode 100644
index 00000000000..e5682ee76b7
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_chest-x-ray-images-with-pneumothorax-masks-512x512.py
@@ -0,0 +1,19 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py',
+ './chest-x-ray-images-with-pneumothorax-masks_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(
+ imports='datasets.chest-x-ray-images-with-pneumothorax-masks_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/datasets/chest-x-ray-images-with-pneumothorax-masks_dataset.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/datasets/chest-x-ray-images-with-pneumothorax-masks_dataset.py
new file mode 100644
index 00000000000..d32f597a5a0
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/datasets/chest-x-ray-images-with-pneumothorax-masks_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class ChestPenumoMaskDataset(BaseSegDataset):
+ """ChestPenumoMaskDataset dataset.
+
+ In segmentation map annotation for ChestPenumoMaskDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'penumothroax'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/tools/prepare_dataset.py
new file mode 100644
index 00000000000..c7de1f19042
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/chest_x_ray_images_with_pneumothorax_masks/tools/prepare_dataset.py
@@ -0,0 +1,36 @@
+import glob
+import os
+import shutil
+
+from PIL import Image
+from sklearn.model_selection import train_test_split
+
+root_path = 'data/'
+img_suffix = '.png'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+all_imgs = glob.glob('data/siim-acr-pneumothorax/png_images/*' + img_suffix)
+x_train, x_test = train_test_split(all_imgs, test_size=0.2, random_state=0)
+
+print(len(x_train), len(x_test))
+os.system('mkdir -p ' + root_path + 'images/train/')
+os.system('mkdir -p ' + root_path + 'images/val/')
+os.system('mkdir -p ' + root_path + 'masks/train/')
+os.system('mkdir -p ' + root_path + 'masks/val/')
+
+part_dir_dict = {0: 'train/', 1: 'val/'}
+for ith, part in enumerate([x_train, x_test]):
+ part_dir = part_dir_dict[ith]
+ for img in part:
+ basename = os.path.basename(img)
+ img_save_path = os.path.join(root_path, 'images', part_dir,
+ basename.split('.')[0] + save_img_suffix)
+ shutil.copy(img, img_save_path)
+ mask_path = 'data/siim-acr-pneumothorax/png_masks/' + basename
+ mask = Image.open(mask_path).convert('L')
+ mask_save_path = os.path.join(
+ root_path, 'masks', part_dir,
+ basename.split('.')[0] + save_seg_map_suffix)
+ mask.save(mask_save_path)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/README.md b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/README.md
new file mode 100644
index 00000000000..8469219effa
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/README.md
@@ -0,0 +1,158 @@
+# Covid-19 CT Chest X-ray Dataset
+
+## Description
+
+This project supports **`Covid-19 CT Chest X-ray Dataset`**, which can be downloaded from [here](https://github.com/ieee8023/covid-chestxray-dataset).
+
+### Dataset Overview
+
+In the context of a COVID-19 pandemic, we want to improve prognostic predictions to triage and manage patient care. Data is the first step to developing any diagnostic/prognostic tool. While there exist large public datasets of more typical chest X-rays from the NIH \[Wang 2017\], Spain \[Bustos 2019\], Stanford \[Irvin 2019\], MIT \[Johnson 2019\] and Indiana University \[Demner-Fushman 2016\], there is no collection of COVID-19 chest X-rays or CT scans designed to be used for computational analysis.
+
+The 2019 novel coronavirus (COVID-19) presents several unique features [Fang, 2020](https://pubs.rsna.org/doi/10.1148/radiol.2020200432) and [Ai 2020](https://pubs.rsna.org/doi/10.1148/radiol.2020200642). While the diagnosis is confirmed using polymerase chain reaction (PCR), infected patients with pneumonia may present on chest X-ray and computed tomography (CT) images with a pattern that is only moderately characteristic for the human eye [Ng, 2020](https://pubs.rsna.org/doi/10.1148/ryct.2020200034). In late January, a Chinese team published a paper detailing the clinical and paraclinical features of COVID-19. They reported that patients present abnormalities in chest CT images with most having bilateral involvement [Huang 2020](). Bilateral multiple lobular and subsegmental areas of consolidation constitute the typical findings in chest CT images of intensive care unit (ICU) patients on admission [Huang 2020](). In comparison, non-ICU patients show bilateral ground-glass opacity and subsegmental areas of consolidation in their chest CT images [Huang 2020](). In these patients, later chest CT images display bilateral ground-glass opacity with resolved consolidation [Huang 2020]().
+
+### Statistic Information
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Nnum. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release date | License |
+| ---------------------------------------------------------------------- | ----------------- | ------------ | -------- | ------------- | --------------------- | ---------------------- | ------------ | --------------------------------------------------------------------- |
+| [Covid-19-ct-cxr](https://github.com/ieee8023/covid-chestxray-dataset) | thorax | segmentation | x_ray | 2 | 205/-/714 | yes/-/no | 2021 | [CC-BY-NC-SA 4.0](https://creativecommons.org/licenses/by-nc-sa/4.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 205 | 72.84 | - | - | - | - |
+| lung | 205 | 27.16 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{cohen2020covidProspective,
+ title={{COVID-19} Image Data Collection: Prospective Predictions Are the Future},
+ author={Joseph Paul Cohen and Paul Morrison and Lan Dao and Karsten Roth and Tim Q Duong and Marzyeh Ghassemi},
+ journal={arXiv 2006.11988},
+ year={2020}
+}
+
+@article{cohen2020covid,
+ title={COVID-19 image data collection},
+ author={Joseph Paul Cohen and Paul Morrison and Lan Dao},
+ journal={arXiv 2003.11597},
+ year={2020}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0 9.3.0
+- scikit-learn(sklearn) v1.2.0 1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `covid_19_ct_cxr/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://github.com/ieee8023/covid-chestxray-dataset) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```shell
+mkdir data && cd data
+git clone git@github.com:ieee8023/covid-chestxray-dataset.git
+cd ..
+python tools/prepare_dataset.py
+python ../../tools/split_seg_dataset.py
+```
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── covid_19_ct_cxr
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 164 | 72.88 | 41 | 72.69 | - | - |
+| lung | 164 | 27.12 | 41 | 27.31 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [x] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/covid-19-ct-cxr_512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/covid-19-ct-cxr_512x512.py
new file mode 100644
index 00000000000..5242d06c374
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/covid-19-ct-cxr_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'Covid19CXRDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='val.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..59a7bedaa06
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet-{use-sigmoid}_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..83b8527d46d
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..10cfcbda6ec
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
new file mode 100644
index 00000000000..aaccc8fd8dd
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_covid-19-ct-cxr-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './covid-19-ct-cxr_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.covid-19-ct-cxr_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/datasets/covid-19-ct-cxr_dataset.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/datasets/covid-19-ct-cxr_dataset.py
new file mode 100644
index 00000000000..68a1bb331f7
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/datasets/covid-19-ct-cxr_dataset.py
@@ -0,0 +1,31 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class Covid19CXRDataset(BaseSegDataset):
+ """Covid19CXRDataset dataset.
+
+ In segmentation map annotation for Covid19CXRDataset,
+ 0 stands for background, which is included in 2 categories.
+ ``reduce_zero_label`` is fixed to False. The ``img_suffix``
+ is fixed to '.png' and ``seg_map_suffix`` is fixed to '.png'.
+
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False.
+ """
+ METAINFO = dict(classes=('background', 'lung'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/covid_19_ct_cxr/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/tools/prepare_dataset.py
new file mode 100644
index 00000000000..72f6435389d
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/covid_19_ct_cxr/tools/prepare_dataset.py
@@ -0,0 +1,52 @@
+import os
+
+import numpy as np
+from PIL import Image
+
+root_path = 'data/'
+src_img_dir = os.path.join(root_path, 'covid-chestxray-dataset', 'images')
+src_mask_dir = os.path.join(root_path, 'covid-chestxray-dataset',
+ 'annotations/lungVAE-masks')
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def convert_label(img, convert_dict):
+ arr = np.zeros_like(img, dtype=np.uint8)
+ for c, i in convert_dict.items():
+ arr[img == c] = i
+ return arr
+
+
+if __name__ == '__main__':
+
+ all_img_names = os.listdir(src_img_dir)
+ all_mask_names = os.listdir(src_mask_dir)
+
+ for img_name in all_img_names:
+ base_name = img_name.replace('.png', '')
+ base_name = base_name.replace('.jpg', '')
+ base_name = base_name.replace('.jpeg', '')
+ mask_name_orig = base_name + '_mask.png'
+ if mask_name_orig in all_mask_names:
+ mask_name = base_name + '.png'
+ src_img_path = os.path.join(src_img_dir, img_name)
+ src_mask_path = os.path.join(src_mask_dir, mask_name_orig)
+ tgt_img_path = os.path.join(tgt_img_train_dir, img_name)
+ tgt_mask_path = os.path.join(tgt_mask_train_dir, mask_name)
+
+ img = Image.open(src_img_path).convert('RGB')
+ img.save(tgt_img_path)
+ mask = np.array(Image.open(src_mask_path))
+ mask = convert_label(mask, {0: 0, 255: 1})
+ mask = Image.fromarray(mask)
+ mask.save(tgt_mask_path)
+ else:
+ src_img_path = os.path.join(src_img_dir, img_name)
+ tgt_img_path = os.path.join(tgt_img_test_dir, img_name)
+ img = Image.open(src_img_path).convert('RGB')
+ img.save(tgt_img_path)
diff --git a/projects/medical/2d_image/x_ray/crass/README.md b/projects/medical/2d_image/x_ray/crass/README.md
new file mode 100644
index 00000000000..0621205be81
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/README.md
@@ -0,0 +1,144 @@
+# Chest Radiograph Anatomical Structure Segmentation (CRASS)
+
+## Description
+
+This project supports **`Chest Radiograph Anatomical Structure Segmentation (CRASS) `**, which can be downloaded from [here](https://crass.grand-challenge.org/).
+
+### Dataset Overview
+
+A set of consecutively obtained posterior-anterior chest radiograph were selected from a database containing images acquired at two sites in sub Saharan Africa with a high tuberculosis incidence. All subjects were 15 years or older. Images from digital chest radiography units were used (Delft Imaging Systems, The Netherlands) of varying resolutions, with a typical resolution of 1800--2000 pixels, the pixel size was 250 lm isotropic. From the total set of images, 225 were considered to be normal by an expert radiologist, while 333 of the images contained abnormalities. Of the abnormal images, 220 contained abnormalities in the upper area of the lung where the clavicle is located. The data was divided into a training and a test set. The training set consisted of 299 images, the test set of 249 images.
+The current data is still incomplete and to be added later.
+
+### Information Statistics
+
+| Dataset Name | Anatomical Region | Task Type | Modality | Num. Classes | Train/Val/Test Images | Train/Val/Test Labeled | Release Date | License |
+| ------------------------------------------- | ----------------- | ------------ | -------- | ------------ | --------------------- | ---------------------- | ------------ | ------------------------------------------------------------- |
+| [crass](https://crass.grand-challenge.org/) | pulmonary | segmentation | x_ray | 2 | 299/-/234 | yes/-/no | 2021 | [CC0 1.0](https://creativecommons.org/publicdomain/zero/1.0/) |
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 299 | 98.38 | - | - | - | - |
+| clavicles | 299 | 1.62 | - | - | - | - |
+
+Note:
+
+- `Pct` means percentage of pixels in this category in all pixels.
+
+### Visualization
+
+
+
+### Dataset Citation
+
+```
+@article{HOGEWEG20121490,
+ title={Clavicle segmentation in chest radiographs},
+ journal={Medical Image Analysis},
+ volume={16},
+ number={8},
+ pages={1490-1502},
+ year={2012}
+}
+```
+
+### Prerequisites
+
+- Python v3.8
+- PyTorch v1.10.0
+- pillow(PIL) v9.3.0
+- scikit-learn(sklearn) v1.2.0
+- [MIM](https://github.com/open-mmlab/mim) v0.3.4
+- [MMCV](https://github.com/open-mmlab/mmcv) v2.0.0rc4
+- [MMEngine](https://github.com/open-mmlab/mmengine) v0.2.0 or higher
+- [MMSegmentation](https://github.com/open-mmlab/mmsegmentation) v1.0.0rc5
+
+All the commands below rely on the correct configuration of `PYTHONPATH`, which should point to the project's directory so that Python can locate the module files. In `crass/` root directory, run the following line to add the current directory to `PYTHONPATH`:
+
+```shell
+export PYTHONPATH=`pwd`:$PYTHONPATH
+```
+
+### Dataset Preparing
+
+- download dataset from [here](https://crass.grand-challenge.org/) and decompress data to path `'data/'`.
+- run script `"python tools/prepare_dataset.py"` to format data and change folder structure as below.
+- run script `"python ../../tools/split_seg_dataset.py"` to split dataset and generate `train.txt`, `val.txt` and `test.txt`. If the label of official validation set and test set cannot be obtained, we generate `train.txt` and `val.txt` from the training set randomly.
+
+```none
+ mmsegmentation
+ ├── mmseg
+ ├── projects
+ │ ├── medical
+ │ │ ├── 2d_image
+ │ │ │ ├── x_ray
+ │ │ │ │ ├── crass
+ │ │ │ │ │ ├── configs
+ │ │ │ │ │ ├── datasets
+ │ │ │ │ │ ├── tools
+ │ │ │ │ │ ├── data
+ │ │ │ │ │ │ ├── train.txt
+ │ │ │ │ │ │ ├── val.txt
+ │ │ │ │ │ │ ├── images
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+ │ │ │ │ │ │ ├── masks
+ │ │ │ │ │ │ │ ├── train
+ │ │ │ │ | │ │ │ ├── xxx.png
+ │ │ │ │ | │ │ │ ├── ...
+ │ │ │ │ | │ │ │ └── xxx.png
+```
+
+### Divided Dataset Information
+
+***Note: The table information below is divided by ourselves.***
+
+| Class Name | Num. Train | Pct. Train | Num. Val | Pct. Val | Num. Test | Pct. Test |
+| :--------: | :--------: | :--------: | :------: | :------: | :-------: | :-------: |
+| background | 227 | 98.38 | 57 | 98.39 | - | - |
+| clavicles | 227 | 1.62 | 57 | 1.61 | - | - |
+
+### Training commands
+
+To train models on a single server with one GPU. (default)
+
+```shell
+mim train mmseg ./configs/${CONFIG_FILE}
+```
+
+### Testing commands
+
+To test models on a single server with one GPU. (default)
+
+```shell
+mim test mmseg ./configs/${CONFIG_FILE} --checkpoint ${CHECKPOINT_PATH}
+```
+
+
+
+## Checklist
+
+- [x] Milestone 1: PR-ready, and acceptable to be one of the `projects/`.
+
+ - [x] Finish the code
+ - [x] Basic docstrings & proper citation
+ - [ ] Test-time correctness
+ - [x] A full README
+
+- [x] Milestone 2: Indicates a successful model implementation.
+
+ - [x] Training-time correctness
+
+- [ ] Milestone 3: Good to be a part of our core package!
+
+ - [ ] Type hints and docstrings
+ - [ ] Unit tests
+ - [ ] Code polishing
+ - [ ] Metafile.yml
+
+- [ ] Move your modules into the core package following the codebase's file hierarchy structure.
+
+- [ ] Refactor your modules into the core package following the codebase's file hierarchy structure.
diff --git a/projects/medical/2d_image/x_ray/crass/configs/crass_512x512.py b/projects/medical/2d_image/x_ray/crass/configs/crass_512x512.py
new file mode 100644
index 00000000000..1425f50cc46
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/crass_512x512.py
@@ -0,0 +1,42 @@
+dataset_type = 'CRASSDataset'
+data_root = 'data/'
+img_scale = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=img_scale, keep_ratio=False),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+train_dataloader = dict(
+ batch_size=16,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='train.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ ann_file='tval.txt',
+ data_prefix=dict(img_path='images/', seg_map_path='masks/'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
+test_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU', 'mDice'])
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_crass-512x512.py
new file mode 100644
index 00000000000..b52bc78f790
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.0001-20k_crass-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.0001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_crass-512x512.py
new file mode 100644
index 00000000000..45242c65b48
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.001-20k_crass-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.001)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_crass-512x512.py
new file mode 100644
index 00000000000..bcf9d0a5ca5
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-0.01-20k_crass-512x512.py
@@ -0,0 +1,17 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(num_classes=2),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-lr0.01-sigmoid-20k_crass-512x512.py b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-lr0.01-sigmoid-20k_crass-512x512.py
new file mode 100644
index 00000000000..0dde736bf76
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/configs/fcn-unet-s5-d16_unet_1xb16-lr0.01-sigmoid-20k_crass-512x512.py
@@ -0,0 +1,18 @@
+_base_ = [
+ 'mmseg::_base_/models/fcn_unet_s5-d16.py', './crass_512x512.py',
+ 'mmseg::_base_/default_runtime.py',
+ 'mmseg::_base_/schedules/schedule_20k.py'
+]
+custom_imports = dict(imports='datasets.crass_dataset')
+img_scale = (512, 512)
+data_preprocessor = dict(size=img_scale)
+optimizer = dict(lr=0.01)
+optim_wrapper = dict(optimizer=optimizer)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ decode_head=dict(
+ num_classes=2, loss_decode=dict(use_sigmoid=True), out_channels=1),
+ auxiliary_head=None,
+ test_cfg=dict(mode='whole', _delete_=True))
+vis_backends = None
+visualizer = dict(vis_backends=vis_backends)
diff --git a/projects/medical/2d_image/x_ray/crass/datasets/crass_dataset.py b/projects/medical/2d_image/x_ray/crass/datasets/crass_dataset.py
new file mode 100644
index 00000000000..f6b5c5228bc
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/datasets/crass_dataset.py
@@ -0,0 +1,30 @@
+from mmseg.datasets import BaseSegDataset
+from mmseg.registry import DATASETS
+
+
+@DATASETS.register_module()
+class CRASSDataset(BaseSegDataset):
+ """CRASSDataset dataset.
+
+ In segmentation map annotation for CRASSDataset, 0 stands for background,
+ which is included in 2 categories. ``reduce_zero_label`` is fixed to
+ False. The ``img_suffix`` is fixed to '.png' and ``seg_map_suffix`` is
+ fixed to '.png'.
+ Args:
+ img_suffix (str): Suffix of images. Default: '.png'
+ seg_map_suffix (str): Suffix of segmentation maps. Default: '.png'
+ reduce_zero_label (bool): Whether to mark label zero as ignored.
+ Default to False..
+ """
+ METAINFO = dict(classes=('background', 'clavicles'))
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=False,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
diff --git a/projects/medical/2d_image/x_ray/crass/tools/prepare_dataset.py b/projects/medical/2d_image/x_ray/crass/tools/prepare_dataset.py
new file mode 100644
index 00000000000..bbd5d8891d4
--- /dev/null
+++ b/projects/medical/2d_image/x_ray/crass/tools/prepare_dataset.py
@@ -0,0 +1,84 @@
+import glob
+import os
+
+import cv2
+import SimpleITK as sitk
+from PIL import Image
+
+root_path = 'data/'
+img_suffix = '.tif'
+seg_map_suffix = '.png'
+save_img_suffix = '.png'
+save_seg_map_suffix = '.png'
+
+src_img_train_dir = os.path.join(root_path, 'CRASS/data_train')
+src_mask_train_dir = os.path.join(root_path, 'CRASS/mask_mhd')
+src_img_test_dir = os.path.join(root_path, 'CRASS/data_test')
+
+tgt_img_train_dir = os.path.join(root_path, 'images/train/')
+tgt_mask_train_dir = os.path.join(root_path, 'masks/train/')
+tgt_img_test_dir = os.path.join(root_path, 'images/test/')
+os.system('mkdir -p ' + tgt_img_train_dir)
+os.system('mkdir -p ' + tgt_mask_train_dir)
+os.system('mkdir -p ' + tgt_img_test_dir)
+
+
+def filter_suffix_recursive(src_dir, suffix):
+ suffix = '.' + suffix if '.' not in suffix else suffix
+ file_paths = glob(
+ os.path.join(src_dir, '**', '*' + suffix), recursive=True)
+ file_names = [_.split('/')[-1] for _ in file_paths]
+ return sorted(file_paths), sorted(file_names)
+
+
+def read_single_array_from_med(path):
+ return sitk.GetArrayFromImage(sitk.ReadImage(path)).squeeze()
+
+
+def convert_meds_into_pngs(src_dir,
+ tgt_dir,
+ suffix='.dcm',
+ norm_min=0,
+ norm_max=255,
+ convert='RGB'):
+ if not os.path.exists(tgt_dir):
+ os.makedirs(tgt_dir)
+
+ src_paths, src_names = filter_suffix_recursive(src_dir, suffix=suffix)
+ num = len(src_paths)
+ for i, (src_name, src_path) in enumerate(zip(src_names, src_paths)):
+ tgt_name = src_name.replace(suffix, '.png')
+ tgt_path = os.path.join(tgt_dir, tgt_name)
+
+ img = read_single_array_from_med(src_path)
+ if norm_min is not None and norm_max is not None:
+ img = cv2.normalize(img, None, norm_min, norm_max, cv2.NORM_MINMAX,
+ cv2.CV_8U)
+ pil = Image.fromarray(img).convert(convert)
+ pil.save(tgt_path)
+ print(f'processed {i+1}/{num}.')
+
+
+convert_meds_into_pngs(
+ src_img_train_dir,
+ tgt_img_train_dir,
+ suffix='.mhd',
+ norm_min=0,
+ norm_max=255,
+ convert='RGB')
+
+convert_meds_into_pngs(
+ src_img_test_dir,
+ tgt_img_test_dir,
+ suffix='.mhd',
+ norm_min=0,
+ norm_max=255,
+ convert='RGB')
+
+convert_meds_into_pngs(
+ src_mask_train_dir,
+ tgt_mask_train_dir,
+ suffix='.mhd',
+ norm_min=0,
+ norm_max=1,
+ convert='L')
diff --git a/projects/pp_mobileseg/README.md b/projects/pp_mobileseg/README.md
new file mode 100644
index 00000000000..c9f9c128e74
--- /dev/null
+++ b/projects/pp_mobileseg/README.md
@@ -0,0 +1,123 @@
+# PP-MobileSeg: Exploring Transformer Blocks for Efficient Mobile Segmentation.
+
+## Reference
+
+> [PP-MobileSeg: Explore the Fast and Accurate Semantic Segmentation Model on Mobile Devices. ](https://arxiv.org/abs/2304.05152)
+
+## Introduction
+
+Official Repo
+
+Code Snippet
+
+##  Abstract
+
+With the success of transformers in computer vision, several attempts have been made to adapt transformers to mobile devices. However, their performance is not satisfied for some real world applications. Therefore, we propose PP-MobileSeg, a SOTA semantic segmentation model for mobile devices.
+
+It is composed of three newly proposed parts, the strideformer backbone, the Aggregated Attention Module(AAM), and the Valid Interpolate Module(VIM):
+
+- With the four-stage MobileNetV3 block as the feature extractor, we manage to extract rich local features of different receptive fields with little parameter overhead. Also, we further efficiently empower features from the last two stages with the global view using strided sea attention.
+- To effectively fuse the features, we use AAM to filter the detail features with ensemble voting and add the semantic feature to it to enhance the semantic information to the most content.
+- At last, we use VIM to upsample the downsampled feature to the original resolution and significantly decrease latency in model inference stage. It only interpolates classes present in the final prediction which only takes around 10% in the ADE20K dataset. This is a common scenario for datasets with large classes. Therefore it significantly decreases the latency of the final upsample process which takes the greatest part of the model's overall latency.
+
+Extensive experiments show that PP-MobileSeg achieves a superior params-accuracy-latency tradeoff compared to other SOTA methods.
+
+
Abstract
+
+With the success of transformers in computer vision, several attempts have been made to adapt transformers to mobile devices. However, their performance is not satisfied for some real world applications. Therefore, we propose PP-MobileSeg, a SOTA semantic segmentation model for mobile devices.
+
+It is composed of three newly proposed parts, the strideformer backbone, the Aggregated Attention Module(AAM), and the Valid Interpolate Module(VIM):
+
+- With the four-stage MobileNetV3 block as the feature extractor, we manage to extract rich local features of different receptive fields with little parameter overhead. Also, we further efficiently empower features from the last two stages with the global view using strided sea attention.
+- To effectively fuse the features, we use AAM to filter the detail features with ensemble voting and add the semantic feature to it to enhance the semantic information to the most content.
+- At last, we use VIM to upsample the downsampled feature to the original resolution and significantly decrease latency in model inference stage. It only interpolates classes present in the final prediction which only takes around 10% in the ADE20K dataset. This is a common scenario for datasets with large classes. Therefore it significantly decreases the latency of the final upsample process which takes the greatest part of the model's overall latency.
+
+Extensive experiments show that PP-MobileSeg achieves a superior params-accuracy-latency tradeoff compared to other SOTA methods.
+
+
+

+
 Performance
+
+### ADE20K
+
+| Model | Backbone | Training Iters | Batchsize | Train Resolution | mIoU(%) | latency(ms)\* | params(M) | config | Links |
+| ----------------- | ----------------- | -------------- | --------- | ---------------- | ------- | ------------- | --------- | ------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| PP-MobileSeg-Base | StrideFormer-Base | 80000 | 32 | 512x512 | 41.57% | 265.5 | 5.62 | [config](https://github.com/Yang-Changhui/mmsegmentation/tree/add_ppmobileseg/projects/pp_mobileseg/configs/pp_mobileseg) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_2xb16_3rdparty-base_512x512-ade20k-f12b44f3.pth)\|[log](https://bj.bcebos.com/paddleseg/dygraph/ade20k/pp_mobileseg_base/train.log) |
+| PP-MobileSeg-Tiny | StrideFormer-Tiny | 80000 | 32 | 512x512 | 36.39% | 215.3 | 1.61 | [config](https://github.com/Yang-Changhui/mmsegmentation/tree/add_ppmobileseg/projects/pp_mobileseg/configs/pp_mobileseg) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_2xb16_3rdparty-tiny_512x512-ade20k-a351ebf5.pth)\|[log](https://bj.bcebos.com/paddleseg/dygraph/ade20k/pp_mobileseg_tiny/train.log) |
+
+## Usage
+
+Same as other models in MMsegmentation, you can run the following command to test the model at ${MMSEG_ROOT}:
+
+```shell
+./tools/dist_test.sh projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py checkpoints/pp_mobileseg_mobilenetv3_2xb16_3rdparty-base_512x512-ade20k-f12b44f3.pth 8
+```
+
+## Inference with ONNXRuntime
+
+### Prerequisites
+
+**1. Install onnxruntime inference engine.**
+
+Choose one of the following ways to install onnxruntime.
+
+- CPU version
+
+```shell
+pip install onnxruntime==1.15.1
+wget https://github.com/microsoft/onnxruntime/releases/download/v1.15.1/onnxruntime-linux-x64-1.15.1.tgz
+tar -zxvf onnxruntime-linux-x64-1.15.1.tgz
+export ONNXRUNTIME_DIR=$(pwd)/onnxruntime-linux-x64-1.15.1
+export LD_LIBRARY_PATH=$ONNXRUNTIME_DIR/lib:$LD_LIBRARY_PATH
+```
+
+**2. Convert model to onnx file**
+
+- Install `mim` and `mmdeploy`.
+
+```shell
+pip install openmim
+mim install mmdeploy
+git clone https://github.com/open-mmlab/mmdeploy.git
+```
+
+- Download pp_mobileseg model.
+
+```shell
+wget https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_2xb16_3rdparty-tiny_512x512-ade20k-a351ebf5.pth
+```
+
+- Convert model to onnx files.
+
+```shell
+python mmdeploy/tools/deploy.py mmdeploy/configs/mmseg/segmentation_onnxruntime_dynamic.py \
+ configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py \
+ pp_mobileseg_mobilenetv3_2xb16_3rdparty-tiny_512x512-ade20k-a351ebf5.pth \
+ ../../demo/demo.png \
+ --work-dir mmdeploy_model/mmseg/ort \
+ --show
+```
+
+**3. Run demo**
+
+```shell
+python inference_onnx.py ${ONNX_FILE_PATH} ${IMAGE_PATH} [${MODEL_INPUT_SIZE} ${DEVICE} ${OUTPUT_IMAGE_PATH}]
+```
+
+Example:
+
+```shell
+python inference_onnx.py mmdeploy_model/mmseg/ort/end2end.onnx ../../demo/demo.png
+```
+
+## Citation
+
+If you find our project useful in your research, please consider citing:
+
+```
+@misc{liu2021paddleseg,
+ title={PaddleSeg: A High-Efficient Development Toolkit for Image Segmentation},
+ author={Yi Liu and Lutao Chu and Guowei Chen and Zewu Wu and Zeyu Chen and Baohua Lai and Yuying Hao},
+ year={2021},
+ eprint={2101.06175},
+ archivePrefix={arXiv},
+ primaryClass={cs.CV}
+}
+
+@misc{paddleseg2019,
+ title={PaddleSeg, End-to-end image segmentation kit based on PaddlePaddle},
+ author={PaddlePaddle Contributors},
+ howpublished = {\url{https://github.com/PaddlePaddle/PaddleSeg}},
+ year={2019}
+}
+```
diff --git a/projects/pp_mobileseg/backbones/__init__.py b/projects/pp_mobileseg/backbones/__init__.py
new file mode 100644
index 00000000000..244b33d37a8
--- /dev/null
+++ b/projects/pp_mobileseg/backbones/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .strideformer import StrideFormer
+
+__all__ = ['StrideFormer']
diff --git a/projects/pp_mobileseg/backbones/strideformer.py b/projects/pp_mobileseg/backbones/strideformer.py
new file mode 100644
index 00000000000..3f09be5225d
--- /dev/null
+++ b/projects/pp_mobileseg/backbones/strideformer.py
@@ -0,0 +1,958 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import math
+import warnings
+
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import ConvModule, build_activation_layer
+from mmcv.cnn.bricks.transformer import build_dropout
+from mmengine.logging import print_log
+from mmengine.model import BaseModule
+from mmengine.runner.checkpoint import CheckpointLoader, load_state_dict
+
+from mmseg.registry import MODELS
+
+
+@MODELS.register_module()
+class StrideFormer(BaseModule):
+ """The StrideFormer implementation based on torch.
+
+ The original article refers to:https://arxiv.org/abs/2304.05152
+ Args:
+ mobileV3_cfg(list): Each sublist describe the config for a
+ MobileNetV3 block.
+ channels(list): The input channels for each MobileNetV3 block.
+ embed_dims(list): The channels of the features input to the sea
+ attention block.
+ key_dims(list, optional): The embeding dims for each head in
+ attention.
+ depths(list, optional): describes the depth of the attention block.
+ i,e: M,N.
+ num_heads(int, optional): The number of heads of the attention
+ blocks.
+ attn_ratios(int, optional): The expand ratio of V.
+ mlp_ratios(list, optional): The ratio of mlp blocks.
+ drop_path_rate(float, optional): The drop path rate in attention
+ block.
+ act_cfg(dict, optional): The activation layer of AAM:
+ Aggregate Attention Module.
+ inj_type(string, optional): The type of injection/AAM.
+ out_channels(int, optional): The output channels of the AAM.
+ dims(list, optional): The dimension of the fusion block.
+ out_feat_chs(list, optional): The input channels of the AAM.
+ stride_attention(bool, optional): whether to stride attention in
+ each attention layer.
+ pretrained(str, optional): the path of pretrained model.
+ """
+
+ def __init__(
+ self,
+ mobileV3_cfg,
+ channels,
+ embed_dims,
+ key_dims=[16, 24],
+ depths=[2, 2],
+ num_heads=8,
+ attn_ratios=2,
+ mlp_ratios=[2, 4],
+ drop_path_rate=0.1,
+ act_cfg=dict(type='ReLU'),
+ inj_type='AAM',
+ out_channels=256,
+ dims=(128, 160),
+ out_feat_chs=None,
+ stride_attention=True,
+ pretrained=None,
+ init_cfg=None,
+ ):
+ super().__init__(init_cfg=init_cfg)
+ assert not (init_cfg and pretrained
+ ), 'init_cfg and pretrained cannot be set at the same time'
+ if isinstance(pretrained, str):
+ warnings.warn('DeprecationWarning: pretrained is deprecated, '
+ 'please use "init_cfg" instead')
+ self.init_cfg = dict(type='Pretrained', checkpoint=pretrained)
+ elif pretrained is not None:
+ raise TypeError('pretrained must be a str or None')
+
+ self.depths = depths
+ self.cfgs = mobileV3_cfg
+ self.dims = dims
+ for i in range(len(self.cfgs)):
+ smb = StackedMV3Block(
+ cfgs=self.cfgs[i],
+ stem=True if i == 0 else False,
+ in_channels=channels[i],
+ )
+ setattr(self, f'smb{i + 1}', smb)
+ for i in range(len(depths)):
+ dpr = [
+ x.item() for x in torch.linspace(0, drop_path_rate, depths[i])
+ ]
+ trans = BasicLayer(
+ block_num=depths[i],
+ embedding_dim=embed_dims[i],
+ key_dim=key_dims[i],
+ num_heads=num_heads,
+ mlp_ratio=mlp_ratios[i],
+ attn_ratio=attn_ratios,
+ drop=0,
+ attn_drop=0.0,
+ drop_path=dpr,
+ act_cfg=act_cfg,
+ stride_attention=stride_attention,
+ )
+ setattr(self, f'trans{i + 1}', trans)
+
+ self.inj_type = inj_type
+ if self.inj_type == 'AAM':
+ self.inj_module = InjectionMultiSumallmultiallsum(
+ in_channels=out_feat_chs, out_channels=out_channels)
+ self.feat_channels = [
+ out_channels,
+ ]
+ elif self.inj_type == 'AAMSx8':
+ self.inj_module = InjectionMultiSumallmultiallsumSimpx8(
+ in_channels=out_feat_chs, out_channels=out_channels)
+ self.feat_channels = [
+ out_channels,
+ ]
+ elif self.inj_type == 'origin':
+ for i in range(len(dims)):
+ fuse = FusionBlock(
+ out_feat_chs[0] if i == 0 else dims[i - 1],
+ out_feat_chs[i + 1],
+ embed_dim=dims[i],
+ act_cfg=None,
+ )
+ setattr(self, f'fuse{i + 1}', fuse)
+ self.feat_channels = [
+ dims[i],
+ ]
+ else:
+ raise NotImplementedError(self.inj_module + ' is not implemented')
+
+ self.pretrained = pretrained
+ # self.init_weights()
+
+ def init_weights(self):
+ if (isinstance(self.init_cfg, dict)
+ and self.init_cfg.get('type') == 'Pretrained'):
+ checkpoint = CheckpointLoader.load_checkpoint(
+ self.init_cfg['checkpoint'], logger=None, map_location='cpu')
+
+ if 'state_dict' in checkpoint:
+ state_dict = checkpoint['state_dict']
+ else:
+ state_dict = checkpoint
+
+ if 'pos_embed' in state_dict.keys():
+ if self.pos_embed.shape != state_dict['pos_embed'].shape:
+ print_log(msg=f'Resize the pos_embed shape from '
+ f'{state_dict["pos_embed"].shape} to '
+ f'{self.pos_embed.shape}')
+ h, w = self.img_size
+ pos_size = int(
+ math.sqrt(state_dict['pos_embed'].shape[1] - 1))
+ state_dict['pos_embed'] = self.resize_pos_embed(
+ state_dict['pos_embed'],
+ (h // self.patch_size, w // self.patch_size),
+ (pos_size, pos_size),
+ self.interpolate_mode,
+ )
+
+ load_state_dict(self, state_dict, strict=False, logger=None)
+
+ def forward(self, x):
+ x_hw = x.shape[2:]
+ outputs = []
+ num_smb_stage = len(self.cfgs)
+ num_trans_stage = len(self.depths)
+
+ for i in range(num_smb_stage):
+ smb = getattr(self, f'smb{i + 1}')
+ x = smb(x)
+
+ # 1/8 shared feat
+ if i == 1:
+ outputs.append(x)
+ if num_trans_stage + i >= num_smb_stage:
+ trans = getattr(
+ self, f'trans{i + num_trans_stage - num_smb_stage + 1}')
+ x = trans(x)
+ outputs.append(x)
+ if self.inj_type == 'origin':
+ x_detail = outputs[0]
+ for i in range(len(self.dims)):
+ fuse = getattr(self, f'fuse{i + 1}')
+
+ x_detail = fuse(x_detail, outputs[i + 1])
+ output = x_detail
+ else:
+ output = self.inj_module(outputs)
+
+ return [output, x_hw]
+
+
+class StackedMV3Block(nn.Module):
+ """The MobileNetV3 block.
+
+ Args:
+ cfgs (list): The MobileNetV3 config list of a stage.
+ stem (bool): Whether is the first stage or not.
+ in_channels (int, optional): The channels of input image. Default: 3.
+ scale: float=1.0.
+ The coefficient that controls the size of network parameters.
+
+ Returns:
+ model: nn.Module.
+ A stage of specific MobileNetV3 model depends on args.
+ """
+
+ def __init__(self,
+ cfgs,
+ stem,
+ in_channels,
+ scale=1.0,
+ norm_cfg=dict(type='BN')):
+ super().__init__()
+
+ self.scale = scale
+ self.stem = stem
+
+ if self.stem:
+ self.conv = ConvModule(
+ in_channels=3,
+ out_channels=_make_divisible(in_channels * self.scale),
+ kernel_size=3,
+ stride=2,
+ padding=1,
+ groups=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=dict(type='HSwish'),
+ )
+
+ self.blocks = nn.ModuleList()
+ for i, (k, exp, c, se, act, s) in enumerate(cfgs):
+ self.blocks.append(
+ ResidualUnit(
+ in_channel=_make_divisible(in_channels * self.scale),
+ mid_channel=_make_divisible(self.scale * exp),
+ out_channel=_make_divisible(self.scale * c),
+ kernel_size=k,
+ stride=s,
+ use_se=se,
+ act=act,
+ dilation=1,
+ ))
+ in_channels = _make_divisible(self.scale * c)
+
+ def forward(self, x):
+ if self.stem:
+ x = self.conv(x)
+ for i, block in enumerate(self.blocks):
+ x = block(x)
+
+ return x
+
+
+class ResidualUnit(nn.Module):
+ """The Residual module.
+
+ Args:
+ in_channel (int, optional): The channels of input feature.
+ mid_channel (int, optional): The channels of middle process.
+ out_channel (int, optional): The channels of output feature.
+ kernel_size (int, optional): The size of the convolving kernel.
+ stride (int, optional): The stride size.
+ use_se (bool, optional): if to use the SEModule.
+ act (string, optional): activation layer.
+ dilation (int, optional): The dilation size.
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN', requires_grad=True).
+ """
+
+ def __init__(
+ self,
+ in_channel,
+ mid_channel,
+ out_channel,
+ kernel_size,
+ stride,
+ use_se,
+ act=None,
+ dilation=1,
+ norm_cfg=dict(type='BN'),
+ ):
+ super().__init__()
+ self.if_shortcut = stride == 1 and in_channel == out_channel
+ self.if_se = use_se
+ self.expand_conv = ConvModule(
+ in_channels=in_channel,
+ out_channels=mid_channel,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=dict(type=act) if act is not None else None,
+ )
+ self.bottleneck_conv = ConvModule(
+ in_channels=mid_channel,
+ out_channels=mid_channel,
+ kernel_size=kernel_size,
+ stride=stride,
+ padding=int((kernel_size - 1) // 2) * dilation,
+ bias=False,
+ groups=mid_channel,
+ dilation=dilation,
+ norm_cfg=norm_cfg,
+ act_cfg=dict(type=act) if act is not None else None,
+ )
+ if self.if_se:
+ self.mid_se = SEModule(mid_channel)
+ self.linear_conv = ConvModule(
+ in_channels=mid_channel,
+ out_channels=out_channel,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+
+ def forward(self, x):
+ identity = x
+ x = self.expand_conv(x)
+ x = self.bottleneck_conv(x)
+ if self.if_se:
+ x = self.mid_se(x)
+ x = self.linear_conv(x)
+ if self.if_shortcut:
+ x = torch.add(identity, x)
+ return x
+
+
+class SEModule(nn.Module):
+ """SE Module.
+
+ Args:
+ channel (int, optional): The channels of input feature.
+ reduction (int, optional): The channel reduction rate.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ """
+
+ def __init__(self, channel, reduction=4, act_cfg=dict(type='ReLU')):
+ super().__init__()
+ self.avg_pool = nn.AdaptiveAvgPool2d(1)
+ self.conv_act1 = ConvModule(
+ in_channels=channel,
+ out_channels=channel // reduction,
+ kernel_size=1,
+ norm_cfg=None,
+ act_cfg=act_cfg,
+ )
+
+ self.conv_act2 = ConvModule(
+ in_channels=channel // reduction,
+ out_channels=channel,
+ kernel_size=1,
+ norm_cfg=None,
+ act_cfg=dict(type='Hardsigmoid', slope=0.2, offset=0.5),
+ )
+
+ def forward(self, x):
+ identity = x
+ x = self.avg_pool(x)
+ x = self.conv_act1(x)
+ x = self.conv_act2(x)
+ return torch.mul(identity, x)
+
+
+class BasicLayer(nn.Module):
+ """The transformer basic layer.
+
+ Args:
+ block_num (int): the block nums of the transformer basic layer.
+ embedding_dim (int): The feature dimension.
+ key_dim (int): the key dim.
+ num_heads (int): Parallel attention heads.
+ mlp_ratio (float): the mlp ratio.
+ attn_ratio (float): the attention ratio.
+ drop (float): Probability of an element to be zeroed
+ after the feed forward layer.Default: 0.0.
+ attn_drop (float): The drop out rate for attention layer.
+ Default: 0.0.
+ drop_path (float): stochastic depth rate. Default 0.0.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ stride_attention (bool, optional): whether to stride attention in
+ each attention layer.
+ """
+
+ def __init__(
+ self,
+ block_num,
+ embedding_dim,
+ key_dim,
+ num_heads,
+ mlp_ratio=4.0,
+ attn_ratio=2.0,
+ drop=0.0,
+ attn_drop=0.0,
+ drop_path=None,
+ act_cfg=None,
+ stride_attention=None,
+ ):
+ super().__init__()
+ self.block_num = block_num
+
+ self.transformer_blocks = nn.ModuleList()
+ for i in range(self.block_num):
+ self.transformer_blocks.append(
+ Block(
+ embedding_dim,
+ key_dim=key_dim,
+ num_heads=num_heads,
+ mlp_ratio=mlp_ratio,
+ attn_ratio=attn_ratio,
+ drop=drop,
+ drop_path=drop_path[i]
+ if isinstance(drop_path, list) else drop_path,
+ act_cfg=act_cfg,
+ stride_attention=stride_attention,
+ ))
+
+ def forward(self, x):
+ for i in range(self.block_num):
+ x = self.transformer_blocks[i](x)
+ return x
+
+
+class Block(nn.Module):
+ """the block of the transformer basic layer.
+
+ Args:
+ dim (int): The feature dimension.
+ key_dim (int): The key dimension.
+ num_heads (int): Parallel attention heads.
+ mlp_ratio (float): the mlp ratio.
+ attn_ratio (float): the attention ratio.
+ drop (float): Probability of an element to be zeroed
+ after the feed forward layer.Default: 0.0.
+ drop_path (float): stochastic depth rate. Default 0.0.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ stride_attention (bool, optional): whether to stride attention in
+ each attention layer.
+ """
+
+ def __init__(
+ self,
+ dim,
+ key_dim,
+ num_heads,
+ mlp_ratio=4.0,
+ attn_ratio=2.0,
+ drop=0.0,
+ drop_path=0.0,
+ act_cfg=None,
+ stride_attention=None,
+ ):
+ super().__init__()
+ self.dim = dim
+ self.num_heads = num_heads
+ self.mlp_ratio = mlp_ratio
+ self.attn = SeaAttention(
+ dim,
+ key_dim=key_dim,
+ num_heads=num_heads,
+ attn_ratio=attn_ratio,
+ act_cfg=act_cfg,
+ stride_attention=stride_attention,
+ )
+ self.drop_path = (
+ build_dropout(dict(type='DropPath', drop_prob=drop_path))
+ if drop_path > 0.0 else nn.Identity())
+ mlp_hidden_dim = int(dim * mlp_ratio)
+ self.mlp = MLP(
+ in_features=dim,
+ hidden_features=mlp_hidden_dim,
+ act_cfg=act_cfg,
+ drop=drop,
+ )
+
+ def forward(self, x1):
+ x1 = x1 + self.drop_path(self.attn(x1))
+ x1 = x1 + self.drop_path(self.mlp(x1))
+
+ return x1
+
+
+class SqueezeAxialPositionalEmbedding(nn.Module):
+ """the Squeeze Axial Positional Embedding.
+
+ Args:
+ dim (int): The feature dimension.
+ shape (int): The patch size.
+ """
+
+ def __init__(self, dim, shape):
+ super().__init__()
+ self.pos_embed = nn.init.normal_(
+ nn.Parameter(torch.zeros(1, dim, shape)))
+
+ def forward(self, x):
+ B, C, N = x.shape
+ x = x + F.interpolate(
+ self.pos_embed, size=(N, ), mode='linear', align_corners=False)
+ return x
+
+
+class SeaAttention(nn.Module):
+ """The sea attention.
+
+ Args:
+ dim (int): The feature dimension.
+ key_dim (int): The key dimension.
+ num_heads (int): number of attention heads.
+ attn_ratio (float): the attention ratio.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='LN')
+ stride_attention (bool, optional): whether to stride attention in
+ each attention layer.
+ """
+
+ def __init__(
+ self,
+ dim,
+ key_dim,
+ num_heads,
+ attn_ratio=4.0,
+ act_cfg=None,
+ norm_cfg=dict(type='BN'),
+ stride_attention=False,
+ ):
+
+ super().__init__()
+ self.num_heads = num_heads
+ self.scale = key_dim**-0.5
+ self.nh_kd = nh_kd = key_dim * num_heads
+ self.d = int(attn_ratio * key_dim)
+ self.dh = int(attn_ratio * key_dim) * num_heads
+ self.attn_ratio = attn_ratio
+
+ self.to_q = ConvModule(
+ dim, nh_kd, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+ self.to_k = ConvModule(
+ dim, nh_kd, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+
+ self.to_v = ConvModule(
+ dim, self.dh, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+ self.stride_attention = stride_attention
+ if self.stride_attention:
+ self.stride_conv = ConvModule(
+ dim,
+ dim,
+ kernel_size=3,
+ stride=2,
+ padding=1,
+ bias=True,
+ groups=dim,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+
+ self.proj = ConvModule(
+ self.dh,
+ dim,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ order=('act', 'conv', 'norm'),
+ )
+ self.proj_encode_row = ConvModule(
+ self.dh,
+ self.dh,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ order=('act', 'conv', 'norm'),
+ )
+ self.pos_emb_rowq = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.pos_emb_rowk = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.proj_encode_column = ConvModule(
+ self.dh,
+ self.dh,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ order=('act', 'conv', 'norm'),
+ )
+ self.pos_emb_columnq = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.pos_emb_columnk = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.dwconv = ConvModule(
+ 2 * self.dh,
+ 2 * self.dh,
+ 3,
+ padding=1,
+ groups=2 * self.dh,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ )
+ self.pwconv = ConvModule(
+ 2 * self.dh, dim, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+ self.sigmoid = build_activation_layer(dict(type='HSigmoid'))
+
+ def forward(self, x):
+ B, C, H_ori, W_ori = x.shape
+ if self.stride_attention:
+ x = self.stride_conv(x)
+ B, C, H, W = x.shape
+
+ q = self.to_q(x) # [B, nhead*dim, H, W]
+ k = self.to_k(x)
+ v = self.to_v(x)
+
+ qkv = torch.cat([q, k, v], dim=1)
+ qkv = self.dwconv(qkv)
+ qkv = self.pwconv(qkv)
+
+ qrow = (self.pos_emb_rowq(q.mean(-1)).reshape(
+ [B, self.num_heads, -1, H]).permute(
+ (0, 1, 3, 2))) # [B, nhead, H, dim]
+ krow = self.pos_emb_rowk(k.mean(-1)).reshape(
+ [B, self.num_heads, -1, H]) # [B, nhead, dim, H]
+ vrow = (v.mean(-1).reshape([B, self.num_heads, -1,
+ H]).permute([0, 1, 3, 2])
+ ) # [B, nhead, H, dim*attn_ratio]
+
+ attn_row = torch.matmul(qrow, krow) * self.scale # [B, nhead, H, H]
+ attn_row = nn.functional.softmax(attn_row, dim=-1)
+
+ xx_row = torch.matmul(attn_row, vrow) # [B, nhead, H, dim*attn_ratio]
+ xx_row = self.proj_encode_row(
+ xx_row.permute([0, 1, 3, 2]).reshape([B, self.dh, H, 1]))
+
+ # squeeze column
+ qcolumn = (
+ self.pos_emb_columnq(q.mean(-2)).reshape(
+ [B, self.num_heads, -1, W]).permute([0, 1, 3, 2]))
+ kcolumn = self.pos_emb_columnk(k.mean(-2)).reshape(
+ [B, self.num_heads, -1, W])
+ vcolumn = (
+ torch.mean(v, -2).reshape([B, self.num_heads, -1,
+ W]).permute([0, 1, 3, 2]))
+
+ attn_column = torch.matmul(qcolumn, kcolumn) * self.scale
+ attn_column = nn.functional.softmax(attn_column, dim=-1)
+
+ xx_column = torch.matmul(attn_column, vcolumn) # B nH W C
+ xx_column = self.proj_encode_column(
+ xx_column.permute([0, 1, 3, 2]).reshape([B, self.dh, 1, W]))
+
+ xx = torch.add(xx_row, xx_column) # [B, self.dh, H, W]
+ xx = torch.add(v, xx)
+
+ xx = self.proj(xx)
+ xx = self.sigmoid(xx) * qkv
+ if self.stride_attention:
+ xx = F.interpolate(xx, size=(H_ori, W_ori), mode='bilinear')
+
+ return xx
+
+
+class MLP(nn.Module):
+ """the Multilayer Perceptron.
+
+ Args:
+ in_features (int): the input feature.
+ hidden_features (int): the hidden feature.
+ out_features (int): the output feature.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ drop (float): Probability of an element to be zeroed.
+ Default 0.0
+ """
+
+ def __init__(
+ self,
+ in_features,
+ hidden_features=None,
+ out_features=None,
+ act_cfg=None,
+ norm_cfg=dict(type='BN'),
+ drop=0.0,
+ ):
+ super().__init__()
+ out_features = out_features or in_features
+ hidden_features = hidden_features or in_features
+ self.fc1 = ConvModule(
+ in_features,
+ hidden_features,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+ self.dwconv = ConvModule(
+ hidden_features,
+ hidden_features,
+ kernel_size=3,
+ padding=1,
+ groups=hidden_features,
+ norm_cfg=None,
+ act_cfg=act_cfg,
+ )
+
+ self.fc2 = ConvModule(
+ hidden_features,
+ out_features,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+ self.drop = build_dropout(dict(type='Dropout', drop_prob=drop))
+
+ def forward(self, x):
+ x = self.fc1(x)
+ x = self.dwconv(x)
+ x = self.drop(x)
+ x = self.fc2(x)
+ x = self.drop(x)
+ return x
+
+
+class FusionBlock(nn.Module):
+ """The feature fusion block.
+
+ Args:
+ in_channel (int): the input channel.
+ out_channel (int): the output channel.
+ embed_dim (int): embedding dimension.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ """
+
+ def __init__(
+ self,
+ in_channel,
+ out_channel,
+ embed_dim,
+ norm_cfg=dict(type='BN'),
+ act_cfg=dict(type='ReLU'),
+ ) -> None:
+ super().__init__()
+ self.local_embedding = ConvModule(
+ in_channels=in_channel,
+ out_channels=embed_dim,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+
+ self.global_act = ConvModule(
+ in_channels=out_channel,
+ out_channels=embed_dim,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg if act_cfg is not None else None,
+ )
+
+ def forward(self, x_l, x_g):
+ """
+ x_g: global features
+ x_l: local features
+ """
+ B, C, H, W = x_l.shape
+
+ local_feat = self.local_embedding(x_l)
+ global_act = self.global_act(x_g)
+ sig_act = F.interpolate(
+ global_act, size=(H, W), mode='bilinear', align_corners=False)
+
+ out = local_feat * sig_act
+
+ return out
+
+
+class InjectionMultiSumallmultiallsum(nn.Module):
+ """the Aggregate Attention Module.
+
+ Args:
+ in_channels (tuple): the input channel.
+ out_channels (int): the output channel.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ """
+
+ def __init__(
+ self,
+ in_channels=(64, 128, 256, 384),
+ out_channels=256,
+ act_cfg=dict(type='Sigmoid'),
+ norm_cfg=dict(type='BN'),
+ ):
+ super().__init__()
+ self.embedding_list = nn.ModuleList()
+ self.act_embedding_list = nn.ModuleList()
+ self.act_list = nn.ModuleList()
+ for i in range(len(in_channels)):
+ self.embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ ))
+ self.act_embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ))
+
+ def forward(self, inputs): # x_x8, x_x16, x_x32, x_x64
+ low_feat1 = F.interpolate(inputs[0], scale_factor=0.5, mode='bilinear')
+ low_feat1_act = self.act_embedding_list[0](low_feat1)
+ low_feat1 = self.embedding_list[0](low_feat1)
+
+ low_feat2 = F.interpolate(
+ inputs[1], size=low_feat1.shape[-2:], mode='bilinear')
+ low_feat2_act = self.act_embedding_list[1](low_feat2) # x16
+ low_feat2 = self.embedding_list[1](low_feat2)
+
+ high_feat_act = F.interpolate(
+ self.act_embedding_list[2](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear',
+ )
+ high_feat = F.interpolate(
+ self.embedding_list[2](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear')
+
+ res = (
+ low_feat1_act * low_feat2_act * high_feat_act *
+ (low_feat1 + low_feat2) + high_feat)
+
+ return res
+
+
+class InjectionMultiSumallmultiallsumSimpx8(nn.Module):
+ """the Aggregate Attention Module.
+
+ Args:
+ in_channels (tuple): the input channel.
+ out_channels (int): the output channel.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ """
+
+ def __init__(
+ self,
+ in_channels=(64, 128, 256, 384),
+ out_channels=256,
+ act_cfg=dict(type='Sigmoid'),
+ norm_cfg=dict(type='BN'),
+ ):
+ super().__init__()
+ self.embedding_list = nn.ModuleList()
+ self.act_embedding_list = nn.ModuleList()
+ self.act_list = nn.ModuleList()
+ for i in range(len(in_channels)):
+ if i != 1:
+ self.embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ ))
+ if i != 0:
+ self.act_embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ))
+
+ def forward(self, inputs):
+ # x_x8, x_x16, x_x32
+ low_feat1 = self.embedding_list[0](inputs[0])
+
+ low_feat2 = F.interpolate(
+ inputs[1], size=low_feat1.shape[-2:], mode='bilinear')
+ low_feat2_act = self.act_embedding_list[0](low_feat2)
+
+ high_feat_act = F.interpolate(
+ self.act_embedding_list[1](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear',
+ )
+ high_feat = F.interpolate(
+ self.embedding_list[1](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear')
+
+ res = low_feat2_act * high_feat_act * low_feat1 + high_feat
+
+ return res
+
+
+def _make_divisible(v, divisor=8, min_value=None):
+ if min_value is None:
+ min_value = divisor
+ new_v = max(min_value, int(v + divisor / 2) // divisor * divisor)
+ if new_v < 0.9 * v:
+ new_v += divisor
+ return new_v
+
+
+@MODELS.register_module()
+class Hardsigmoid(nn.Module):
+ """the hardsigmoid activation.
+
+ Args:
+ slope (float, optional): The slope of hardsigmoid function.
+ Default is 0.1666667.
+ offset (float, optional): The offset of hardsigmoid function.
+ Default is 0.5.
+ inplace (bool): can optionally do the operation in-place.
+ Default: ``False``
+ """
+
+ def __init__(self, slope=0.1666667, offset=0.5, inplace=False):
+ super().__init__()
+ self.slope = slope
+ self.offset = offset
+
+ def forward(self, x):
+ return (x * self.slope + self.offset).clamp(0, 1)
diff --git a/projects/pp_mobileseg/configs/_base_/datasets/ade20k.py b/projects/pp_mobileseg/configs/_base_/datasets/ade20k.py
new file mode 100644
index 00000000000..48340d11eea
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/datasets/ade20k.py
@@ -0,0 +1,68 @@
+# dataset settings
+dataset_type = 'ADE20KDataset'
+data_root = 'data/ade/ADEChallengeData2016'
+crop_size = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations', reduce_zero_label=True),
+ dict(
+ type='RandomResize',
+ scale=(2048, 512),
+ ratio_range=(0.5, 2.0),
+ keep_ratio=True),
+ dict(type='RandomCrop', crop_size=crop_size, cat_max_ratio=0.75),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 512), keep_ratio=True),
+ # add loading annotation after ``Resize`` because ground truth
+ # does not need to do resize data transform
+ dict(type='LoadAnnotations', reduce_zero_label=True),
+ dict(type='PackSegInputs')
+]
+img_ratios = [0.5, 0.75, 1.0, 1.25, 1.5, 1.75]
+tta_pipeline = [
+ dict(type='LoadImageFromFile', backend_args=None),
+ dict(
+ type='TestTimeAug',
+ transforms=[
+ [
+ dict(type='Resize', scale_factor=r, keep_ratio=True)
+ for r in img_ratios
+ ],
+ [
+ dict(type='RandomFlip', prob=0., direction='horizontal'),
+ dict(type='RandomFlip', prob=1., direction='horizontal')
+ ], [dict(type='LoadAnnotations')], [dict(type='PackSegInputs')]
+ ])
+]
+train_dataloader = dict(
+ batch_size=4,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/training', seg_map_path='annotations/training'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/validation',
+ seg_map_path='annotations/validation'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU'])
+test_evaluator = val_evaluator
diff --git a/projects/pp_mobileseg/configs/_base_/default_runtime.py b/projects/pp_mobileseg/configs/_base_/default_runtime.py
new file mode 100644
index 00000000000..272b4d24679
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/default_runtime.py
@@ -0,0 +1,15 @@
+default_scope = 'mmseg'
+env_cfg = dict(
+ cudnn_benchmark=True,
+ mp_cfg=dict(mp_start_method='fork', opencv_num_threads=0),
+ dist_cfg=dict(backend='nccl'),
+)
+vis_backends = [dict(type='LocalVisBackend')]
+visualizer = dict(
+ type='SegLocalVisualizer', vis_backends=vis_backends, name='visualizer')
+log_processor = dict(by_epoch=False)
+log_level = 'INFO'
+load_from = None
+resume = False
+
+tta_model = dict(type='SegTTAModel')
diff --git a/projects/pp_mobileseg/configs/_base_/models/pp_mobile.py b/projects/pp_mobileseg/configs/_base_/models/pp_mobile.py
new file mode 100644
index 00000000000..0c7695636f6
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/models/pp_mobile.py
@@ -0,0 +1,47 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255)
+
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ # pretrained='open-mmlab://resnet50_v1c',
+ backbone=dict(
+ type='StrideFormer',
+ mobileV3_cfg=[
+ # k t c, s
+ [[3, 16, 16, True, 'ReLU', 1], [3, 64, 32, False, 'ReLU', 2],
+ [3, 96, 32, False, 'ReLU', 1]], # cfg1
+ [[5, 128, 64, True, 'HSwish', 2], [5, 240, 64, True, 'HSwish',
+ 1]], # cfg2
+ [[5, 384, 128, True, 'HSwish', 2],
+ [5, 384, 128, True, 'HSwish', 1]], # cfg3
+ [[5, 768, 192, True, 'HSwish', 2],
+ [5, 768, 192, True, 'HSwish', 1]], # cfg4
+ ],
+ channels=[16, 32, 64, 128, 192],
+ depths=[3, 3],
+ embed_dims=[128, 192],
+ num_heads=8,
+ inj_type='AAMSx8',
+ out_feat_chs=[64, 128, 192],
+ act_cfg=dict(type='ReLU6'),
+ ),
+ decode_head=dict(
+ type='PPMobileSegHead',
+ num_classes=150,
+ in_channels=256,
+ dropout_ratio=0.1,
+ use_dw=True,
+ act_cfg=dict(type='ReLU'),
+ align_corners=False),
+
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/pp_mobileseg/configs/_base_/schedules/schedule_80k.py b/projects/pp_mobileseg/configs/_base_/schedules/schedule_80k.py
new file mode 100644
index 00000000000..0dcd6c4d1bc
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/schedules/schedule_80k.py
@@ -0,0 +1,24 @@
+# optimizer
+optimizer = dict(type='SGD', lr=0.01, momentum=0.9, weight_decay=0.0005)
+optim_wrapper = dict(type='OptimWrapper', optimizer=optimizer, clip_grad=None)
+# learning policy
+param_scheduler = [
+ dict(
+ type='PolyLR',
+ eta_min=1e-4,
+ power=0.9,
+ begin=0,
+ end=80000,
+ by_epoch=False)
+]
+# training schedule for 80k
+train_cfg = dict(type='IterBasedTrainLoop', max_iters=80000, val_interval=8000)
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
+default_hooks = dict(
+ timer=dict(type='IterTimerHook'),
+ logger=dict(type='LoggerHook', interval=50, log_metric_by_epoch=False),
+ param_scheduler=dict(type='ParamSchedulerHook'),
+ checkpoint=dict(type='CheckpointHook', by_epoch=False, interval=8000),
+ sampler_seed=dict(type='DistSamplerSeedHook'),
+ visualization=dict(type='SegVisualizationHook'))
diff --git a/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py
new file mode 100644
index 00000000000..4b68a927e20
--- /dev/null
+++ b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py
@@ -0,0 +1,18 @@
+_base_ = [
+ '../_base_/models/pp_mobile.py', '../_base_/datasets/ade20k.py',
+ '../_base_/default_runtime.py', '../_base_/schedules/schedule_80k.py'
+]
+# the custom import path is determined by your workspace path (i.e., where you run the command from) # noqa
+custom_imports = dict(
+ imports=[
+ 'projects.pp_mobileseg.backbones', 'projects.pp_mobileseg.decode_head'
+ ],
+ allow_failed_imports=False)
+checkpoint = 'https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_3rdparty-base-ed0be681.pth' # noqa
+crop_size = (512, 512)
+data_preprocessor = dict(size=crop_size, test_cfg=dict(size_divisor=32))
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(init_cfg=dict(type='Pretrained', checkpoint=checkpoint)),
+ decode_head=dict(num_classes=150, upsample='intepolate'))
diff --git a/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py
new file mode 100644
index 00000000000..b78869e517a
--- /dev/null
+++ b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py
@@ -0,0 +1,45 @@
+_base_ = [
+ '../_base_/models/pp_mobile.py', '../_base_/datasets/ade20k.py',
+ '../_base_/default_runtime.py', '../_base_/schedules/schedule_80k.py'
+]
+# the custom import path is determined by your workspace path (i.e., where you run the command from) # noqa
+custom_imports = dict(
+ imports=[
+ 'projects.pp_mobileseg.backbones', 'projects.pp_mobileseg.decode_head'
+ ],
+ allow_failed_imports=False)
+checkpoint = 'https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_3rdparty-tiny-e4b35e96.pth' # noqa
+crop_size = (512, 512)
+data_preprocessor = dict(size=crop_size, test_cfg=dict(size_divisor=32))
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ init_cfg=dict(type='Pretrained', checkpoint=checkpoint),
+ type='StrideFormer',
+ mobileV3_cfg=[
+ # k t c, s
+ [[3, 16, 16, True, 'ReLU', 1], [3, 64, 32, False, 'ReLU', 2],
+ [3, 48, 24, False, 'ReLU', 1]], # cfg1
+ [[5, 96, 32, True, 'HSwish', 2], [5, 96, 32, True, 'HSwish',
+ 1]], # cfg2
+ [[5, 160, 64, True, 'HSwish', 2], [5, 160, 64, True, 'HSwish',
+ 1]], # cfg3
+ [[3, 384, 128, True, 'HSwish', 2],
+ [3, 384, 128, True, 'HSwish', 1]], # cfg4
+ ],
+ channels=[16, 24, 32, 64, 128],
+ depths=[2, 2],
+ embed_dims=[64, 128],
+ num_heads=4,
+ inj_type='AAM',
+ out_feat_chs=[32, 64, 128],
+ act_cfg=dict(type='ReLU6'),
+ ),
+ decode_head=dict(
+ num_classes=150,
+ in_channels=256,
+ use_dw=True,
+ act_cfg=dict(type='ReLU'),
+ upsample='intepolate'),
+)
diff --git a/projects/pp_mobileseg/decode_head/__init__.py b/projects/pp_mobileseg/decode_head/__init__.py
new file mode 100644
index 00000000000..6f71b784e15
--- /dev/null
+++ b/projects/pp_mobileseg/decode_head/__init__.py
@@ -0,0 +1,6 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .pp_mobileseg_head import PPMobileSegHead
+
+__all__ = [
+ 'PPMobileSegHead',
+]
diff --git a/projects/pp_mobileseg/decode_head/pp_mobileseg_head.py b/projects/pp_mobileseg/decode_head/pp_mobileseg_head.py
new file mode 100644
index 00000000000..243f0263729
--- /dev/null
+++ b/projects/pp_mobileseg/decode_head/pp_mobileseg_head.py
@@ -0,0 +1,94 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from typing import List
+
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import ConvModule, build_conv_layer
+from torch import Tensor
+
+from mmseg.registry import MODELS
+
+
+@MODELS.register_module()
+class PPMobileSegHead(nn.Module):
+ """the segmentation head.
+
+ Args:
+ num_classes (int): the classes num.
+ in_channels (int): the input channels.
+ use_dw (bool): if to use deepwith convolution.
+ dropout_ratio (float): Probability of an element to be zeroed.
+ Default 0.0。
+ align_corners (bool, optional): Geometrically, we consider the pixels
+ of the input and output as squares rather than points.
+ upsample (str): the upsample method.
+ out_channels (int): the output channel.
+ conv_cfg (dict): Config dict for convolution layer.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN').
+ """
+
+ def __init__(self,
+ num_classes,
+ in_channels,
+ use_dw=True,
+ dropout_ratio=0.1,
+ align_corners=False,
+ upsample='intepolate',
+ out_channels=None,
+ conv_cfg=dict(type='Conv'),
+ act_cfg=dict(type='ReLU'),
+ norm_cfg=dict(type='BN')):
+
+ super().__init__()
+ self.align_corners = align_corners
+ self.last_channels = in_channels
+ self.upsample = upsample
+ self.num_classes = num_classes
+ self.out_channels = out_channels
+ self.linear_fuse = ConvModule(
+ in_channels=self.last_channels,
+ out_channels=self.last_channels,
+ kernel_size=1,
+ bias=False,
+ groups=self.last_channels if use_dw else 1,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg)
+ self.dropout = nn.Dropout2d(dropout_ratio)
+ self.conv_seg = build_conv_layer(
+ conv_cfg, self.last_channels, self.num_classes, kernel_size=1)
+
+ def forward(self, x):
+ x, x_hw = x[0], x[1]
+ x = self.linear_fuse(x)
+ x = self.dropout(x)
+ x = self.conv_seg(x)
+ if self.upsample == 'intepolate' or self.training or \
+ self.num_classes < 30:
+ x = F.interpolate(
+ x, x_hw, mode='bilinear', align_corners=self.align_corners)
+ elif self.upsample == 'vim':
+ labelset = torch.unique(torch.argmax(x, 1))
+ x = torch.gather(x, 1, labelset)
+ x = F.interpolate(
+ x, x_hw, mode='bilinear', align_corners=self.align_corners)
+
+ pred = torch.argmax(x, 1)
+ pred_retrieve = torch.zeros(pred.shape, dtype=torch.int32)
+ for i, val in enumerate(labelset):
+ pred_retrieve[pred == i] = labelset[i].cast('int32')
+
+ x = pred_retrieve
+ else:
+ raise NotImplementedError(self.upsample, ' is not implemented')
+
+ return [x]
+
+ def predict(self, inputs, batch_img_metas: List[dict], test_cfg,
+ **kwargs) -> List[Tensor]:
+ """Forward function for testing, only ``pam_cam`` is used."""
+ seg_logits = self.forward(inputs)[0]
+ return seg_logits
diff --git a/projects/pp_mobileseg/inference_onnx.py b/projects/pp_mobileseg/inference_onnx.py
new file mode 100644
index 00000000000..139d1b13243
--- /dev/null
+++ b/projects/pp_mobileseg/inference_onnx.py
@@ -0,0 +1,203 @@
+import argparse
+import time
+from typing import List, Tuple
+
+import cv2
+import loguru
+import numpy as np
+import onnxruntime as ort
+
+logger = loguru.logger
+
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='PP_Mobileseg ONNX inference demo.')
+ parser.add_argument('onnx_file', help='ONNX file path')
+ parser.add_argument('image_file', help='Input image file path')
+ parser.add_argument(
+ '--input-size',
+ type=int,
+ nargs='+',
+ default=[512, 512],
+ help='input image size')
+ parser.add_argument(
+ '--device', help='device type for inference', default='cpu')
+ parser.add_argument(
+ '--save-path',
+ help='path to save the output image',
+ default='output.jpg')
+ args = parser.parse_args()
+ return args
+
+
+def preprocess(
+ img: np.ndarray, input_size: Tuple[int, int] = (512, 512)
+) -> Tuple[np.ndarray, np.ndarray]:
+ """Preprocess image for inference."""
+ img_shape = img.shape[:2]
+ # Resize
+ resized_img = cv2.resize(img, input_size)
+
+ # Normalize
+ mean = np.array([123.575, 116.28, 103.53], dtype=np.float32)
+ std = np.array([58.395, 57.12, 57.375], dtype=np.float32)
+ resized_img = (resized_img - mean) / std
+
+ return resized_img, img_shape
+
+
+def build_session(onnx_file: str, device: str = 'cpu') -> ort.InferenceSession:
+ """Build onnxruntime session.
+
+ Args:
+ onnx_file (str): ONNX file path.
+ device (str): Device type for inference.
+
+ Returns:
+ sess (ort.InferenceSession): ONNXRuntime session.
+ """
+ providers = ['CPUExecutionProvider'
+ ] if device == 'cpu' else ['CUDAExecutionProvider']
+ sess = ort.InferenceSession(path_or_bytes=onnx_file, providers=providers)
+
+ return sess
+
+
+def inference(sess: ort.InferenceSession, img: np.ndarray) -> np.ndarray:
+ """Inference RTMPose model.
+
+ Args:
+ sess (ort.InferenceSession): ONNXRuntime session.
+ img (np.ndarray): Input image in shape.
+
+ Returns:
+ outputs (np.ndarray): Output of RTMPose model.
+ """
+ # build input
+ input_img = [img.transpose(2, 0, 1).astype(np.float32)]
+
+ # build output
+ sess_input = {sess.get_inputs()[0].name: input_img}
+ sess_output = []
+ for out in sess.get_outputs():
+ sess_output.append(out.name)
+
+ # inference
+ outputs = sess.run(output_names=sess_output, input_feed=sess_input)
+
+ return outputs
+
+
+def postprocess(outputs: List[np.ndarray],
+ origin_shape: Tuple[int, int]) -> np.ndarray:
+ """Postprocess outputs of PP_Mobileseg model.
+
+ Args:
+ outputs (List[np.ndarray]): Outputs of PP_Mobileseg model.
+ origin_shape (Tuple[int, int]): Input size of PP_Mobileseg model.
+
+ Returns:
+ seg_map (np.ndarray): Segmentation map.
+ """
+ seg_map = outputs[0][0][0]
+ seg_map = cv2.resize(seg_map.astype(np.float32), origin_shape)
+ return seg_map
+
+
+def visualize(img: np.ndarray,
+ seg_map: np.ndarray,
+ filename: str = 'output.jpg',
+ opacity: float = 0.8) -> np.ndarray:
+ assert 0.0 <= opacity <= 1.0, 'opacity should be in range [0, 1]'
+ palette = np.array(PALETTE)
+ color_seg = np.zeros((seg_map.shape[0], seg_map.shape[1], 3),
+ dtype=np.uint8)
+ for label, color in enumerate(palette):
+ color_seg[seg_map == label, :] = color
+ # convert to BGR
+ color_seg = color_seg[..., ::-1]
+
+ img = img * (1 - opacity) + color_seg * opacity
+ cv2.imwrite(filename, img)
+
+ return img
+
+
+def main():
+ args = parse_args()
+ logger.info('Start running model inference...')
+
+ # read image from file
+ logger.info(f'1. Read image from file {args.image_file}...')
+ img = cv2.imread(args.image_file)
+
+ # build onnx model
+ logger.info(f'2. Build onnx model from {args.onnx_file}...')
+ sess = build_session(args.onnx_file, args.device)
+
+ # preprocess
+ logger.info('3. Preprocess image...')
+ model_input_size = tuple(args.input_size)
+ assert len(model_input_size) == 2
+ resized_img, origin_shape = preprocess(img, model_input_size)
+
+ # inference
+ logger.info('4. Inference...')
+ start = time.time()
+ outputs = inference(sess, resized_img)
+ logger.info(f'Inference time: {time.time() - start:.4f}s')
+
+ # postprocess
+ logger.info('5. Postprocess...')
+ h, w = origin_shape
+ seg_map = postprocess(outputs, (w, h))
+
+ # visualize
+ logger.info('6. Visualize...')
+ visualize(img, seg_map, args.save_path)
+
+ logger.info('Done...')
+
+
+PALETTE = [[120, 120, 120], [180, 120, 120], [6, 230, 230], [80, 50, 50],
+ [4, 200, 3], [120, 120, 80], [140, 140, 140], [204, 5, 255],
+ [230, 230, 230], [4, 250, 7], [224, 5, 255], [235, 255, 7],
+ [150, 5, 61], [120, 120, 70], [8, 255, 51], [255, 6, 82],
+ [143, 255, 140], [204, 255, 4], [255, 51, 7], [204, 70, 3],
+ [0, 102, 200], [61, 230, 250], [255, 6, 51], [11, 102, 255],
+ [255, 7, 71], [255, 9, 224], [9, 7, 230], [220, 220, 220],
+ [255, 9, 92], [112, 9, 255], [8, 255, 214], [7, 255, 224],
+ [255, 184, 6], [10, 255, 71], [255, 41, 10], [7, 255, 255],
+ [224, 255, 8], [102, 8, 255], [255, 61, 6], [255, 194, 7],
+ [255, 122, 8], [0, 255, 20], [255, 8, 41], [255, 5, 153],
+ [6, 51, 255], [235, 12, 255], [160, 150, 20], [0, 163, 255],
+ [140, 140, 140], [250, 10, 15], [20, 255, 0], [31, 255, 0],
+ [255, 31, 0], [255, 224, 0], [153, 255, 0], [0, 0, 255],
+ [255, 71, 0], [0, 235, 255], [0, 173, 255], [31, 0, 255],
+ [11, 200, 200], [255, 82, 0], [0, 255, 245], [0, 61, 255],
+ [0, 255, 112], [0, 255, 133], [255, 0, 0], [255, 163, 0],
+ [255, 102, 0], [194, 255, 0], [0, 143, 255], [51, 255, 0],
+ [0, 82, 255], [0, 255, 41], [0, 255, 173], [10, 0, 255],
+ [173, 255, 0], [0, 255, 153], [255, 92, 0], [255, 0, 255],
+ [255, 0, 245], [255, 0, 102], [255, 173, 0], [255, 0, 20],
+ [255, 184, 184], [0, 31, 255], [0, 255, 61], [0, 71, 255],
+ [255, 0, 204], [0, 255, 194], [0, 255, 82], [0, 10, 255],
+ [0, 112, 255], [51, 0, 255], [0, 194, 255], [0, 122, 255],
+ [0, 255, 163], [255, 153, 0], [0, 255, 10], [255, 112, 0],
+ [143, 255, 0], [82, 0, 255], [163, 255, 0], [255, 235, 0],
+ [8, 184, 170], [133, 0, 255], [0, 255, 92], [184, 0, 255],
+ [255, 0, 31], [0, 184, 255], [0, 214, 255], [255, 0, 112],
+ [92, 255, 0], [0, 224, 255], [112, 224, 255], [70, 184, 160],
+ [163, 0, 255], [153, 0, 255], [71, 255, 0], [255, 0, 163],
+ [255, 204, 0], [255, 0, 143], [0, 255, 235], [133, 255, 0],
+ [255, 0, 235], [245, 0, 255], [255, 0, 122], [255, 245, 0],
+ [10, 190, 212], [214, 255, 0], [0, 204, 255], [20, 0, 255],
+ [255, 255, 0], [0, 153, 255], [0, 41, 255], [0, 255, 204],
+ [41, 0, 255], [41, 255, 0], [173, 0, 255], [0, 245, 255],
+ [71, 0, 255], [122, 0, 255], [0, 255, 184], [0, 92, 255],
+ [184, 255, 0], [0, 133, 255], [255, 214, 0], [25, 194, 194],
+ [102, 255, 0], [92, 0, 255]]
+
+if __name__ == '__main__':
+ main()
diff --git a/projects/sam_inference_demo/README.md b/projects/sam_inference_demo/README.md
new file mode 100644
index 00000000000..f8077b8729f
--- /dev/null
+++ b/projects/sam_inference_demo/README.md
@@ -0,0 +1,40 @@
+# Introducing the Segment Anything Model (SAM) Inference Demo!
+
+Welcome to the Segment Anything (SA) Inference Demo, a user-friendly implementation based on the original Segment Anything project. Our demo allows you to experience the power and versatility of the Segment Anything Model (SAM) through an easy-to-use API.
+
+With this inference demo, you can explore the capabilities of the Segment Anything Model and witness its effectiveness in various tasks and image distributions. For more information on the original project, dataset, and model, please visit the official website at https://segment-anything.com.
+
+### Prerequisites
+
+- Python 3.10
+- PyTorch 1.13
+- MMEngine >= v0.7.2
+- MMCV >= v2.0.0
+
+### Installation
+
+We assume that you have already installed PyTorch. If not, please follow the instructions on the [PyTorch website](https://pytorch.org/).
+
+**1. Install MMEngine & MMCV**
+
+```shell
+pip install openmim
+mim install mmengine
+mim install 'mmcv>=2.0.0'
+```
+
+**2. Install MMPretrain**
+
+```shell
+pip install git+https://github.com/open-mmlab/mmpretrain.git@dev
+```
+
+**3. Install MMSegmentation**
+
+```shell
+pip install mmsegmentation
+```
+
+### Usage
+
+Open the `sam_image_demo.ipynb` notebook and follow the instructions to run the demo.
diff --git a/projects/sam_inference_demo/sam/__init__.py b/projects/sam_inference_demo/sam/__init__.py
new file mode 100644
index 00000000000..82b6b78469c
--- /dev/null
+++ b/projects/sam_inference_demo/sam/__init__.py
@@ -0,0 +1,2 @@
+from .modeling import * # noqa
+from .utils import * # noqa
diff --git a/projects/sam_inference_demo/sam/modeling/__init__.py b/projects/sam_inference_demo/sam/modeling/__init__.py
new file mode 100644
index 00000000000..9892a6b085d
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/__init__.py
@@ -0,0 +1,12 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+from .mask_decoder import MaskDecoder
+from .prompt_encoder import PromptEncoder
+from .sam import SAM
+from .transformer import TwoWayTransformer
+
+__all__ = ['SAM', 'MaskDecoder', 'PromptEncoder', 'TwoWayTransformer']
diff --git a/projects/sam_inference_demo/sam/modeling/common.py b/projects/sam_inference_demo/sam/modeling/common.py
new file mode 100644
index 00000000000..d2892761122
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/common.py
@@ -0,0 +1,45 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+from typing import Type
+
+import torch
+import torch.nn as nn
+
+
+class MLPBlock(nn.Module):
+
+ def __init__(
+ self,
+ embedding_dim: int,
+ mlp_dim: int,
+ act: Type[nn.Module] = nn.GELU,
+ ) -> None:
+ super().__init__()
+ self.lin1 = nn.Linear(embedding_dim, mlp_dim)
+ self.lin2 = nn.Linear(mlp_dim, embedding_dim)
+ self.act = act()
+
+ def forward(self, x: torch.Tensor) -> torch.Tensor:
+ return self.lin2(self.act(self.lin1(x)))
+
+
+# From https://github.com/facebookresearch/detectron2/blob/main/detectron2/layers/batch_norm.py # noqa
+# Itself from https://github.com/facebookresearch/ConvNeXt/blob/d1fa8f6fef0a165b27399986cc2bdacc92777e40/models/convnext.py#L119 # noqa
+class LayerNorm2d(nn.Module):
+
+ def __init__(self, num_channels: int, eps: float = 1e-6) -> None:
+ super().__init__()
+ self.weight = nn.Parameter(torch.ones(num_channels))
+ self.bias = nn.Parameter(torch.zeros(num_channels))
+ self.eps = eps
+
+ def forward(self, x: torch.Tensor) -> torch.Tensor:
+ u = x.mean(1, keepdim=True)
+ s = (x - u).pow(2).mean(1, keepdim=True)
+ x = (x - u) / torch.sqrt(s + self.eps)
+ x = self.weight[:, None, None] * x + self.bias[:, None, None]
+ return x
diff --git a/projects/sam_inference_demo/sam/modeling/mask_decoder.py b/projects/sam_inference_demo/sam/modeling/mask_decoder.py
new file mode 100644
index 00000000000..9ad616b5899
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/mask_decoder.py
@@ -0,0 +1,196 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# Borrowed from https://github.com/facebookresearch/segment-anything
+
+from typing import List, Tuple
+
+import torch
+from torch import Tensor, nn
+from torch.nn import functional as F
+
+from mmseg.registry import MODELS
+from .common import LayerNorm2d
+
+
+@MODELS.register_module()
+class MaskDecoder(nn.Module):
+
+ def __init__(
+ self,
+ *,
+ transformer_dim: int,
+ transformer: dict,
+ num_multimask_outputs: int = 3,
+ act_cfg: dict = dict(type='GELU'),
+ iou_head_depth: int = 3,
+ iou_head_hidden_dim: int = 256,
+ ) -> None:
+ """Predicts masks given an image and prompt embeddings, using a
+ tranformer architecture.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ transformer_dim (int): the channel dimension of the transformer
+ transformer (nn.Module): the transformer used to predict masks
+ num_multimask_outputs (int): the number of masks to predict
+ when disambiguating masks
+ activation (nn.Module): the type of activation to use when
+ upscaling masks
+ iou_head_depth (int): the depth of the MLP used to predict
+ mask quality
+ iou_head_hidden_dim (int): the hidden dimension of the MLP
+ used to predict mask quality
+ """
+ super().__init__()
+ self.transformer_dim = transformer_dim
+ self.transformer = MODELS.build(transformer)
+
+ self.num_multimask_outputs = num_multimask_outputs
+
+ self.iou_token = nn.Embedding(1, transformer_dim)
+ self.num_mask_tokens = num_multimask_outputs + 1
+ self.mask_tokens = nn.Embedding(self.num_mask_tokens, transformer_dim)
+
+ activation = MODELS.build(act_cfg)
+ self.output_upscaling = nn.Sequential(
+ nn.ConvTranspose2d(
+ transformer_dim, transformer_dim // 4, kernel_size=2,
+ stride=2),
+ LayerNorm2d(transformer_dim // 4),
+ activation,
+ nn.ConvTranspose2d(
+ transformer_dim // 4,
+ transformer_dim // 8,
+ kernel_size=2,
+ stride=2),
+ activation,
+ )
+ self.output_hypernetworks_mlps = nn.ModuleList([
+ MLP(transformer_dim, transformer_dim, transformer_dim // 8, 3)
+ for i in range(self.num_mask_tokens)
+ ])
+
+ self.iou_prediction_head = MLP(transformer_dim, iou_head_hidden_dim,
+ self.num_mask_tokens, iou_head_depth)
+
+ def forward(
+ self,
+ image_embeddings: Tensor,
+ image_pe: Tensor,
+ sparse_prompt_embeddings: Tensor,
+ dense_prompt_embeddings: Tensor,
+ multimask_output: bool,
+ ) -> Tuple[Tensor, Tensor]:
+ """Predict masks given image and prompt embeddings.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ image_embeddings (Tensor): the embeddings from the image encoder
+ image_pe (Tensor): positional encoding with the shape of
+ image_embeddings
+ sparse_prompt_embeddings (Tensor): the embeddings of
+ the points and boxes
+ dense_prompt_embeddings (Tensor): the embeddings of the mask inputs
+ multimask_output (bool): Whether to return multiple masks or a single
+ mask.
+
+ Returns:
+ Tensor: batched predicted masks
+ Tensor: batched predictions of mask quality
+ """
+ masks, iou_pred = self.predict_masks(
+ image_embeddings=image_embeddings,
+ image_pe=image_pe,
+ sparse_prompt_embeddings=sparse_prompt_embeddings,
+ dense_prompt_embeddings=dense_prompt_embeddings,
+ )
+
+ # Select the correct mask or masks for output
+ if multimask_output:
+ mask_slice = slice(1, None)
+ else:
+ mask_slice = slice(0, 1)
+ masks = masks[:, mask_slice, :, :]
+ iou_pred = iou_pred[:, mask_slice]
+
+ # Prepare output
+ return masks, iou_pred
+
+ def predict_masks(
+ self,
+ image_embeddings: Tensor,
+ image_pe: Tensor,
+ sparse_prompt_embeddings: Tensor,
+ dense_prompt_embeddings: Tensor,
+ ) -> Tuple[Tensor, Tensor]:
+ """Predicts masks.
+
+ See 'forward' for more details.
+ """
+ # Concatenate output tokens
+ output_tokens = torch.cat(
+ [self.iou_token.weight, self.mask_tokens.weight], dim=0)
+ output_tokens = output_tokens.unsqueeze(0).expand(
+ sparse_prompt_embeddings.size(0), -1, -1)
+ tokens = torch.cat((output_tokens, sparse_prompt_embeddings), dim=1)
+
+ # Expand per-image data in batch direction to be per-mask
+ src = torch.repeat_interleave(image_embeddings, tokens.shape[0], dim=0)
+ src = src + dense_prompt_embeddings
+ pos_src = torch.repeat_interleave(image_pe, tokens.shape[0], dim=0)
+ b, c, h, w = src.shape
+
+ # Run the transformer
+ hs, src = self.transformer(src, pos_src, tokens)
+ iou_token_out = hs[:, 0, :]
+ mask_tokens_out = hs[:, 1:(1 + self.num_mask_tokens), :]
+
+ # Upscale mask embeddings and predict masks using the mask tokens
+ src = src.transpose(1, 2).view(b, c, h, w)
+ upscaled_embedding = self.output_upscaling(src)
+ hyper_in_list: List[Tensor] = []
+ for i in range(self.num_mask_tokens):
+ hyper_in_list.append(self.output_hypernetworks_mlps[i](
+ mask_tokens_out[:, i, :]))
+ hyper_in = torch.stack(hyper_in_list, dim=1)
+ b, c, h, w = upscaled_embedding.shape
+ masks = (hyper_in @ upscaled_embedding.view(b, c, h * w)).view(
+ b, -1, h, w)
+
+ # Generate mask quality predictions
+ iou_pred = self.iou_prediction_head(iou_token_out)
+
+ return masks, iou_pred
+
+
+# Lightly adapted from
+# https://github.com/facebookresearch/MaskFormer/blob/main/mask_former/modeling/transformer/transformer_predictor.py # noqa
+class MLP(nn.Module):
+
+ def __init__(
+ self,
+ input_dim: int,
+ hidden_dim: int,
+ output_dim: int,
+ num_layers: int,
+ sigmoid_output: bool = False,
+ ) -> None:
+ super().__init__()
+ self.num_layers = num_layers
+ h = [hidden_dim] * (num_layers - 1)
+ self.layers = nn.ModuleList(
+ nn.Linear(n, k) for n, k in zip([input_dim] + h, h + [output_dim]))
+ self.sigmoid_output = sigmoid_output
+
+ def forward(self, x):
+ for i, layer in enumerate(self.layers):
+ x = F.relu(layer(x)) if i < self.num_layers - 1 else layer(x)
+ if self.sigmoid_output:
+ x = F.sigmoid(x)
+ return x
diff --git a/projects/sam_inference_demo/sam/modeling/prompt_encoder.py b/projects/sam_inference_demo/sam/modeling/prompt_encoder.py
new file mode 100644
index 00000000000..6b7c0833871
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/prompt_encoder.py
@@ -0,0 +1,227 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# Borrowed from https://github.com/facebookresearch/segment-anything
+
+from typing import Any, Optional, Tuple, Type
+
+import numpy as np
+import torch
+from torch import nn
+
+from mmseg.registry import MODELS
+from .common import LayerNorm2d
+
+
+@MODELS.register_module()
+class PromptEncoder(nn.Module):
+
+ def __init__(
+ self,
+ embed_dim: int,
+ image_embedding_size: Tuple[int, int],
+ input_image_size: Tuple[int, int],
+ mask_in_chans: int,
+ activation: Type[nn.Module] = nn.GELU,
+ ) -> None:
+ """Encodes prompts for input to SAM's mask decoder.
+
+ Arguments:
+ embed_dim (int): The prompts' embedding dimension
+ image_embedding_size (tuple(int, int)): The spatial size of the
+ image embedding, as (H, W).
+ input_image_size (int): The padded size of the image as input
+ to the image encoder, as (H, W).
+ mask_in_chans (int): The number of hidden channels used for
+ encoding input masks.
+ activation (nn.Module): The activation to use when encoding
+ input masks.
+ """
+ super().__init__()
+ self.embed_dim = embed_dim
+ self.input_image_size = input_image_size
+ self.image_embedding_size = image_embedding_size
+ self.pe_layer = PositionEmbeddingRandom(embed_dim // 2)
+
+ self.num_point_embeddings: int = 4 # pos/neg point + 2 box corners
+ point_embeddings = [
+ nn.Embedding(1, embed_dim)
+ for i in range(self.num_point_embeddings)
+ ]
+ self.point_embeddings = nn.ModuleList(point_embeddings)
+ self.not_a_point_embed = nn.Embedding(1, embed_dim)
+
+ self.mask_input_size = (4 * image_embedding_size[0],
+ 4 * image_embedding_size[1])
+ self.mask_downscaling = nn.Sequential(
+ nn.Conv2d(1, mask_in_chans // 4, kernel_size=2, stride=2),
+ LayerNorm2d(mask_in_chans // 4),
+ activation(),
+ nn.Conv2d(
+ mask_in_chans // 4, mask_in_chans, kernel_size=2, stride=2),
+ LayerNorm2d(mask_in_chans),
+ activation(),
+ nn.Conv2d(mask_in_chans, embed_dim, kernel_size=1),
+ )
+ self.no_mask_embed = nn.Embedding(1, embed_dim)
+
+ def get_dense_pe(self) -> torch.Tensor:
+ """Returns the positional encoding used to encode point prompts,
+ applied to a dense set of points the shape of the image encoding.
+
+ Returns:
+ torch.Tensor: Positional encoding with shape
+ 1x(embed_dim)x(embedding_h)x(embedding_w)
+ """
+ return self.pe_layer(self.image_embedding_size).unsqueeze(0)
+
+ def _embed_points(
+ self,
+ points: torch.Tensor,
+ labels: torch.Tensor,
+ pad: bool,
+ ) -> torch.Tensor:
+ """Embeds point prompts."""
+ points = points + 0.5 # Shift to center of pixel
+ if pad:
+ padding_point = torch.zeros((points.shape[0], 1, 2),
+ device=points.device)
+ padding_label = -torch.ones(
+ (labels.shape[0], 1), device=labels.device)
+ points = torch.cat([points, padding_point], dim=1)
+ labels = torch.cat([labels, padding_label], dim=1)
+ point_embedding = self.pe_layer.forward_with_coords(
+ points, self.input_image_size)
+ point_embedding[labels == -1] = 0.0
+ point_embedding[labels == -1] += self.not_a_point_embed.weight
+ point_embedding[labels == 0] += self.point_embeddings[0].weight
+ point_embedding[labels == 1] += self.point_embeddings[1].weight
+ return point_embedding
+
+ def _embed_boxes(self, boxes: torch.Tensor) -> torch.Tensor:
+ """Embeds box prompts."""
+ boxes = boxes + 0.5 # Shift to center of pixel
+ coords = boxes.reshape(-1, 2, 2)
+ corner_embedding = self.pe_layer.forward_with_coords(
+ coords, self.input_image_size)
+ corner_embedding[:, 0, :] += self.point_embeddings[2].weight
+ corner_embedding[:, 1, :] += self.point_embeddings[3].weight
+ return corner_embedding
+
+ def _embed_masks(self, masks: torch.Tensor) -> torch.Tensor:
+ """Embeds mask inputs."""
+ mask_embedding = self.mask_downscaling(masks)
+ return mask_embedding
+
+ def _get_batch_size(
+ self,
+ points: Optional[Tuple[torch.Tensor, torch.Tensor]],
+ boxes: Optional[torch.Tensor],
+ masks: Optional[torch.Tensor],
+ ) -> int:
+ """Gets the batch size of the output given the batch size of the input
+ prompts."""
+ if points is not None:
+ return points[0].shape[0]
+ elif boxes is not None:
+ return boxes.shape[0]
+ elif masks is not None:
+ return masks.shape[0]
+ else:
+ return 1
+
+ def _get_device(self) -> torch.device:
+ return self.point_embeddings[0].weight.device
+
+ def forward(
+ self,
+ points: Optional[Tuple[torch.Tensor, torch.Tensor]],
+ boxes: Optional[torch.Tensor],
+ masks: Optional[torch.Tensor],
+ ) -> Tuple[torch.Tensor, torch.Tensor]:
+ """Embeds different types of prompts, returning both sparse and dense
+ embeddings.
+
+ Arguments:
+ points (tuple(torch.Tensor, torch.Tensor) or none): point coordinates
+ and labels to embed.
+ boxes (torch.Tensor or none): boxes to embed
+ masks (torch.Tensor or none): masks to embed
+
+ Returns:
+ torch.Tensor: sparse embeddings for the points and boxes, with shape
+ BxNx(embed_dim), where N is determined by the number of input points
+ and boxes.
+ torch.Tensor: dense embeddings for the masks, in the shape
+ Bx(embed_dim)x(embed_H)x(embed_W)
+ """ # noqa
+ bs = self._get_batch_size(points, boxes, masks)
+ sparse_embeddings = torch.empty((bs, 0, self.embed_dim),
+ device=self._get_device())
+ if points is not None:
+ coords, labels = points
+ point_embeddings = self._embed_points(
+ coords, labels, pad=(boxes is None))
+ sparse_embeddings = torch.cat(
+ [sparse_embeddings, point_embeddings], dim=1)
+ if boxes is not None:
+ box_embeddings = self._embed_boxes(boxes)
+ sparse_embeddings = torch.cat([sparse_embeddings, box_embeddings],
+ dim=1)
+
+ if masks is not None:
+ dense_embeddings = self._embed_masks(masks)
+ else:
+ dense_embeddings = self.no_mask_embed.weight.reshape(
+ 1, -1, 1, 1).expand(bs, -1, self.image_embedding_size[0],
+ self.image_embedding_size[1])
+
+ return sparse_embeddings, dense_embeddings
+
+
+class PositionEmbeddingRandom(nn.Module):
+ """Positional encoding using random spatial frequencies."""
+
+ def __init__(self,
+ num_pos_feats: int = 64,
+ scale: Optional[float] = None) -> None:
+ super().__init__()
+ if scale is None or scale <= 0.0:
+ scale = 1.0
+ self.register_buffer(
+ 'positional_encoding_gaussian_matrix',
+ scale * torch.randn((2, num_pos_feats)),
+ )
+
+ def _pe_encoding(self, coords: torch.Tensor) -> torch.Tensor:
+ """Positionally encode points that are normalized to [0,1]."""
+ # assuming coords are in [0, 1]^2 square and have d_1 x ... x d_n x 2 shape # noqa
+ coords = 2 * coords - 1
+ coords = coords @ self.positional_encoding_gaussian_matrix
+ coords = 2 * np.pi * coords
+ # outputs d_1 x ... x d_n x C shape
+ return torch.cat([torch.sin(coords), torch.cos(coords)], dim=-1)
+
+ def forward(self, size: Tuple[int, int]) -> torch.Tensor:
+ """Generate positional encoding for a grid of the specified size."""
+ h, w = size
+ device: Any = self.positional_encoding_gaussian_matrix.device
+ grid = torch.ones((h, w), device=device, dtype=torch.float32)
+ y_embed = grid.cumsum(dim=0) - 0.5
+ x_embed = grid.cumsum(dim=1) - 0.5
+ y_embed = y_embed / h
+ x_embed = x_embed / w
+
+ pe = self._pe_encoding(torch.stack([x_embed, y_embed], dim=-1))
+ return pe.permute(2, 0, 1) # C x H x W
+
+ def forward_with_coords(self, coords_input: torch.Tensor,
+ image_size: Tuple[int, int]) -> torch.Tensor:
+ """Positionally encode points that are not normalized to [0,1]."""
+ coords = coords_input.clone()
+ coords[:, :, 0] = coords[:, :, 0] / image_size[1]
+ coords[:, :, 1] = coords[:, :, 1] / image_size[0]
+ return self._pe_encoding(coords.to(torch.float)) # B x N x C
diff --git a/projects/sam_inference_demo/sam/modeling/sam.py b/projects/sam_inference_demo/sam/modeling/sam.py
new file mode 100644
index 00000000000..c61c1eca4e7
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/sam.py
@@ -0,0 +1,188 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# Borrowed from https://github.com/facebookresearch/segment-anything
+
+from typing import Any, Dict, List, Tuple
+
+import torch
+from torch import nn
+from torch.nn import functional as F
+
+from mmseg.registry import MODELS
+from .mask_decoder import MaskDecoder
+from .prompt_encoder import PromptEncoder
+
+
+@MODELS.register_module()
+class SAM(nn.Module):
+ mask_threshold: float = 0.0
+ image_format: str = 'RGB'
+
+ def __init__(
+ self,
+ image_encoder_cfg: dict,
+ prompt_encoder_cfg: dict,
+ mask_decoder_cfg: dict,
+ pixel_mean: List[float] = [123.675, 116.28, 103.53],
+ pixel_std: List[float] = [58.395, 57.12, 57.375],
+ ) -> None:
+ """SAM predicts object masks from an image and input prompts. Borrowed
+ from https://github.com/facebookresearch/segment-anything.
+
+ Arguments:
+ image_encoder (ViTSAM): The backbone used to encode the
+ image into image embeddings that allow for efficient mask
+ prediction.
+ prompt_encoder (PromptEncoder): Encodes various types of input
+ prompts.
+ mask_decoder (MaskDecoder): Predicts masks from the image embeddings
+ and encoded prompts.
+ pixel_mean (list(float)): Mean values for normalizing pixels in the
+ input image.
+ pixel_std (list(float)): Std values for normalizing pixels in the
+ input image.
+ """
+ super().__init__()
+ self.image_encoder = MODELS.build(image_encoder_cfg)
+ self.prompt_encoder: PromptEncoder = MODELS.build(prompt_encoder_cfg)
+ self.mask_decoder: MaskDecoder = MODELS.build(mask_decoder_cfg)
+ self.register_buffer('pixel_mean',
+ torch.Tensor(pixel_mean).view(-1, 1, 1), False)
+ self.register_buffer('pixel_std',
+ torch.Tensor(pixel_std).view(-1, 1, 1), False)
+
+ @property
+ def device(self) -> Any:
+ return self.pixel_mean.device
+
+ @torch.no_grad()
+ def forward(
+ self,
+ batched_input: List[Dict[str, Any]],
+ multimask_output: bool,
+ ) -> List[Dict[str, torch.Tensor]]:
+ """Predicts masks end-to-end from provided images and prompts. If
+ prompts are not known in advance, using SamPredictor is recommended
+ over calling the model directly.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ batched_input (list(dict)): A list over input images, each a
+ dictionary with the following keys. A prompt key can be
+ excluded if it is not present.
+ 'image': The image as a torch tensor in 3xHxW format,
+ already transformed for input to the model.
+ 'original_size': (tuple(int, int)) The original size of
+ the image before transformation, as (H, W).
+ 'point_coords': (torch.Tensor) Batched point prompts for
+ this image, with shape BxNx2. Already transformed to the
+ input frame of the model.
+ 'point_labels': (torch.Tensor) Batched labels for point prompts,
+ with shape BxN.
+ 'boxes': (torch.Tensor) Batched box inputs, with shape Bx4.
+ Already transformed to the input frame of the model.
+ 'mask_inputs': (torch.Tensor) Batched mask inputs to the model,
+ in the form Bx1xHxW.
+ multimask_output (bool): Whether the model should predict multiple
+ disambiguating masks, or return a single mask.
+
+ Returns:
+ (list(dict)): A list over input images, where each element is
+ as dictionary with the following keys.
+ 'masks': (torch.Tensor) Batched binary mask predictions,
+ with shape BxCxHxW, where B is the number of input prompts,
+ C is determiend by multimask_output, and (H, W) is the
+ original size of the image.
+ 'iou_predictions': (torch.Tensor) The model's predictions
+ of mask quality, in shape BxC.
+ 'low_res_logits': (torch.Tensor) Low resolution logits with
+ shape BxCxHxW, where H=W=256. Can be passed as mask input
+ to subsequent iterations of prediction.
+ """
+ input_images = torch.stack(
+ [self.preprocess(x['image']) for x in batched_input], dim=0)
+ image_embeddings = self.image_encoder(input_images)
+
+ outputs = []
+ for image_record, curr_embedding in zip(batched_input,
+ image_embeddings):
+ if 'point_coords' in image_record:
+ points = (image_record['point_coords'],
+ image_record['point_labels'])
+ else:
+ points = None
+ sparse_embeddings, dense_embeddings = self.prompt_encoder(
+ points=points,
+ boxes=image_record.get('boxes', None),
+ masks=image_record.get('mask_inputs', None),
+ )
+ low_res_masks, iou_predictions = self.mask_decoder(
+ image_embeddings=curr_embedding.unsqueeze(0),
+ image_pe=self.prompt_encoder.get_dense_pe(),
+ sparse_prompt_embeddings=sparse_embeddings,
+ dense_prompt_embeddings=dense_embeddings,
+ multimask_output=multimask_output,
+ )
+ masks = self.postprocess_masks(
+ low_res_masks,
+ input_size=image_record['image'].shape[-2:],
+ original_size=image_record['original_size'],
+ )
+ masks = masks > self.mask_threshold
+ outputs.append({
+ 'masks': masks,
+ 'iou_predictions': iou_predictions,
+ 'low_res_logits': low_res_masks,
+ })
+ return outputs
+
+ def postprocess_masks(
+ self,
+ masks: torch.Tensor,
+ input_size: Tuple[int, ...],
+ original_size: Tuple[int, ...],
+ ) -> torch.Tensor:
+ """Remove padding and upscale masks to the original image size.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ masks (torch.Tensor): Batched masks from the mask_decoder,
+ in BxCxHxW format.
+ input_size (tuple(int, int)): The size of the image input to the
+ model, in (H, W) format. Used to remove padding.
+ original_size (tuple(int, int)): The original size of the image
+ before resizing for input to the model, in (H, W) format.
+
+ Returns:
+ (torch.Tensor): Batched masks in BxCxHxW format, where (H, W)
+ is given by original_size.
+ """
+ masks = F.interpolate(
+ masks,
+ self.image_encoder.img_size,
+ mode='bilinear',
+ align_corners=False,
+ )
+ masks = masks[..., :input_size[0], :input_size[1]]
+ masks = F.interpolate(
+ masks, original_size, mode='bilinear', align_corners=False)
+ return masks
+
+ def preprocess(self, x: torch.Tensor) -> torch.Tensor:
+ """Normalize pixel values and pad to a square input."""
+ # Normalize colors
+ x = (x - self.pixel_mean) / self.pixel_std
+
+ # Pad
+ h, w = x.shape[-2:]
+ img_size = max(self.image_encoder.img_size)
+ padh = img_size - h
+ padw = img_size - w
+ x = F.pad(x, (0, padw, 0, padh))
+ return x
diff --git a/projects/sam_inference_demo/sam/modeling/transformer.py b/projects/sam_inference_demo/sam/modeling/transformer.py
new file mode 100644
index 00000000000..c56f602487d
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/transformer.py
@@ -0,0 +1,241 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+import math
+from typing import Tuple, Type
+
+import torch
+from torch import Tensor, nn
+
+from mmseg.registry import MODELS
+from .common import MLPBlock
+
+
+@MODELS.register_module()
+class TwoWayTransformer(nn.Module):
+
+ def __init__(
+ self,
+ depth: int,
+ embedding_dim: int,
+ num_heads: int,
+ mlp_dim: int,
+ activation: Type[nn.Module] = nn.ReLU,
+ attention_downsample_rate: int = 2,
+ ) -> None:
+ """A transformer decoder that attends to an input image using queries
+ whose positional embedding is supplied.
+
+ Args:
+ depth (int): number of layers in the transformer
+ embedding_dim (int): the channel dimension for the input embeddings
+ num_heads (int): the number of heads for multihead attention. Must
+ divide embedding_dim
+ mlp_dim (int): the channel dimension internal to the MLP block
+ activation (nn.Module): the activation to use in the MLP block
+ """
+ super().__init__()
+ self.depth = depth
+ self.embedding_dim = embedding_dim
+ self.num_heads = num_heads
+ self.mlp_dim = mlp_dim
+ self.layers = nn.ModuleList()
+
+ for i in range(depth):
+ self.layers.append(
+ TwoWayAttentionBlock(
+ embedding_dim=embedding_dim,
+ num_heads=num_heads,
+ mlp_dim=mlp_dim,
+ activation=activation,
+ attention_downsample_rate=attention_downsample_rate,
+ skip_first_layer_pe=(i == 0),
+ ))
+
+ self.final_attn_token_to_image = Attention(
+ embedding_dim,
+ num_heads,
+ downsample_rate=attention_downsample_rate)
+ self.norm_final_attn = nn.LayerNorm(embedding_dim)
+
+ def forward(
+ self,
+ image_embedding: Tensor,
+ image_pe: Tensor,
+ point_embedding: Tensor,
+ ) -> Tuple[Tensor, Tensor]:
+ """
+ Args:
+ image_embedding (torch.Tensor): image to attend to. Should be shape
+ B x embedding_dim x h x w for any h and w.
+ image_pe (torch.Tensor): the positional encoding to add to the image. Must
+ have the same shape as image_embedding.
+ point_embedding (torch.Tensor): the embedding to add to the query points.
+ Must have shape B x N_points x embedding_dim for any N_points.
+
+ Returns:
+ torch.Tensor: the processed point_embedding
+ torch.Tensor: the processed image_embedding
+ """ # noqa E501
+ # BxCxHxW -> BxHWxC == B x N_image_tokens x C
+ bs, c, h, w = image_embedding.shape
+ image_embedding = image_embedding.flatten(2).permute(0, 2, 1)
+ image_pe = image_pe.flatten(2).permute(0, 2, 1)
+
+ # Prepare queries
+ queries = point_embedding
+ keys = image_embedding
+
+ # Apply transformer blocks and final layernorm
+ for layer in self.layers:
+ queries, keys = layer(
+ queries=queries,
+ keys=keys,
+ query_pe=point_embedding,
+ key_pe=image_pe,
+ )
+
+ # Apply the final attenion layer from the points to the image
+ q = queries + point_embedding
+ k = keys + image_pe
+ attn_out = self.final_attn_token_to_image(q=q, k=k, v=keys)
+ queries = queries + attn_out
+ queries = self.norm_final_attn(queries)
+
+ return queries, keys
+
+
+class TwoWayAttentionBlock(nn.Module):
+
+ def __init__(
+ self,
+ embedding_dim: int,
+ num_heads: int,
+ mlp_dim: int = 2048,
+ activation: Type[nn.Module] = nn.ReLU,
+ attention_downsample_rate: int = 2,
+ skip_first_layer_pe: bool = False,
+ ) -> None:
+ """A transformer block with four layers: (1) self-attention of sparse
+ inputs, (2) cross attention of sparse inputs to dense inputs, (3) mlp
+ block on sparse inputs, and (4) cross attention of dense inputs to
+ sparse inputs.
+
+ Arguments:
+ embedding_dim (int): the channel dimension of the embeddings
+ num_heads (int): the number of heads in the attention layers
+ mlp_dim (int): the hidden dimension of the mlp block
+ activation (nn.Module): the activation of the mlp block
+ skip_first_layer_pe (bool): skip the PE on the first layer
+ """
+ super().__init__()
+ self.self_attn = Attention(embedding_dim, num_heads)
+ self.norm1 = nn.LayerNorm(embedding_dim)
+
+ self.cross_attn_token_to_image = Attention(
+ embedding_dim,
+ num_heads,
+ downsample_rate=attention_downsample_rate)
+ self.norm2 = nn.LayerNorm(embedding_dim)
+
+ self.mlp = MLPBlock(embedding_dim, mlp_dim, activation)
+ self.norm3 = nn.LayerNorm(embedding_dim)
+
+ self.norm4 = nn.LayerNorm(embedding_dim)
+ self.cross_attn_image_to_token = Attention(
+ embedding_dim,
+ num_heads,
+ downsample_rate=attention_downsample_rate)
+
+ self.skip_first_layer_pe = skip_first_layer_pe
+
+ def forward(self, queries: Tensor, keys: Tensor, query_pe: Tensor,
+ key_pe: Tensor) -> Tuple[Tensor, Tensor]:
+ # Self attention block
+ if self.skip_first_layer_pe:
+ queries = self.self_attn(q=queries, k=queries, v=queries)
+ else:
+ q = queries + query_pe
+ attn_out = self.self_attn(q=q, k=q, v=queries)
+ queries = queries + attn_out
+ queries = self.norm1(queries)
+
+ # Cross attention block, tokens attending to image embedding
+ q = queries + query_pe
+ k = keys + key_pe
+ attn_out = self.cross_attn_token_to_image(q=q, k=k, v=keys)
+ queries = queries + attn_out
+ queries = self.norm2(queries)
+
+ # MLP block
+ mlp_out = self.mlp(queries)
+ queries = queries + mlp_out
+ queries = self.norm3(queries)
+
+ # Cross attention block, image embedding attending to tokens
+ q = queries + query_pe
+ k = keys + key_pe
+ attn_out = self.cross_attn_image_to_token(q=k, k=q, v=queries)
+ keys = keys + attn_out
+ keys = self.norm4(keys)
+
+ return queries, keys
+
+
+class Attention(nn.Module):
+ """An attention layer that allows for downscaling the size of the embedding
+ after projection to queries, keys, and values."""
+
+ def __init__(
+ self,
+ embedding_dim: int,
+ num_heads: int,
+ downsample_rate: int = 1,
+ ) -> None:
+ super().__init__()
+ self.embedding_dim = embedding_dim
+ self.internal_dim = embedding_dim // downsample_rate
+ self.num_heads = num_heads
+ assert self.internal_dim % num_heads == 0, 'num_heads must divide embedding_dim.' # noqa E501
+
+ self.q_proj = nn.Linear(embedding_dim, self.internal_dim)
+ self.k_proj = nn.Linear(embedding_dim, self.internal_dim)
+ self.v_proj = nn.Linear(embedding_dim, self.internal_dim)
+ self.out_proj = nn.Linear(self.internal_dim, embedding_dim)
+
+ def _separate_heads(self, x: Tensor, num_heads: int) -> Tensor:
+ b, n, c = x.shape
+ x = x.reshape(b, n, num_heads, c // num_heads)
+ return x.transpose(1, 2) # B x N_heads x N_tokens x C_per_head
+
+ def _recombine_heads(self, x: Tensor) -> Tensor:
+ b, n_heads, n_tokens, c_per_head = x.shape
+ x = x.transpose(1, 2)
+ return x.reshape(b, n_tokens, n_heads * c_per_head) # B x N_tokens x C
+
+ def forward(self, q: Tensor, k: Tensor, v: Tensor) -> Tensor:
+ # Input projections
+ q = self.q_proj(q)
+ k = self.k_proj(k)
+ v = self.v_proj(v)
+
+ # Separate into heads
+ q = self._separate_heads(q, self.num_heads)
+ k = self._separate_heads(k, self.num_heads)
+ v = self._separate_heads(v, self.num_heads)
+
+ # Attention
+ _, _, _, c_per_head = q.shape
+ attn = q @ k.permute(0, 1, 3, 2) # B x N_heads x N_tokens x N_tokens
+ attn = attn / math.sqrt(c_per_head)
+ attn = torch.softmax(attn, dim=-1)
+
+ # Get output
+ out = attn @ v
+ out = self._recombine_heads(out)
+ out = self.out_proj(out)
+
+ return out
diff --git a/projects/sam_inference_demo/sam/sam_inferencer.py b/projects/sam_inference_demo/sam/sam_inferencer.py
new file mode 100644
index 00000000000..2da2e959c09
--- /dev/null
+++ b/projects/sam_inference_demo/sam/sam_inferencer.py
@@ -0,0 +1,688 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from typing import Any, Dict, List, Optional, Tuple
+
+import numpy as np
+import torch
+from mmengine.runner.checkpoint import load_checkpoint
+# yapf: disable
+from sam.utils import (MaskData, area_from_rle, batch_iterator,
+ batched_mask_to_box, box_xyxy_to_xywh,
+ build_all_layer_point_grids, calculate_stability_score,
+ coco_encode_rle, generate_crop_boxes,
+ is_box_near_crop_edge, mask_to_rle_pytorch,
+ remove_small_regions, rle_to_mask, uncrop_boxes_xyxy,
+ uncrop_masks, uncrop_points)
+from torchvision.ops.boxes import batched_nms, box_area
+
+from mmseg.registry import MODELS, TRANSFORMS
+
+# yapf: enable
+
+model_zoo = {
+ 'base':
+ 'https://download.openmmlab.com/mmsegmentation/v0.5/sam/sam_vit-base-p16_3rdparty_sa1b-1024x1024_20230413-78a25eed.pth', # noqa
+ 'large':
+ 'https://download.openmmlab.com/mmsegmentation/v0.5/sam/sam_vit-large-p16_3rdparty_sa1b-1024x1024_20230413-940520da.pth', # noqa
+ 'huge':
+ 'https://download.openmmlab.com/mmsegmentation/v0.5/sam/sam_vit-huge-p16_3rdparty_sa1b-1024x1024_20230413-faaf96f6.pth', # noqa
+}
+
+
+class SAMInferencer:
+
+ def __init__(self, arch: str = 'base') -> None:
+ assert arch in ['base', 'large', 'huge']
+ self.model = self.init_model(arch)
+ self.transform = TRANSFORMS.build(
+ dict(
+ type='ResizeLongestSide',
+ target_length=max(self.model.image_encoder.img_size)))
+
+ def set_image(
+ self,
+ image: np.ndarray,
+ image_format: str = 'RGB',
+ ) -> None:
+ """Calculates the image embeddings for the provided image, allowing
+ masks to be predicted with the 'predict' method.
+
+ Arguments:
+ image (np.ndarray): The image for calculating masks. Expects an
+ image in HWC uint8 format, with pixel values in [0, 255].
+ image_format (str): The color format of the image, in ['RGB', 'BGR'].
+ """
+ assert image_format in [
+ 'RGB',
+ 'BGR',
+ ], f"image_format must be in ['RGB', 'BGR'], is {image_format}."
+ if image_format != self.model.image_format:
+ image = image[..., ::-1]
+
+ # Transform the image to the form expected by the model
+ input_image = self.transform.apply_image(image)
+ input_image_torch = torch.as_tensor(input_image, device=self.device)
+ input_image_torch = input_image_torch.permute(
+ 2, 0, 1).contiguous()[None, :, :, :]
+
+ self.set_torch_image(input_image_torch, image.shape[:2])
+
+ @torch.no_grad()
+ def set_torch_image(
+ self,
+ transformed_image: torch.Tensor,
+ original_image_size: Tuple[int, ...],
+ ) -> None:
+ """Calculates the image embeddings for the provided image, allowing
+ masks to be predicted with the 'predict' method. Expects the input
+ image to be already transformed to the format expected by the model.
+
+ Arguments:
+ transformed_image (torch.Tensor): The input image, with shape
+ 1x3xHxW, which has been transformed with ResizeLongestSide.
+ original_image_size (tuple(int, int)): The size of the image
+ before transformation, in (H, W) format.
+ """
+ assert (len(transformed_image.shape) == 4
+ and transformed_image.shape[1] == 3
+ and max(*transformed_image.shape[2:]) == max(
+ self.model.image_encoder.img_size)
+ ), 'set_torch_image input must be BCHW with long side'
+ f' {self.model.image_encoder.img_size}.'
+ self.reset_image()
+
+ self.original_size = original_image_size
+ self.input_size = tuple(transformed_image.shape[-2:])
+ input_image = self.model.preprocess(transformed_image)
+ self.features = self.model.image_encoder(input_image)[0]
+ self.is_image_set = True
+
+ def predict(
+ self,
+ point_coords: Optional[np.ndarray] = None,
+ point_labels: Optional[np.ndarray] = None,
+ box: Optional[np.ndarray] = None,
+ mask_input: Optional[np.ndarray] = None,
+ multimask_output: bool = True,
+ return_logits: bool = False,
+ ) -> Tuple[torch.Tensor, torch.Tensor, torch.Tensor]:
+ """Predict masks for the given input prompts, using the currently set
+ image.
+
+ Arguments:
+ point_coords (np.ndarray or None): A Nx2 array of point prompts to the
+ model. Each point is in (X,Y) in pixels.
+ point_labels (np.ndarray or None): A length N array of labels for the
+ point prompts. 1 indicates a foreground point and 0 indicates a
+ background point.
+ box (np.ndarray or None): A length 4 array given a box prompt to the
+ model, in XYXY format.
+ mask_input (np.ndarray): A low resolution mask input to the model, typically
+ coming from a previous prediction iteration. Has form 1xHxW, where
+ for SAM, H=W=256.
+ multimask_output (bool): If true, the model will return three masks.
+ For ambiguous input prompts (such as a single click), this will often
+ produce better masks than a single prediction. If only a single
+ mask is needed, the model's predicted quality score can be used
+ to select the best mask. For non-ambiguous prompts, such as multiple
+ input prompts, multimask_output=False can give better results.
+ return_logits (bool): If true, returns un-thresholded masks logits
+ instead of a binary mask.
+
+ Returns:
+ (np.ndarray): The output masks in CxHxW format, where C is the
+ number of masks, and (H, W) is the original image size.
+ (np.ndarray): An array of length C containing the model's
+ predictions for the quality of each mask.
+ (np.ndarray): An array of shape CxHxW, where C is the number
+ of masks and H=W=256. These low resolution logits can be passed to
+ a subsequent iteration as mask input.
+ """ # noqa
+ if not self.is_image_set:
+ raise RuntimeError(
+ 'An image must be set with .set_image(...) before mask'
+ 'prediction.')
+
+ # Transform input prompts
+ coords_torch = None
+ labels_torch = None
+ box_torch = None
+ mask_input_torch = None
+
+ if point_coords is not None:
+ assert (
+ point_labels is not None
+ ), 'point_labels must be supplied if point_coords is supplied.'
+ point_coords = self.transform.apply_coords(point_coords,
+ self.original_size)
+ coords_torch = torch.as_tensor(
+ point_coords, dtype=torch.float, device=self.device)
+ labels_torch = torch.as_tensor(
+ point_labels, dtype=torch.int, device=self.device)
+ coords_torch, labels_torch = coords_torch[
+ None, :, :], labels_torch[None, :]
+ if box is not None:
+ box = self.transform.apply_boxes(box, self.original_size)
+ box_torch = torch.as_tensor(
+ box, dtype=torch.float, device=self.device)
+ box_torch = box_torch[None, :]
+ if mask_input is not None:
+ mask_input_torch = torch.as_tensor(
+ mask_input, dtype=torch.float, device=self.device)
+ mask_input_torch = mask_input_torch[None, :, :, :]
+
+ masks, iou_predictions, low_res_masks = self.predict_torch(
+ coords_torch,
+ labels_torch,
+ box_torch,
+ mask_input_torch,
+ multimask_output,
+ return_logits=return_logits,
+ )
+
+ masks = masks[0].detach().cpu().numpy()
+ iou_predictions = iou_predictions[0].detach().cpu().numpy()
+ low_res_masks = low_res_masks[0].detach().cpu().numpy()
+ return masks, iou_predictions, low_res_masks
+
+ @torch.no_grad()
+ def predict_torch(
+ self,
+ point_coords: Optional[torch.Tensor],
+ point_labels: Optional[torch.Tensor],
+ boxes: Optional[torch.Tensor] = None,
+ mask_input: Optional[torch.Tensor] = None,
+ multimask_output: bool = True,
+ return_logits: bool = False,
+ ) -> Tuple[torch.Tensor, torch.Tensor, torch.Tensor]:
+ """Predict masks for the given input prompts, using the currently set
+ image. Input prompts are batched torch tensors and are expected to
+ already be transformed to the input frame using ResizeLongestSide.
+
+ Arguments:
+ point_coords (torch.Tensor or None): A BxNx2 array of point prompts to the
+ model. Each point is in (X,Y) in pixels.
+ point_labels (torch.Tensor or None): A BxN array of labels for the
+ point prompts. 1 indicates a foreground point and 0 indicates a
+ background point.
+ box (np.ndarray or None): A Bx4 array given a box prompt to the
+ model, in XYXY format.
+ mask_input (np.ndarray): A low resolution mask input to the model, typically
+ coming from a previous prediction iteration. Has form Bx1xHxW, where
+ for SAM, H=W=256. Masks returned by a previous iteration of the
+ predict method do not need further transformation.
+ multimask_output (bool): If true, the model will return three masks.
+ For ambiguous input prompts (such as a single click), this will often
+ produce better masks than a single prediction. If only a single
+ mask is needed, the model's predicted quality score can be used
+ to select the best mask. For non-ambiguous prompts, such as multiple
+ input prompts, multimask_output=False can give better results.
+ return_logits (bool): If true, returns un-thresholded masks logits
+ instead of a binary mask.
+
+ Returns:
+ (torch.Tensor): The output masks in BxCxHxW format, where C is the
+ number of masks, and (H, W) is the original image size.
+ (torch.Tensor): An array of shape BxC containing the model's
+ predictions for the quality of each mask.
+ (torch.Tensor): An array of shape BxCxHxW, where C is the number
+ of masks and H=W=256. These low res logits can be passed to
+ a subsequent iteration as mask input.
+ """ # noqa
+ if not self.is_image_set:
+ raise RuntimeError(
+ 'An image must be set with .set_image(...) before mask '
+ 'prediction.')
+
+ if point_coords is not None:
+ points = (point_coords, point_labels)
+ else:
+ points = None
+
+ # Embed prompts
+ sparse_embeddings, dense_embeddings = self.model.prompt_encoder(
+ points=points,
+ boxes=boxes,
+ masks=mask_input,
+ )
+
+ # Predict masks
+ low_res_masks, iou_predictions = self.model.mask_decoder(
+ image_embeddings=self.features,
+ image_pe=self.model.prompt_encoder.get_dense_pe(),
+ sparse_prompt_embeddings=sparse_embeddings,
+ dense_prompt_embeddings=dense_embeddings,
+ multimask_output=multimask_output,
+ )
+
+ # Upscale the masks to the original image resolution
+ masks = self.model.postprocess_masks(low_res_masks, self.input_size,
+ self.original_size)
+
+ if not return_logits:
+ masks = masks > self.model.mask_threshold
+
+ return masks, iou_predictions, low_res_masks
+
+ def get_image_embedding(self) -> torch.Tensor:
+ """Returns the image embeddings for the currently set image, with shape
+ 1xCxHxW, where C is the embedding dimension and (H,W) are the embedding
+ spatial dimension of SAM (typically C=256, H=W=64)."""
+ if not self.is_image_set:
+ raise RuntimeError(
+ 'An image must be set with .set_image(...) to generate an '
+ 'embedding.')
+ assert self.features is not None, 'Features must exist if an image has'
+ ' been set.'
+ return self.features
+
+ @property
+ def device(self) -> torch.device:
+ return self.model.device
+
+ def reset_image(self) -> None:
+ """Resets the currently set image."""
+ self.is_image_set = False
+ self.features = None
+ self.orig_h = None
+ self.orig_w = None
+ self.input_h = None
+ self.input_w = None
+
+ def init_model(self, arch: str):
+ model = MODELS.build(
+ dict(
+ type='SAM',
+ image_encoder_cfg=dict(
+ type='mmpretrain.ViTSAM',
+ arch=arch,
+ img_size=1024,
+ patch_size=16,
+ out_channels=256,
+ use_abs_pos=True,
+ use_rel_pos=True,
+ window_size=14,
+ ),
+ prompt_encoder_cfg=dict(
+ type='PromptEncoder',
+ embed_dim=256,
+ image_embedding_size=(64, 64),
+ input_image_size=(1024, 1024),
+ mask_in_chans=16,
+ ),
+ mask_decoder_cfg=dict(
+ type='MaskDecoder',
+ num_multimask_outputs=3,
+ transformer=dict(
+ type='TwoWayTransformer',
+ depth=2,
+ embedding_dim=256,
+ mlp_dim=2048,
+ num_heads=8,
+ ),
+ transformer_dim=256,
+ iou_head_depth=3,
+ iou_head_hidden_dim=256,
+ )))
+ load_checkpoint(model, model_zoo.get(arch), strict=True)
+ if torch.cuda.is_available():
+ model = model.cuda()
+ return model
+
+
+class SamAutomaticMaskGenerator:
+
+ def __init__(
+ self,
+ arch: str = 'base',
+ points_per_side: Optional[int] = 32,
+ points_per_batch: int = 64,
+ pred_iou_thresh: float = 0.88,
+ stability_score_thresh: float = 0.95,
+ stability_score_offset: float = 1.0,
+ box_nms_thresh: float = 0.7,
+ crop_n_layers: int = 0,
+ crop_nms_thresh: float = 0.7,
+ crop_overlap_ratio: float = 512 / 1500,
+ crop_n_points_downscale_factor: int = 1,
+ point_grids: Optional[List[np.ndarray]] = None,
+ min_mask_region_area: int = 0,
+ output_mode: str = 'binary_mask',
+ ) -> None:
+ """Using a SAM model, generates masks for the entire image. Generates a
+ grid of point prompts over the image, then filters low quality and
+ duplicate masks. The default settings are chosen for SAM with a ViT-H
+ backbone.
+
+ Arguments:
+ arch (str): The SAM model to use for mask prediction.
+ points_per_side (int or None): The number of points to be sampled
+ along one side of the image. The total number of points is
+ points_per_side**2. If None, 'point_grids' must provide explicit
+ point sampling.
+ points_per_batch (int): Sets the number of points run simultaneously
+ by the model. Higher numbers may be faster but use more GPU memory.
+ pred_iou_thresh (float): A filtering threshold in [0,1], using the
+ model's predicted mask quality.
+ stability_score_thresh (float): A filtering threshold in [0,1], using
+ the stability of the mask under changes to the cutoff used to binarize
+ the model's mask predictions.
+ stability_score_offset (float): The amount to shift the cutoff when
+ calculated the stability score.
+ box_nms_thresh (float): The box IoU cutoff used by non-maximal
+ suppression to filter duplicate masks.
+ crops_n_layers (int): If >0, mask prediction will be run again on
+ crops of the image. Sets the number of layers to run, where each
+ layer has 2**i_layer number of image crops.
+ crops_nms_thresh (float): The box IoU cutoff used by non-maximal
+ suppression to filter duplicate masks between different crops.
+ crop_overlap_ratio (float): Sets the degree to which crops overlap.
+ In the first crop layer, crops will overlap by this fraction of
+ the image length. Later layers with more crops scale down this overlap.
+ crop_n_points_downscale_factor (int): The number of points-per-side
+ sampled in layer n is scaled down by crop_n_points_downscale_factor**n.
+ point_grids (list(np.ndarray) or None): A list over explicit grids
+ of points used for sampling, normalized to [0,1]. The nth grid in the
+ list is used in the nth crop layer. Exclusive with points_per_side.
+ min_mask_region_area (int): If >0, postprocessing will be applied
+ to remove disconnected regions and holes in masks with area smaller
+ than min_mask_region_area. Requires opencv.
+ output_mode (str): The form masks are returned in. Can be 'binary_mask',
+ 'uncompressed_rle', or 'coco_rle'. 'coco_rle' requires pycocotools.
+ For large resolutions, 'binary_mask' may consume large amounts of
+ memory.
+ """ # noqa
+
+ assert (points_per_side is None) != (
+ point_grids is None
+ ), 'Exactly one of points_per_side or point_grid must be provided.'
+ if points_per_side is not None:
+ self.point_grids = build_all_layer_point_grids(
+ points_per_side,
+ crop_n_layers,
+ crop_n_points_downscale_factor,
+ )
+ elif point_grids is not None:
+ self.point_grids = point_grids
+ else:
+ raise ValueError(
+ "Can't have both points_per_side and point_grid be None.")
+
+ assert output_mode in [
+ 'binary_mask',
+ 'uncompressed_rle',
+ 'coco_rle',
+ ], f'Unknown output_mode {output_mode}.'
+ if output_mode == 'coco_rle':
+ from pycocotools import \
+ mask as mask_utils # type: ignore # noqa: F401
+
+ if min_mask_region_area > 0:
+ import cv2 # type: ignore # noqa: F401
+
+ self.predictor = SAMInferencer(arch)
+ self.points_per_batch = points_per_batch
+ self.pred_iou_thresh = pred_iou_thresh
+ self.stability_score_thresh = stability_score_thresh
+ self.stability_score_offset = stability_score_offset
+ self.box_nms_thresh = box_nms_thresh
+ self.crop_n_layers = crop_n_layers
+ self.crop_nms_thresh = crop_nms_thresh
+ self.crop_overlap_ratio = crop_overlap_ratio
+ self.crop_n_points_downscale_factor = crop_n_points_downscale_factor
+ self.min_mask_region_area = min_mask_region_area
+ self.output_mode = output_mode
+
+ @torch.no_grad()
+ def generate(self, image: np.ndarray) -> List[Dict[str, Any]]:
+ """Generates masks for the given image.
+
+ Arguments:
+ image (np.ndarray): The image to generate masks for, in HWC uint8 format.
+
+ Returns:
+ list(dict(str, any)): A list over records for masks. Each record is
+ a dict containing the following keys:
+ segmentation (dict(str, any) or np.ndarray): The mask. If
+ output_mode='binary_mask', is an array of shape HW. Otherwise,
+ is a dictionary containing the RLE.
+ bbox (list(float)): The box around the mask, in XYWH format.
+ area (int): The area in pixels of the mask.
+ predicted_iou (float): The model's own prediction of the mask's
+ quality. This is filtered by the pred_iou_thresh parameter.
+ point_coords (list(list(float))): The point coordinates input
+ to the model to generate this mask.
+ stability_score (float): A measure of the mask's quality. This
+ is filtered on using the stability_score_thresh parameter.
+ crop_box (list(float)): The crop of the image used to generate
+ the mask, given in XYWH format.
+ """ # noqa
+
+ # Generate masks
+ mask_data = self._generate_masks(image)
+
+ # Filter small disconnected regions and holes in masks
+ if self.min_mask_region_area > 0:
+ mask_data = self.postprocess_small_regions(
+ mask_data,
+ self.min_mask_region_area,
+ max(self.box_nms_thresh, self.crop_nms_thresh),
+ )
+
+ # Encode masks
+ if self.output_mode == 'coco_rle':
+ mask_data['segmentations'] = [
+ coco_encode_rle(rle) for rle in mask_data['rles']
+ ]
+ elif self.output_mode == 'binary_mask':
+ mask_data['segmentations'] = [
+ rle_to_mask(rle) for rle in mask_data['rles']
+ ]
+ else:
+ mask_data['segmentations'] = mask_data['rles']
+
+ # Write mask records
+ curr_anns = []
+ for idx in range(len(mask_data['segmentations'])):
+ ann = {
+ 'segmentation':
+ mask_data['segmentations'][idx],
+ 'area':
+ area_from_rle(mask_data['rles'][idx]),
+ 'bbox':
+ box_xyxy_to_xywh(mask_data['boxes'][idx]).tolist(),
+ 'predicted_iou':
+ mask_data['iou_preds'][idx].item(),
+ 'point_coords': [mask_data['points'][idx].tolist()],
+ 'stability_score':
+ mask_data['stability_score'][idx].item(),
+ 'crop_box':
+ box_xyxy_to_xywh(mask_data['crop_boxes'][idx]).tolist(),
+ }
+ curr_anns.append(ann)
+
+ return curr_anns
+
+ def _generate_masks(self, image: np.ndarray) -> MaskData:
+ orig_size = image.shape[:2]
+ crop_boxes, layer_idxs = generate_crop_boxes(orig_size,
+ self.crop_n_layers,
+ self.crop_overlap_ratio)
+
+ # Iterate over image crops
+ data = MaskData()
+ for crop_box, layer_idx in zip(crop_boxes, layer_idxs):
+ crop_data = self._process_crop(image, crop_box, layer_idx,
+ orig_size)
+ data.cat(crop_data)
+
+ # Remove duplicate masks between crops
+ if len(crop_boxes) > 1:
+ # Prefer masks from smaller crops
+ scores = 1 / box_area(data['crop_boxes'])
+ scores = scores.to(data['boxes'].device)
+ keep_by_nms = batched_nms(
+ data['boxes'].float(),
+ scores,
+ torch.zeros(len(data['boxes'])), # categories
+ iou_threshold=self.crop_nms_thresh,
+ )
+ data.filter(keep_by_nms)
+
+ data.to_numpy()
+ return data
+
+ def _process_crop(
+ self,
+ image: np.ndarray,
+ crop_box: List[int],
+ crop_layer_idx: int,
+ orig_size: Tuple[int, ...],
+ ) -> MaskData:
+ # Crop the image and calculate embeddings
+ x0, y0, x1, y1 = crop_box
+ cropped_im = image[y0:y1, x0:x1, :]
+ cropped_im_size = cropped_im.shape[:2]
+ self.predictor.set_image(cropped_im)
+
+ # Get points for this crop
+ points_scale = np.array(cropped_im_size)[None, ::-1]
+ points_for_image = self.point_grids[crop_layer_idx] * points_scale
+
+ # Generate masks for this crop in batches
+ data = MaskData()
+ for (points, ) in batch_iterator(self.points_per_batch,
+ points_for_image):
+ batch_data = self._process_batch(points, cropped_im_size, crop_box,
+ orig_size)
+ data.cat(batch_data)
+ del batch_data
+ self.predictor.reset_image()
+
+ # Remove duplicates within this crop.
+ keep_by_nms = batched_nms(
+ data['boxes'].float(),
+ data['iou_preds'],
+ torch.zeros(len(data['boxes'])), # categories
+ iou_threshold=self.box_nms_thresh,
+ )
+ data.filter(keep_by_nms)
+
+ # Return to the original image frame
+ data['boxes'] = uncrop_boxes_xyxy(data['boxes'], crop_box)
+ data['points'] = uncrop_points(data['points'], crop_box)
+ data['crop_boxes'] = torch.tensor(
+ [crop_box for _ in range(len(data['rles']))])
+
+ return data
+
+ def _process_batch(
+ self,
+ points: np.ndarray,
+ im_size: Tuple[int, ...],
+ crop_box: List[int],
+ orig_size: Tuple[int, ...],
+ ) -> MaskData:
+ orig_h, orig_w = orig_size
+
+ # Run model on this batch
+ transformed_points = self.predictor.transform.apply_coords(
+ points, im_size)
+ in_points = torch.as_tensor(
+ transformed_points, device=self.predictor.device)
+ in_labels = torch.ones(
+ in_points.shape[0], dtype=torch.int, device=in_points.device)
+ masks, iou_preds, _ = self.predictor.predict_torch(
+ in_points[:, None, :],
+ in_labels[:, None],
+ multimask_output=True,
+ return_logits=True,
+ )
+
+ # Serialize predictions and store in MaskData
+ data = MaskData(
+ masks=masks.flatten(0, 1),
+ iou_preds=iou_preds.flatten(0, 1),
+ points=torch.as_tensor(points.repeat(masks.shape[1], axis=0)),
+ )
+ del masks
+
+ # Filter by predicted IoU
+ if self.pred_iou_thresh > 0.0:
+ keep_mask = data['iou_preds'] > self.pred_iou_thresh
+ data.filter(keep_mask)
+
+ # Calculate stability score
+ data['stability_score'] = calculate_stability_score(
+ data['masks'], self.predictor.model.mask_threshold,
+ self.stability_score_offset)
+ if self.stability_score_thresh > 0.0:
+ keep_mask = data['stability_score'] >= self.stability_score_thresh
+ data.filter(keep_mask)
+
+ # Threshold masks and calculate boxes
+ data['masks'] = data['masks'] > self.predictor.model.mask_threshold
+ data['boxes'] = batched_mask_to_box(data['masks'])
+
+ # Filter boxes that touch crop boundaries
+ keep_mask = ~is_box_near_crop_edge(data['boxes'], crop_box,
+ [0, 0, orig_w, orig_h])
+ if not torch.all(keep_mask):
+ data.filter(keep_mask)
+
+ # Compress to RLE
+ data['masks'] = uncrop_masks(data['masks'], crop_box, orig_h, orig_w)
+ data['rles'] = mask_to_rle_pytorch(data['masks'])
+ del data['masks']
+
+ return data
+
+ @staticmethod
+ def postprocess_small_regions(mask_data: MaskData, min_area: int,
+ nms_thresh: float) -> MaskData:
+ """Removes small disconnected regions and holes in masks, then reruns
+ box NMS to remove any new duplicates.
+
+ Edits mask_data in place.
+
+ Requires open-cv as a dependency.
+ """
+ if len(mask_data['rles']) == 0:
+ return mask_data
+
+ # Filter small disconnected regions and holes
+ new_masks = []
+ scores = []
+ for rle in mask_data['rles']:
+ mask = rle_to_mask(rle)
+
+ mask, changed = remove_small_regions(mask, min_area, mode='holes')
+ unchanged = not changed
+ mask, changed = remove_small_regions(
+ mask, min_area, mode='islands')
+ unchanged = unchanged and not changed
+
+ new_masks.append(torch.as_tensor(mask).unsqueeze(0))
+ # Give score=0 to changed masks and score=1 to unchanged masks
+ # so NMS will prefer ones that didn't need postprocessing
+ scores.append(float(unchanged))
+
+ # Recalculate boxes and remove any new duplicates
+ masks = torch.cat(new_masks, dim=0)
+ boxes = batched_mask_to_box(masks)
+ keep_by_nms = batched_nms(
+ boxes.float(),
+ torch.as_tensor(scores),
+ torch.zeros(len(boxes)), # categories
+ iou_threshold=nms_thresh,
+ )
+
+ # Only recalculate RLEs for masks that have changed
+ for i_mask in keep_by_nms:
+ if scores[i_mask] == 0.0:
+ mask_torch = masks[i_mask].unsqueeze(0)
+ mask_data['rles'][i_mask] = mask_to_rle_pytorch(mask_torch)[0]
+ mask_data['boxes'][i_mask] = boxes[
+ i_mask] # update res directly
+ mask_data.filter(keep_by_nms)
+
+ return mask_data
diff --git a/projects/sam_inference_demo/sam/utils/__init__.py b/projects/sam_inference_demo/sam/utils/__init__.py
new file mode 100644
index 00000000000..5d33e33aeef
--- /dev/null
+++ b/projects/sam_inference_demo/sam/utils/__init__.py
@@ -0,0 +1,2 @@
+from .amg import * # noqa: F403 F401
+from .transforms import ResizeLongestSide # noqa: F403 F401
diff --git a/projects/sam_inference_demo/sam/utils/amg.py b/projects/sam_inference_demo/sam/utils/amg.py
new file mode 100644
index 00000000000..3ba359901f7
--- /dev/null
+++ b/projects/sam_inference_demo/sam/utils/amg.py
@@ -0,0 +1,355 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# https://github.com/facebookresearch/segment-anything
+
+import math
+from copy import deepcopy
+from itertools import product
+from typing import Any, Dict, Generator, ItemsView, List, Tuple
+
+import numpy as np
+import torch
+
+
+class MaskData:
+ """A structure for storing masks and their related data in batched format.
+
+ Implements basic filtering and concatenation.
+ """
+
+ def __init__(self, **kwargs) -> None:
+ for v in kwargs.values():
+ assert isinstance(
+ v, (list, np.ndarray, torch.Tensor)
+ ), 'MaskData only supports list, numpy arrays, and torch tensors.'
+ self._stats = dict(**kwargs)
+
+ def __setitem__(self, key: str, item: Any) -> None:
+ assert isinstance(
+ item, (list, np.ndarray, torch.Tensor)
+ ), 'MaskData only supports list, numpy arrays, and torch tensors.'
+ self._stats[key] = item
+
+ def __delitem__(self, key: str) -> None:
+ del self._stats[key]
+
+ def __getitem__(self, key: str) -> Any:
+ return self._stats[key]
+
+ def items(self) -> ItemsView[str, Any]:
+ return self._stats.items()
+
+ def filter(self, keep: torch.Tensor) -> None:
+ for k, v in self._stats.items():
+ if v is None:
+ self._stats[k] = None
+ elif isinstance(v, torch.Tensor):
+ self._stats[k] = v[torch.as_tensor(keep, device=v.device)]
+ elif isinstance(v, np.ndarray):
+ self._stats[k] = v[keep.detach().cpu().numpy()]
+ elif isinstance(v, list) and keep.dtype == torch.bool:
+ self._stats[k] = [a for i, a in enumerate(v) if keep[i]]
+ elif isinstance(v, list):
+ self._stats[k] = [v[i] for i in keep]
+ else:
+ raise TypeError(
+ f'MaskData key {k} has an unsupported type {type(v)}.')
+
+ def cat(self, new_stats: 'MaskData') -> None:
+ for k, v in new_stats.items():
+ if k not in self._stats or self._stats[k] is None:
+ self._stats[k] = deepcopy(v)
+ elif isinstance(v, torch.Tensor):
+ self._stats[k] = torch.cat([self._stats[k], v], dim=0)
+ elif isinstance(v, np.ndarray):
+ self._stats[k] = np.concatenate([self._stats[k], v], axis=0)
+ elif isinstance(v, list):
+ self._stats[k] = self._stats[k] + deepcopy(v)
+ else:
+ raise TypeError(
+ f'MaskData key {k} has an unsupported type {type(v)}.')
+
+ def to_numpy(self) -> None:
+ for k, v in self._stats.items():
+ if isinstance(v, torch.Tensor):
+ self._stats[k] = v.detach().cpu().numpy()
+
+
+def is_box_near_crop_edge(boxes: torch.Tensor,
+ crop_box: List[int],
+ orig_box: List[int],
+ atol: float = 20.0) -> torch.Tensor:
+ """Filter masks at the edge of a crop, but not at the edge of the original
+ image."""
+ crop_box_torch = torch.as_tensor(
+ crop_box, dtype=torch.float, device=boxes.device)
+ orig_box_torch = torch.as_tensor(
+ orig_box, dtype=torch.float, device=boxes.device)
+ boxes = uncrop_boxes_xyxy(boxes, crop_box).float()
+ near_crop_edge = torch.isclose(
+ boxes, crop_box_torch[None, :], atol=atol, rtol=0)
+ near_image_edge = torch.isclose(
+ boxes, orig_box_torch[None, :], atol=atol, rtol=0)
+ near_crop_edge = torch.logical_and(near_crop_edge, ~near_image_edge)
+ return torch.any(near_crop_edge, dim=1)
+
+
+def box_xyxy_to_xywh(box_xyxy: torch.Tensor) -> torch.Tensor:
+ box_xywh = deepcopy(box_xyxy)
+ box_xywh[2] = box_xywh[2] - box_xywh[0]
+ box_xywh[3] = box_xywh[3] - box_xywh[1]
+ return box_xywh
+
+
+def batch_iterator(batch_size: int, *args) -> Generator[List[Any], None, None]:
+ assert len(args) > 0 and all(
+ len(a) == len(args[0]) for a in
+ args), 'Batched iteration must have inputs of all the same size.'
+ n_batches = len(args[0]) // batch_size + int(
+ len(args[0]) % batch_size != 0)
+ for b in range(n_batches):
+ yield [arg[b * batch_size:(b + 1) * batch_size] for arg in args]
+
+
+def mask_to_rle_pytorch(tensor: torch.Tensor) -> List[Dict[str, Any]]:
+ """Encodes masks to an uncompressed RLE, in the format expected by pycoco
+ tools."""
+ # Put in fortran order and flatten h,w
+ b, h, w = tensor.shape
+ tensor = tensor.permute(0, 2, 1).flatten(1)
+
+ # Compute change indices
+ diff = tensor[:, 1:] ^ tensor[:, :-1]
+ change_indices = diff.nonzero()
+
+ # Encode run length
+ out = []
+ for i in range(b):
+ cur_idxs = change_indices[change_indices[:, 0] == i, 1]
+ cur_idxs = torch.cat([
+ torch.tensor([0], dtype=cur_idxs.dtype, device=cur_idxs.device),
+ cur_idxs + 1,
+ torch.tensor([h * w], dtype=cur_idxs.dtype,
+ device=cur_idxs.device),
+ ])
+ btw_idxs = cur_idxs[1:] - cur_idxs[:-1]
+ counts = [] if tensor[i, 0] == 0 else [0]
+ counts.extend(btw_idxs.detach().cpu().tolist())
+ out.append({'size': [h, w], 'counts': counts})
+ return out
+
+
+def rle_to_mask(rle: Dict[str, Any]) -> np.ndarray:
+ """Compute a binary mask from an uncompressed RLE."""
+ h, w = rle['size']
+ mask = np.empty(h * w, dtype=bool)
+ idx = 0
+ parity = False
+ for count in rle['counts']:
+ mask[idx:idx + count] = parity
+ idx += count
+ parity ^= True
+ mask = mask.reshape(w, h)
+ return mask.transpose() # Put in C order
+
+
+def area_from_rle(rle: Dict[str, Any]) -> int:
+ return sum(rle['counts'][1::2])
+
+
+def calculate_stability_score(masks: torch.Tensor, mask_threshold: float,
+ threshold_offset: float) -> torch.Tensor:
+ """Computes the stability score for a batch of masks.
+
+ The stability score is the IoU between the binary masks obtained by
+ thresholding the predicted mask logits at high and low values.
+ """
+ # One mask is always contained inside the other.
+ # Save memory by preventing unnecessary cast to torch.int64
+ intersections = ((masks > (mask_threshold + threshold_offset)).sum(
+ -1, dtype=torch.int16).sum(-1, dtype=torch.int32))
+ unions = ((masks > (mask_threshold - threshold_offset)).sum(
+ -1, dtype=torch.int16).sum(-1, dtype=torch.int32))
+ return intersections / unions
+
+
+def build_point_grid(n_per_side: int) -> np.ndarray:
+ """Generates a 2D grid of points evenly spaced in [0,1]x[0,1]."""
+ offset = 1 / (2 * n_per_side)
+ points_one_side = np.linspace(offset, 1 - offset, n_per_side)
+ points_x = np.tile(points_one_side[None, :], (n_per_side, 1))
+ points_y = np.tile(points_one_side[:, None], (1, n_per_side))
+ points = np.stack([points_x, points_y], axis=-1).reshape(-1, 2)
+ return points
+
+
+def build_all_layer_point_grids(n_per_side: int, n_layers: int,
+ scale_per_layer: int) -> List[np.ndarray]:
+ """Generates point grids for all crop layers."""
+ points_by_layer = []
+ for i in range(n_layers + 1):
+ n_points = int(n_per_side / (scale_per_layer**i))
+ points_by_layer.append(build_point_grid(n_points))
+ return points_by_layer
+
+
+def generate_crop_boxes(
+ im_size: Tuple[int, ...], n_layers: int,
+ overlap_ratio: float) -> Tuple[List[List[int]], List[int]]:
+ """Generates a list of crop boxes of different sizes.
+
+ Each layer has (2**i)**2 boxes for the ith layer.
+ """
+ crop_boxes, layer_idxs = [], []
+ im_h, im_w = im_size
+ short_side = min(im_h, im_w)
+
+ # Original image
+ crop_boxes.append([0, 0, im_w, im_h])
+ layer_idxs.append(0)
+
+ def crop_len(orig_len, n_crops, overlap):
+ return int(math.ceil((overlap * (n_crops - 1) + orig_len) / n_crops))
+
+ for i_layer in range(n_layers):
+ n_crops_per_side = 2**(i_layer + 1)
+ overlap = int(overlap_ratio * short_side * (2 / n_crops_per_side))
+
+ crop_w = crop_len(im_w, n_crops_per_side, overlap)
+ crop_h = crop_len(im_h, n_crops_per_side, overlap)
+
+ crop_box_x0 = [
+ int((crop_w - overlap) * i) for i in range(n_crops_per_side)
+ ]
+ crop_box_y0 = [
+ int((crop_h - overlap) * i) for i in range(n_crops_per_side)
+ ]
+
+ # Crops in XYWH format
+ for x0, y0 in product(crop_box_x0, crop_box_y0):
+ box = [x0, y0, min(x0 + crop_w, im_w), min(y0 + crop_h, im_h)]
+ crop_boxes.append(box)
+ layer_idxs.append(i_layer + 1)
+
+ return crop_boxes, layer_idxs
+
+
+def uncrop_boxes_xyxy(boxes: torch.Tensor,
+ crop_box: List[int]) -> torch.Tensor:
+ x0, y0, _, _ = crop_box
+ offset = torch.tensor([[x0, y0, x0, y0]], device=boxes.device)
+ # Check if boxes has a channel dimension
+ if len(boxes.shape) == 3:
+ offset = offset.unsqueeze(1)
+ return boxes + offset
+
+
+def uncrop_points(points: torch.Tensor, crop_box: List[int]) -> torch.Tensor:
+ x0, y0, _, _ = crop_box
+ offset = torch.tensor([[x0, y0]], device=points.device)
+ # Check if points has a channel dimension
+ if len(points.shape) == 3:
+ offset = offset.unsqueeze(1)
+ return points + offset
+
+
+def uncrop_masks(masks: torch.Tensor, crop_box: List[int], orig_h: int,
+ orig_w: int) -> torch.Tensor:
+ x0, y0, x1, y1 = crop_box
+ if x0 == 0 and y0 == 0 and x1 == orig_w and y1 == orig_h:
+ return masks
+ # Coordinate transform masks
+ pad_x, pad_y = orig_w - (x1 - x0), orig_h - (y1 - y0)
+ pad = (x0, pad_x - x0, y0, pad_y - y0)
+ return torch.nn.functional.pad(masks, pad, value=0)
+
+
+def remove_small_regions(mask: np.ndarray, area_thresh: float,
+ mode: str) -> Tuple[np.ndarray, bool]:
+ """Removes small disconnected regions and holes in a mask.
+
+ Returns the mask and an indicator of if the mask has been modified.
+ """
+ import cv2 # type: ignore
+
+ assert mode in ['holes', 'islands']
+ correct_holes = mode == 'holes'
+ working_mask = (correct_holes ^ mask).astype(np.uint8)
+ n_labels, regions, stats, _ = cv2.connectedComponentsWithStats(
+ working_mask, 8)
+ sizes = stats[:, -1][1:] # Row 0 is background label
+ small_regions = [i + 1 for i, s in enumerate(sizes) if s < area_thresh]
+ if len(small_regions) == 0:
+ return mask, False
+ fill_labels = [0] + small_regions
+ if not correct_holes:
+ fill_labels = [i for i in range(n_labels) if i not in fill_labels]
+ # If every region is below threshold, keep largest
+ if len(fill_labels) == 0:
+ fill_labels = [int(np.argmax(sizes)) + 1]
+ mask = np.isin(regions, fill_labels)
+ return mask, True
+
+
+def coco_encode_rle(uncompressed_rle: Dict[str, Any]) -> Dict[str, Any]:
+ from pycocotools import mask as mask_utils # type: ignore
+
+ h, w = uncompressed_rle['size']
+ rle = mask_utils.frPyObjects(uncompressed_rle, h, w)
+ rle['counts'] = rle['counts'].decode(
+ 'utf-8') # Necessary to serialize with json
+ return rle
+
+
+def batched_mask_to_box(masks: torch.Tensor) -> torch.Tensor:
+ """Calculates boxes in XYXY format around masks.
+
+ Return [0,0,0,0] for an empty mask. For input shape C1xC2x...xHxW, the
+ output shape is C1xC2x...x4.
+ """
+ # torch.max below raises an error on empty inputs, just skip in this case
+ if torch.numel(masks) == 0:
+ return torch.zeros(*masks.shape[:-2], 4, device=masks.device)
+
+ # Normalize shape to CxHxW
+ shape = masks.shape
+ h, w = shape[-2:]
+ if len(shape) > 2:
+ masks = masks.flatten(0, -3)
+ else:
+ masks = masks.unsqueeze(0)
+
+ # Get top and bottom edges
+ in_height, _ = torch.max(masks, dim=-1)
+ in_height_coords = in_height * torch.arange(
+ h, device=in_height.device)[None, :]
+ bottom_edges, _ = torch.max(in_height_coords, dim=-1)
+ in_height_coords = in_height_coords + h * (~in_height)
+ top_edges, _ = torch.min(in_height_coords, dim=-1)
+
+ # Get left and right edges
+ in_width, _ = torch.max(masks, dim=-2)
+ in_width_coords = in_width * torch.arange(
+ w, device=in_width.device)[None, :]
+ right_edges, _ = torch.max(in_width_coords, dim=-1)
+ in_width_coords = in_width_coords + w * (~in_width)
+ left_edges, _ = torch.min(in_width_coords, dim=-1)
+
+ # If the mask is empty the right edge will be to the left of the left edge.
+ # Replace these boxes with [0, 0, 0, 0]
+ empty_filter = (right_edges < left_edges) | (bottom_edges < top_edges)
+ out = torch.stack([left_edges, top_edges, right_edges, bottom_edges],
+ dim=-1)
+ out = out * (~empty_filter).unsqueeze(-1)
+
+ # Return to original shape
+ if len(shape) > 2:
+ out = out.reshape(*shape[:-2], 4)
+ else:
+ out = out[0]
+
+ return out
diff --git a/projects/sam_inference_demo/sam/utils/transforms.py b/projects/sam_inference_demo/sam/utils/transforms.py
new file mode 100644
index 00000000000..484fd6691cf
--- /dev/null
+++ b/projects/sam_inference_demo/sam/utils/transforms.py
@@ -0,0 +1,110 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+from copy import deepcopy
+from typing import Tuple
+
+import numpy as np
+import torch
+from torch.nn import functional as F
+from torchvision.transforms.functional import resize # type: ignore
+from torchvision.transforms.functional import to_pil_image
+
+from mmseg.registry import TRANSFORMS
+
+
+@TRANSFORMS.register_module()
+class ResizeLongestSide:
+ """Resizes images to longest side 'target_length', as well as provides
+ methods for resizing coordinates and boxes.
+
+ Provides methods for transforming both numpy array and batched torch
+ tensors.
+ """
+
+ def __init__(self, target_length: int) -> None:
+ self.target_length = target_length
+
+ def apply_image(self, image: np.ndarray) -> np.ndarray:
+ """Expects a numpy array with shape HxWxC in uint8 format."""
+ target_size = self.get_preprocess_shape(image.shape[0], image.shape[1],
+ self.target_length)
+ return np.array(resize(to_pil_image(image), target_size))
+
+ def apply_coords(self, coords: np.ndarray,
+ original_size: Tuple[int, ...]) -> np.ndarray:
+ """Expects a numpy array of length 2 in the final dimension.
+
+ Requires the original image size in (H, W) format.
+ """
+ old_h, old_w = original_size
+ new_h, new_w = self.get_preprocess_shape(original_size[0],
+ original_size[1],
+ self.target_length)
+ coords = deepcopy(coords).astype(float)
+ coords[..., 0] = coords[..., 0] * (new_w / old_w)
+ coords[..., 1] = coords[..., 1] * (new_h / old_h)
+ return coords
+
+ def apply_boxes(self, boxes: np.ndarray,
+ original_size: Tuple[int, ...]) -> np.ndarray:
+ """Expects a numpy array shape Bx4.
+
+ Requires the original image size in (H, W) format.
+ """
+ boxes = self.apply_coords(boxes.reshape(-1, 2, 2), original_size)
+ return boxes.reshape(-1, 4)
+
+ def apply_image_torch(self, image: torch.Tensor) -> torch.Tensor:
+ """Expects batched images with shape BxCxHxW and float format.
+
+ This transformation may not exactly match apply_image. apply_image is
+ the transformation expected by the model.
+ """
+ # Expects an image in BCHW format. May not exactly match apply_image.
+ target_size = self.get_preprocess_shape(image.shape[0], image.shape[1],
+ self.target_length)
+ return F.interpolate(
+ image,
+ target_size,
+ mode='bilinear',
+ align_corners=False,
+ antialias=True)
+
+ def apply_coords_torch(self, coords: torch.Tensor,
+ original_size: Tuple[int, ...]) -> torch.Tensor:
+ """Expects a torch tensor with length 2 in the last dimension.
+
+ Requires the original image size in (H, W) format.
+ """
+ old_h, old_w = original_size
+ new_h, new_w = self.get_preprocess_shape(original_size[0],
+ original_size[1],
+ self.target_length)
+ coords = deepcopy(coords).to(torch.float)
+ coords[..., 0] = coords[..., 0] * (new_w / old_w)
+ coords[..., 1] = coords[..., 1] * (new_h / old_h)
+ return coords
+
+ def apply_boxes_torch(self, boxes: torch.Tensor,
+ original_size: Tuple[int, ...]) -> torch.Tensor:
+ """Expects a torch tensor with shape Bx4.
+
+ Requires the original image size in (H, W) format.
+ """
+ boxes = self.apply_coords_torch(boxes.reshape(-1, 2, 2), original_size)
+ return boxes.reshape(-1, 4)
+
+ @staticmethod
+ def get_preprocess_shape(oldh: int, oldw: int,
+ long_side_length: int) -> Tuple[int, int]:
+ """Compute the output size given input size and target long side
+ length."""
+ scale = long_side_length * 1.0 / max(oldh, oldw)
+ newh, neww = oldh * scale, oldw * scale
+ neww = int(neww + 0.5)
+ newh = int(newh + 0.5)
+ return (newh, neww)
diff --git a/projects/sam_inference_demo/sam_image_demo.ipynb b/projects/sam_inference_demo/sam_image_demo.ipynb
new file mode 100644
index 00000000000..1cb433fae9b
--- /dev/null
+++ b/projects/sam_inference_demo/sam_image_demo.ipynb
@@ -0,0 +1,122 @@
+{
+ "cells": [
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import numpy as np\n",
+ "import matplotlib.pyplot as plt\n",
+ "import cv2\n",
+ "\n",
+ "import sam # noqa: F401\n",
+ "from sam.sam_inferencer import SAMInferencer\n",
+ "\n",
+ "\n",
+ "def show_mask(mask, ax, random_color=False):\n",
+ " if random_color:\n",
+ " color = np.concatenate([np.random.random(3), np.array([0.6])], axis=0)\n",
+ " else:\n",
+ " color = np.array([30/255, 144/255, 255/255, 0.6])\n",
+ " h, w = mask.shape[-2:]\n",
+ " mask_image = mask.reshape(h, w, 1) * color.reshape(1, 1, -1)\n",
+ " ax.imshow(mask_image)\n",
+ " \n",
+ "def show_points(coords, labels, ax, marker_size=375):\n",
+ " pos_points = coords[labels==1]\n",
+ " neg_points = coords[labels==0]\n",
+ " ax.scatter(pos_points[:, 0], pos_points[:, 1], color='green', marker='*', s=marker_size, edgecolor='white', linewidth=1.25)\n",
+ " ax.scatter(neg_points[:, 0], neg_points[:, 1], color='red', marker='*', s=marker_size, edgecolor='white', linewidth=1.25) \n",
+ " \n",
+ "def show_box(box, ax):\n",
+ " x0, y0 = box[0], box[1]\n",
+ " w, h = box[2] - box[0], box[3] - box[1]\n",
+ " ax.add_patch(plt.Rectangle((x0, y0), w, h, edgecolor='green', facecolor=(0,0,0,0), lw=2))\n",
+ "\n",
+ "image = cv2.imread('../../demo/demo.png')\n",
+ "image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)\n",
+ "plt.figure(figsize=(10,10))\n",
+ "plt.imshow(image)\n",
+ "plt.axis('on')\n",
+ "plt.show()\n",
+ "print(image.shape)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "inferencer = SAMInferencer(arch='huge')\n",
+ "inferencer.set_image(image)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "input_point = np.array([[280, 230], [500, 300]])\n",
+ "input_label = np.array([1, 1])\n",
+ "plt.figure(figsize=(10,10))\n",
+ "plt.imshow(image)\n",
+ "show_points(input_point, input_label, plt.gca())\n",
+ "plt.axis('on')\n",
+ "plt.show() "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "masks, scores, logits = inferencer.predict(\n",
+ " point_coords=input_point,\n",
+ " point_labels=input_label,\n",
+ " multimask_output=True,\n",
+ ")\n",
+ "for i, (mask, score) in enumerate(zip(masks, scores)):\n",
+ " plt.figure(figsize=(10,10))\n",
+ " plt.imshow(image)\n",
+ " show_mask(mask, plt.gca(), random_color=True)\n",
+ " show_points(input_point, input_label, plt.gca())\n",
+ " plt.title(f\"Mask {i+1}, Score: {score:.3f}\", fontsize=18)\n",
+ " plt.axis('off')\n",
+ " plt.show()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": []
+ }
+ ],
+ "metadata": {
+ "kernelspec": {
+ "display_name": "pt1.13",
+ "language": "python",
+ "name": "python3"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.10.9"
+ },
+ "orig_nbformat": 4
+ },
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
diff --git a/projects/van/README.md b/projects/van/README.md
new file mode 100644
index 00000000000..be0ba362faa
--- /dev/null
+++ b/projects/van/README.md
@@ -0,0 +1,101 @@
+# Visual Attention Network (VAN) for Segmentation
+
+This repo is a PyTorch implementation of applying **VAN** (**Visual Attention Network**) to semantic segmentation.
+
+The code is an integration from [VAN-Segmentation](https://github.com/Visual-Attention-Network/VAN-Segmentation/blob/main/README.md?plain=1)
+
+More details can be found in [**Visual Attention Network**](https://arxiv.org/abs/2202.09741).
+
+## Citation
+
+```bib
+@article{guo2022visual,
+ title={Visual Attention Network},
+ author={Guo, Meng-Hao and Lu, Cheng-Ze and Liu, Zheng-Ning and Cheng, Ming-Ming and Hu, Shi-Min},
+ journal={arXiv preprint arXiv:2202.09741},
+ year={2022}
+}
+```
+
+## Results
+
+**Notes**: Pre-trained models can be found in [TsingHua Cloud](https://cloud.tsinghua.edu.cn/d/0100f0cea37d41ba8d08/).
+
+Results can be found in [VAN-Segmentation](https://github.com/Visual-Attention-Network/VAN-Segmentation/blob/main/README.md?plain=1)
+
+We provide evaluation results of the converted weights.
+
+| Method | Backbone | mIoU | Download |
+| :-----: | :----------: | :---: | :--------------------------------------------------------------------------------------------------------------------------------------------: |
+| UPerNet | VAN-B2 | 49.35 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2-in1kpre_upernet_3rdparty_512x512-ade20k_20230522-19c58aee.pth) |
+| UPerNet | VAN-B3 | 49.71 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3-in1kpre_upernet_3rdparty_512x512-ade20k_20230522-653bd6b7.pth) |
+| UPerNet | VAN-B4 | 51.56 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4-in1kpre_upernet_3rdparty_512x512-ade20k_20230522-653bd6b7.pth) |
+| UPerNet | VAN-B4-in22k | 52.61 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4-in22kpre_upernet_3rdparty_512x512-ade20k_20230522-4a4d744a.pth) |
+| UPerNet | VAN-B5-in22k | 53.11 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b5-in22kpre_upernet_3rdparty_512x512-ade20k_20230522-5bb6f2b4.pth) |
+| UPerNet | VAN-B6-in22k | 54.25 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b6-in22kpre_upernet_3rdparty_512x512-ade20k_20230522-e226b363.pth) |
+| FPN | VAN-B0 | 38.65 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b0-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-75a76298.pth) |
+| FPN | VAN-B1 | 43.22 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b1-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-104499ff.pth) |
+| FPN | VAN-B2 | 46.84 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-7074e6f8.pth) |
+| FPN | VAN-B3 | 48.32 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-2c3b7f5e.pth) |
+
+## Preparation
+
+Install MMSegmentation and download ADE20K according to the guidelines in MMSegmentation.
+
+## Requirement
+
+**Step 0.** Install [MMCV](https://github.com/open-mmlab/mmcv) using [MIM](https://github.com/open-mmlab/mim).
+
+```shell
+pip install -U openmim
+mim install mmengine
+mim install "mmcv>=2.0.0"
+```
+
+**Step 1.** Install MMSegmentation.
+
+Case a: If you develop and run mmseg directly, install it from source:
+
+```shell
+git clone -b main https://github.com/open-mmlab/mmsegmentation.git
+cd mmsegmentation
+pip install -v -e .
+```
+
+Case b: If you use mmsegmentation as a dependency or third-party package, install it with pip:
+
+```shell
+pip install "mmsegmentation>=1.0.0"
+```
+
+## Training
+
+If you use 4 GPUs for training by default. Run:
+
+```bash
+bash tools/dist_train.sh projects/van/configs/van/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512.py 4
+```
+
+## Evaluation
+
+To evaluate the model, an example is:
+
+```bash
+bash tools/dist_train.sh projects/van/configs/van/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512.py work_dirs/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512/iter_160000.pth 4 --eval mIoU
+```
+
+## FLOPs
+
+To calculate FLOPs for a model, run:
+
+```bash
+bash tools/analysis_tools/get_flops.py projects/van/configs/van/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512.py --shape 512 512
+```
+
+## Acknowledgment
+
+Our implementation is mainly based on [mmsegmentation](https://github.com/open-mmlab/mmsegmentation/tree/v0.12.0), [Swin-Transformer](https://github.com/SwinTransformer/Swin-Transformer-Semantic-Segmentation), [PoolFormer](https://github.com/sail-sg/poolformer), [Enjoy-Hamburger](https://github.com/Gsunshine/Enjoy-Hamburger) and [VAN-Segmentation](https://github.com/Visual-Attention-Network/VAN-Segmentation/blob/main/README.md?plain=1). Thanks for their authors.
+
+## LICENSE
+
+This repo is under the Apache-2.0 license. For commercial use, please contact the authors.
diff --git a/projects/van/backbones/__init__.py b/projects/van/backbones/__init__.py
new file mode 100644
index 00000000000..071995de294
--- /dev/null
+++ b/projects/van/backbones/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .van import VAN
+
+__all__ = ['VAN']
diff --git a/projects/van/backbones/van.py b/projects/van/backbones/van.py
new file mode 100644
index 00000000000..301834a7587
--- /dev/null
+++ b/projects/van/backbones/van.py
@@ -0,0 +1,124 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import warnings
+
+import torch
+import torch.nn as nn
+from mmengine.model import BaseModule
+
+from mmseg.models.backbones.mscan import (MSCAN, MSCABlock,
+ MSCASpatialAttention,
+ OverlapPatchEmbed)
+from mmseg.registry import MODELS
+
+
+class VANAttentionModule(BaseModule):
+
+ def __init__(self, in_channels):
+ super().__init__()
+ self.conv0 = nn.Conv2d(
+ in_channels, in_channels, 5, padding=2, groups=in_channels)
+ self.conv_spatial = nn.Conv2d(
+ in_channels,
+ in_channels,
+ 7,
+ stride=1,
+ padding=9,
+ groups=in_channels,
+ dilation=3)
+ self.conv1 = nn.Conv2d(in_channels, in_channels, 1)
+
+ def forward(self, x):
+ u = x.clone()
+ attn = self.conv0(x)
+ attn = self.conv_spatial(attn)
+ attn = self.conv1(attn)
+ return u * attn
+
+
+class VANSpatialAttention(MSCASpatialAttention):
+
+ def __init__(self, in_channels, act_cfg=dict(type='GELU')):
+ super().__init__(in_channels, act_cfg=act_cfg)
+ self.spatial_gating_unit = VANAttentionModule(in_channels)
+
+
+class VANBlock(MSCABlock):
+
+ def __init__(self,
+ channels,
+ mlp_ratio=4.,
+ drop=0.,
+ drop_path=0.,
+ act_cfg=dict(type='GELU'),
+ norm_cfg=dict(type='SyncBN', requires_grad=True)):
+ super().__init__(
+ channels,
+ mlp_ratio=mlp_ratio,
+ drop=drop,
+ drop_path=drop_path,
+ act_cfg=act_cfg,
+ norm_cfg=norm_cfg)
+ self.attn = VANSpatialAttention(channels)
+
+
+@MODELS.register_module()
+class VAN(MSCAN):
+
+ def __init__(self,
+ in_channels=3,
+ embed_dims=[64, 128, 256, 512],
+ mlp_ratios=[8, 8, 4, 4],
+ drop_rate=0.,
+ drop_path_rate=0.,
+ depths=[3, 4, 6, 3],
+ num_stages=4,
+ act_cfg=dict(type='GELU'),
+ norm_cfg=dict(type='SyncBN', requires_grad=True),
+ pretrained=None,
+ init_cfg=None):
+ super(MSCAN, self).__init__(init_cfg=init_cfg)
+
+ assert not (init_cfg and pretrained), \
+ 'init_cfg and pretrained cannot be set at the same time'
+ if isinstance(pretrained, str):
+ warnings.warn('DeprecationWarning: pretrained is deprecated, '
+ 'please use "init_cfg" instead')
+ self.init_cfg = dict(type='Pretrained', checkpoint=pretrained)
+ elif pretrained is not None:
+ raise TypeError('pretrained must be a str or None')
+
+ self.depths = depths
+ self.num_stages = num_stages
+
+ # stochastic depth decay rule
+ dpr = [
+ x.item() for x in torch.linspace(0, drop_path_rate, sum(depths))
+ ]
+ cur = 0
+
+ for i in range(num_stages):
+ patch_embed = OverlapPatchEmbed(
+ patch_size=7 if i == 0 else 3,
+ stride=4 if i == 0 else 2,
+ in_channels=in_channels if i == 0 else embed_dims[i - 1],
+ embed_dim=embed_dims[i],
+ norm_cfg=norm_cfg)
+
+ block = nn.ModuleList([
+ VANBlock(
+ channels=embed_dims[i],
+ mlp_ratio=mlp_ratios[i],
+ drop=drop_rate,
+ drop_path=dpr[cur + j],
+ act_cfg=act_cfg,
+ norm_cfg=norm_cfg) for j in range(depths[i])
+ ])
+ norm = nn.LayerNorm(embed_dims[i])
+ cur += depths[i]
+
+ setattr(self, f'patch_embed{i + 1}', patch_embed)
+ setattr(self, f'block{i + 1}', block)
+ setattr(self, f'norm{i + 1}', norm)
+
+ def init_weights(self):
+ return super().init_weights()
diff --git a/projects/van/configs/_base_/datasets/ade20k.py b/projects/van/configs/_base_/datasets/ade20k.py
new file mode 100644
index 00000000000..69b3c2a73b8
--- /dev/null
+++ b/projects/van/configs/_base_/datasets/ade20k.py
@@ -0,0 +1,14 @@
+# dataset settings
+_base_ = '../../../../../configs/_base_/datasets/ade20k.py'
+
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 512), keep_ratio=True),
+ dict(type='ResizeToMultiple', size_divisor=32),
+ # add loading annotation after ``Resize`` because ground truth
+ # does not need to do resize data transform
+ dict(type='LoadAnnotations', reduce_zero_label=True),
+ dict(type='PackSegInputs')
+]
+val_dataloader = dict(dataset=dict(pipeline=test_pipeline))
+test_dataloader = val_dataloader
diff --git a/projects/van/configs/_base_/models/van_fpn.py b/projects/van/configs/_base_/models/van_fpn.py
new file mode 100644
index 00000000000..c7fd7391f77
--- /dev/null
+++ b/projects/van/configs/_base_/models/van_fpn.py
@@ -0,0 +1,43 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255,
+ size=(512, 512))
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ type='VAN',
+ embed_dims=[32, 64, 160, 256],
+ drop_rate=0.0,
+ drop_path_rate=0.1,
+ depths=[3, 3, 5, 2],
+ act_cfg=dict(type='GELU'),
+ norm_cfg=norm_cfg,
+ init_cfg=dict()),
+ neck=dict(
+ type='FPN',
+ in_channels=[32, 64, 160, 256],
+ out_channels=256,
+ num_outs=4),
+ decode_head=dict(
+ type='FPNHead',
+ in_channels=[256, 256, 256, 256],
+ in_index=[0, 1, 2, 3],
+ feature_strides=[4, 8, 16, 32],
+ channels=128,
+ dropout_ratio=0.1,
+ num_classes=150,
+ norm_cfg=norm_cfg,
+ align_corners=False,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0)),
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/van/configs/_base_/models/van_upernet.py b/projects/van/configs/_base_/models/van_upernet.py
new file mode 100644
index 00000000000..8f94c0d9d84
--- /dev/null
+++ b/projects/van/configs/_base_/models/van_upernet.py
@@ -0,0 +1,51 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255,
+ size=(512, 512))
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ type='VAN',
+ embed_dims=[32, 64, 160, 256],
+ drop_rate=0.0,
+ drop_path_rate=0.1,
+ depths=[3, 3, 5, 2],
+ act_cfg=dict(type='GELU'),
+ norm_cfg=norm_cfg,
+ init_cfg=dict()),
+ decode_head=dict(
+ type='UPerHead',
+ in_channels=[32, 64, 160, 256],
+ in_index=[0, 1, 2, 3],
+ pool_scales=(1, 2, 3, 6),
+ channels=512,
+ dropout_ratio=0.1,
+ num_classes=150,
+ norm_cfg=norm_cfg,
+ align_corners=False,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0)),
+ auxiliary_head=dict(
+ type='FCNHead',
+ in_channels=160,
+ in_index=2,
+ channels=256,
+ num_convs=1,
+ concat_input=False,
+ dropout_ratio=0.1,
+ num_classes=150,
+ norm_cfg=norm_cfg,
+ align_corners=False,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=0.4)),
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/van/configs/van/van-b0_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b0_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..2faf3788a71
--- /dev/null
+++ b/projects/van/configs/van/van-b0_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,8 @@
+_base_ = './van-b2_fpn_8xb4-40k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b0_3rdparty_20230522-956f5e0d.pth' # noqa
+model = dict(
+ backbone=dict(
+ embed_dims=[32, 64, 160, 256],
+ depths=[3, 3, 5, 2],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path)),
+ neck=dict(in_channels=[32, 64, 160, 256]))
diff --git a/projects/van/configs/van/van-b1_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b1_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..cf64a7138b2
--- /dev/null
+++ b/projects/van/configs/van/van-b1_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,6 @@
+_base_ = './van-b2_fpn_8xb4-40k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b1_3rdparty_20230522-3adb117f.pth' # noqa
+model = dict(
+ backbone=dict(
+ depths=[2, 2, 4, 2],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path)))
diff --git a/projects/van/configs/van/van-b2_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b2_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..965fa1cd363
--- /dev/null
+++ b/projects/van/configs/van/van-b2_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,53 @@
+_base_ = [
+ '../_base_/models/van_fpn.py',
+ '../_base_/datasets/ade20k.py',
+ '../../../../configs/_base_/default_runtime.py',
+]
+custom_imports = dict(imports=['projects.van.backbones'])
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2_3rdparty_20230522-636fac93.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ embed_dims=[64, 128, 320, 512],
+ depths=[3, 3, 12, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.2),
+ neck=dict(in_channels=[64, 128, 320, 512]),
+ decode_head=dict(num_classes=150))
+
+train_dataloader = dict(batch_size=4)
+
+# we use 8 gpu instead of 4 in mmsegmentation, so lr*2 and max_iters/2
+gpu_multiples = 2
+max_iters = 80000 // gpu_multiples
+interval = 8000 // gpu_multiples
+optim_wrapper = dict(
+ type='OptimWrapper',
+ optimizer=dict(
+ type='AdamW',
+ lr=0.0001 * gpu_multiples,
+ # betas=(0.9, 0.999),
+ weight_decay=0.0001),
+ clip_grad=None)
+# learning policy
+param_scheduler = [
+ dict(
+ type='PolyLR',
+ power=0.9,
+ eta_min=0.0,
+ begin=0,
+ end=max_iters,
+ by_epoch=False,
+ )
+]
+train_cfg = dict(
+ type='IterBasedTrainLoop', max_iters=max_iters, val_interval=interval)
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
+default_hooks = dict(
+ timer=dict(type='IterTimerHook'),
+ logger=dict(type='LoggerHook', interval=50, log_metric_by_epoch=False),
+ param_scheduler=dict(type='ParamSchedulerHook'),
+ checkpoint=dict(type='CheckpointHook', by_epoch=False, interval=interval),
+ sampler_seed=dict(type='DistSamplerSeedHook'),
+ visualization=dict(type='SegVisualizationHook'))
diff --git a/projects/van/configs/van/van-b2_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b2_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..c529606a202
--- /dev/null
+++ b/projects/van/configs/van/van-b2_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,46 @@
+_base_ = [
+ '../_base_/models/van_upernet.py', '../_base_/datasets/ade20k.py',
+ '../../../../configs/_base_/default_runtime.py',
+ '../../../../configs/_base_/schedules/schedule_160k.py'
+]
+custom_imports = dict(imports=['projects.van.backbones'])
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2_3rdparty_20230522-636fac93.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ embed_dims=[64, 128, 320, 512],
+ depths=[3, 3, 12, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path)),
+ decode_head=dict(in_channels=[64, 128, 320, 512], num_classes=150),
+ auxiliary_head=dict(in_channels=320, num_classes=150))
+
+# AdamW optimizer
+# no weight decay for position embedding & layer norm in backbone
+optim_wrapper = dict(
+ _delete_=True,
+ type='OptimWrapper',
+ optimizer=dict(
+ type='AdamW', lr=0.00006, betas=(0.9, 0.999), weight_decay=0.01),
+ clip_grad=None,
+ paramwise_cfg=dict(
+ custom_keys={
+ 'absolute_pos_embed': dict(decay_mult=0.),
+ 'relative_position_bias_table': dict(decay_mult=0.),
+ 'norm': dict(decay_mult=0.)
+ }))
+# learning policy
+param_scheduler = [
+ dict(
+ type='LinearLR', start_factor=1e-6, by_epoch=False, begin=0, end=1500),
+ dict(
+ type='PolyLR',
+ power=1.0,
+ begin=1500,
+ end=_base_.train_cfg.max_iters,
+ eta_min=0.0,
+ by_epoch=False,
+ )
+]
+
+# By default, models are trained on 8 GPUs with 2 images per GPU
+train_dataloader = dict(batch_size=2)
diff --git a/projects/van/configs/van/van-b3_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b3_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..b0493fe4f9f
--- /dev/null
+++ b/projects/van/configs/van/van-b3_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,11 @@
+_base_ = './van-b2_fpn_8xb4-40k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3_3rdparty_20230522-a184e051.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ embed_dims=[64, 128, 320, 512],
+ depths=[3, 5, 27, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.3),
+ neck=dict(in_channels=[64, 128, 320, 512]))
+train_dataloader = dict(batch_size=4)
diff --git a/projects/van/configs/van/van-b3_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b3_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..8201801d992
--- /dev/null
+++ b/projects/van/configs/van/van-b3_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,8 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3_3rdparty_20230522-a184e051.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ depths=[3, 5, 27, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.3))
diff --git a/projects/van/configs/van/van-b4-in22kpre_upernet_4xb4-160k_ade20k-512x512.py b/projects/van/configs/van/van-b4-in22kpre_upernet_4xb4-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..15c8f7ca6ee
--- /dev/null
+++ b/projects/van/configs/van/van-b4-in22kpre_upernet_4xb4-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4-in22k_3rdparty_20230522-5e31cafb.pth' # noqa
+model = dict(
+ backbone=dict(
+ depths=[3, 6, 40, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.4))
+
+# By default, models are trained on 8 GPUs with 2 images per GPU
+train_dataloader = dict(batch_size=4)
diff --git a/projects/van/configs/van/van-b4_upernet_4xb4-160k_ade20k-512x512.py b/projects/van/configs/van/van-b4_upernet_4xb4-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..33ae049d0c0
--- /dev/null
+++ b/projects/van/configs/van/van-b4_upernet_4xb4-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4_3rdparty_20230522-1d71c077.pth' # noqa
+model = dict(
+ backbone=dict(
+ depths=[3, 6, 40, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.4))
+
+# By default, models are trained on 4 GPUs with 4 images per GPU
+train_dataloader = dict(batch_size=4)
diff --git a/projects/van/configs/van/van-b5-in22kpre_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b5-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..f36c6242bdf
--- /dev/null
+++ b/projects/van/configs/van/van-b5-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b5-in22k_3rdparty_20230522-b26134d7.pth' # noqa
+model = dict(
+ backbone=dict(
+ embed_dims=[96, 192, 480, 768],
+ depths=[3, 3, 24, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.4),
+ decode_head=dict(in_channels=[96, 192, 480, 768], num_classes=150),
+ auxiliary_head=dict(in_channels=480, num_classes=150))
diff --git a/projects/van/configs/van/van-b6-in22kpre_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b6-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..aa529efed8c
--- /dev/null
+++ b/projects/van/configs/van/van-b6-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b6-in22k_3rdparty_20230522-5e5172a3.pth' # noqa
+model = dict(
+ backbone=dict(
+ embed_dims=[96, 192, 384, 768],
+ depths=[6, 6, 90, 6],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.5),
+ decode_head=dict(in_channels=[96, 192, 384, 768], num_classes=150),
+ auxiliary_head=dict(in_channels=384, num_classes=150))
diff --git a/requirements.txt b/requirements.txt
index 6da5adea757..501bddc8843 100644
--- a/requirements.txt
+++ b/requirements.txt
@@ -1,3 +1,4 @@
-r requirements/optional.txt
-r requirements/runtime.txt
-r requirements/tests.txt
+-r requirements/multimodal.txt
diff --git a/requirements/albu.txt b/requirements/albu.txt
new file mode 100644
index 00000000000..f421fbbdc47
--- /dev/null
+++ b/requirements/albu.txt
@@ -0,0 +1 @@
+albumentations>=0.3.2 --no-binary qudida,albumentations
diff --git a/requirements/docs.txt b/requirements/docs.txt
index 8e98c16fc72..19632d36aba 100644
--- a/requirements/docs.txt
+++ b/requirements/docs.txt
@@ -4,3 +4,4 @@ myst-parser
sphinx==4.0.2
sphinx_copybutton
sphinx_markdown_tables
+urllib3<2.0.0
diff --git a/requirements/multimodal.txt b/requirements/multimodal.txt
new file mode 100644
index 00000000000..2195d0d9ef8
--- /dev/null
+++ b/requirements/multimodal.txt
@@ -0,0 +1,2 @@
+ftfy
+regex
diff --git a/requirements/optional.txt b/requirements/optional.txt
index 5eca649247e..b0310f52960 100644
--- a/requirements/optional.txt
+++ b/requirements/optional.txt
@@ -1,2 +1,22 @@
cityscapesscripts
+-e git+https://github.com/openai/CLIP.git@main#egg=clip
+
+# for vpd model
+diffusers
+einops==0.3.0
+imageio==2.9.0
+imageio-ffmpeg==0.4.2
+invisible-watermark
+kornia==0.6
+-e git+https://github.com/CompVis/stable-diffusion@21f890f#egg=latent-diffusion
nibabel
+omegaconf==2.1.1
+pudb==2019.2
+pytorch-lightning==1.4.2
+streamlit>=0.73.1
+-e git+https://github.com/CompVis/taming-transformers.git@master#egg=taming-transformers
+test-tube>=0.7.5
+timm
+torch-fidelity==0.3.0
+torchmetrics==0.6.0
+transformers==4.19.2
diff --git a/requirements/tests.txt b/requirements/tests.txt
index 74fc76146d8..3fff2520d7c 100644
--- a/requirements/tests.txt
+++ b/requirements/tests.txt
@@ -1,6 +1,8 @@
codecov
flake8
+ftfy
interrogate
pytest
+regex
xdoctest>=0.10.0
yapf
diff --git a/resources/miaomiao_qrcode.jpg b/resources/miaomiao_qrcode.jpg
new file mode 100644
index 00000000000..d34cbae6fd1
Binary files /dev/null and b/resources/miaomiao_qrcode.jpg differ
diff --git a/setup.cfg b/setup.cfg
index dc5ea071110..2ea07600c0b 100644
--- a/setup.cfg
+++ b/setup.cfg
@@ -16,4 +16,4 @@ default_section = THIRDPARTY
skip = *.po,*.ts,*.ipynb
count =
quiet-level = 3
-ignore-words-list = formating,sur,hist,dota,warmup
+ignore-words-list = formating,sur,hist,dota,warmup,damon
diff --git a/setup.py b/setup.py
index 854dd186054..45d923db60e 100755
--- a/setup.py
+++ b/setup.py
@@ -123,7 +123,7 @@ def add_mim_extension():
else:
return
- filenames = ['tools', 'configs', 'model-index.yml']
+ filenames = ['tools', 'configs', 'model-index.yml', 'dataset-index.yml']
repo_path = osp.dirname(__file__)
mim_path = osp.join(repo_path, 'mmseg', '.mim')
os.makedirs(mim_path, exist_ok=True)
@@ -175,7 +175,7 @@ def add_mim_extension():
author='MMSegmentation Contributors',
author_email='openmmlab@gmail.com',
keywords='computer vision, semantic segmentation',
- url='http://github.com/open-mmlab/mmsegmentation',
+ url='https://github.com/open-mmlab/mmsegmentation',
packages=find_packages(exclude=('configs', 'tools', 'demo')),
include_package_data=True,
classifiers=[
@@ -194,6 +194,7 @@ def add_mim_extension():
'tests': parse_requirements('requirements/tests.txt'),
'optional': parse_requirements('requirements/optional.txt'),
'mim': parse_requirements('requirements/mminstall.txt'),
+ 'multimodal': parse_requirements('requirements/multimodal.txt'),
},
ext_modules=[],
zip_safe=False)
diff --git a/tests/data/dsdl_seg/config.py b/tests/data/dsdl_seg/config.py
new file mode 100755
index 00000000000..8eed751c2f7
--- /dev/null
+++ b/tests/data/dsdl_seg/config.py
@@ -0,0 +1,13 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+local = dict(
+ type='LocalFileReader',
+ working_dir='/nvme/share_data/VOC2012',
+)
+
+ali_oss = dict(
+ type='AliOSSFileReader',
+ access_key_secret='your secret key of aliyun oss',
+ endpoint='your endpoint of aliyun oss',
+ access_key_id='your access key of aliyun oss',
+ bucket_name='your bucket name of aliyun oss',
+ working_dir='the relative path of your media dir in the bucket')
diff --git a/tests/data/dsdl_seg/defs/class-dom.yaml b/tests/data/dsdl_seg/defs/class-dom.yaml
new file mode 100755
index 00000000000..e5dd598c4ae
--- /dev/null
+++ b/tests/data/dsdl_seg/defs/class-dom.yaml
@@ -0,0 +1,24 @@
+$dsdl-version: "0.5.0"
+VOCClassDom:
+ $def: class_domain
+ classes:
+ - aeroplane
+ - bicycle
+ - bird
+ - boat
+ - bottle
+ - bus
+ - car
+ - cat
+ - chair
+ - cow
+ - diningtable
+ - dog
+ - horse
+ - motorbike
+ - person
+ - pottedplant
+ - sheep
+ - sofa
+ - train
+ - tvmonitor
diff --git a/tests/data/dsdl_seg/defs/segmentation-def.yaml b/tests/data/dsdl_seg/defs/segmentation-def.yaml
new file mode 100755
index 00000000000..057139ed57e
--- /dev/null
+++ b/tests/data/dsdl_seg/defs/segmentation-def.yaml
@@ -0,0 +1,15 @@
+$dsdl-version: "0.5.0"
+
+ImageMedia:
+ $def: struct
+ $fields:
+ image: Image
+ image_shape: ImageShape
+
+SegmentationSample:
+ $def: struct
+ $params: ['cdom']
+ $fields:
+ media: ImageMedia
+ label_map: LabelMap[dom=$cdom]
+ instance_map: InstanceMap
diff --git a/tests/data/dsdl_seg/set-train/train.yaml b/tests/data/dsdl_seg/set-train/train.yaml
new file mode 100755
index 00000000000..69872445a53
--- /dev/null
+++ b/tests/data/dsdl_seg/set-train/train.yaml
@@ -0,0 +1,15 @@
+$dsdl-version: "0.5.0"
+$import:
+ - ../defs/segmentation-def
+ - ../defs/class-dom
+meta:
+ dataset_name: "VOC2012"
+ sub_dataset_name: "train"
+ task_type: "Segmentation"
+ dataset_homepage: "http://host.robots.ox.ac.uk/pascal/VOC/voc2012/index.html"
+ dataset_publisher: "University of Leeds | ETHZ, Zurich | University of Edinburgh\
+ \ |Microsoft Research Cambridge | University of Oxford"
+ OpenDataLab_address: "https://opendatalab.com/PASCAL_VOC2012/download"
+data:
+ sample-type: SegmentationSample[cdom=VOCClassDom]
+ sample-path: train_samples.json
diff --git a/tests/data/dsdl_seg/set-train/train_samples.json b/tests/data/dsdl_seg/set-train/train_samples.json
new file mode 100755
index 00000000000..559f5845721
--- /dev/null
+++ b/tests/data/dsdl_seg/set-train/train_samples.json
@@ -0,0 +1 @@
+{"samples": [{"media": {"image": "JPEGImages/2007_000032.jpg", "image_shape": [281, 500]}, "label_map": "SegmentationClass/2007_000032.png", "instance_map": "SegmentationObject/2007_000032.png"}, {"media": {"image": "JPEGImages/2007_000039.jpg", "image_shape": [375, 500]}, "label_map": "SegmentationClass/2007_000039.png", "instance_map": "SegmentationObject/2007_000039.png"}, {"media": {"image": "JPEGImages/2007_000063.jpg", "image_shape": [375, 500]}, "label_map": "SegmentationClass/2007_000063.png", "instance_map": "SegmentationObject/2007_000063.png"}]}
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/train/0004a4c0-d4dff0ad.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/train/0004a4c0-d4dff0ad.jpg
new file mode 100644
index 00000000000..4724a3d9302
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/train/0004a4c0-d4dff0ad.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/train/00054602-3bf57337.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00054602-3bf57337.jpg
new file mode 100644
index 00000000000..5efe06b99a4
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00054602-3bf57337.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/train/00067cfb-e535423e.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00067cfb-e535423e.jpg
new file mode 100644
index 00000000000..2233c03c763
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00067cfb-e535423e.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d06fefd-f7be05a6.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d06fefd-f7be05a6.jpg
new file mode 100644
index 00000000000..535087b5a95
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d06fefd-f7be05a6.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d128593-0ccfea4c.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d128593-0ccfea4c.jpg
new file mode 100644
index 00000000000..7f2971afdee
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d128593-0ccfea4c.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d15b18b-1e0d6e3f.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d15b18b-1e0d6e3f.jpg
new file mode 100644
index 00000000000..31a951d4830
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d15b18b-1e0d6e3f.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/0004a4c0-d4dff0ad.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/0004a4c0-d4dff0ad.png
new file mode 100644
index 00000000000..086a8d50647
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/0004a4c0-d4dff0ad.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00054602-3bf57337.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00054602-3bf57337.png
new file mode 100644
index 00000000000..43338c283cc
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00054602-3bf57337.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00067cfb-e535423e.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00067cfb-e535423e.png
new file mode 100644
index 00000000000..7c0ad1d5d90
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00067cfb-e535423e.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d128593-0ccfea4c.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d128593-0ccfea4c.png
new file mode 100644
index 00000000000..43338c283cc
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d128593-0ccfea4c.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d15b18b-1e0d6e3f.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d15b18b-1e0d6e3f.png
new file mode 100644
index 00000000000..43338c283cc
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d15b18b-1e0d6e3f.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d2f7975-e0c1c5a7.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d2f7975-e0c1c5a7.png
new file mode 100644
index 00000000000..7c0ad1d5d90
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d2f7975-e0c1c5a7.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/0004a4c0-d4dff0ad.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/0004a4c0-d4dff0ad.png
new file mode 100644
index 00000000000..5c6bf5e1581
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/0004a4c0-d4dff0ad.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00054602-3bf57337.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00054602-3bf57337.png
new file mode 100644
index 00000000000..c525a768885
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00054602-3bf57337.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00067cfb-e535423e.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00067cfb-e535423e.png
new file mode 100644
index 00000000000..7dfd3af4e3a
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00067cfb-e535423e.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d06fefd-f7be05a6.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d06fefd-f7be05a6.png
new file mode 100644
index 00000000000..7dfd3af4e3a
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d06fefd-f7be05a6.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d128593-0ccfea4c.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d128593-0ccfea4c.png
new file mode 100644
index 00000000000..c525a768885
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d128593-0ccfea4c.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d15b18b-1e0d6e3f.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d15b18b-1e0d6e3f.png
new file mode 100644
index 00000000000..c525a768885
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d15b18b-1e0d6e3f.png differ
diff --git a/tests/data/pseudo_nyu_dataset/annotations/bookstore_0001d_00001.png b/tests/data/pseudo_nyu_dataset/annotations/bookstore_0001d_00001.png
new file mode 100644
index 00000000000..77e343603ad
Binary files /dev/null and b/tests/data/pseudo_nyu_dataset/annotations/bookstore_0001d_00001.png differ
diff --git a/tests/data/pseudo_nyu_dataset/images/bookstore_0001d_00001.jpg b/tests/data/pseudo_nyu_dataset/images/bookstore_0001d_00001.jpg
new file mode 100644
index 00000000000..7892ed47e7d
Binary files /dev/null and b/tests/data/pseudo_nyu_dataset/images/bookstore_0001d_00001.jpg differ
diff --git a/tests/test_apis/test_inferencer.py b/tests/test_apis/test_inferencer.py
index 497eae4a011..d8dbce8f385 100644
--- a/tests/test_apis/test_inferencer.py
+++ b/tests/test_apis/test_inferencer.py
@@ -3,76 +3,16 @@
import numpy as np
import torch
-import torch.nn as nn
from mmengine import ConfigDict
-from torch.utils.data import DataLoader, Dataset
+from utils import * # noqa: F401, F403
from mmseg.apis import MMSegInferencer
-from mmseg.models import EncoderDecoder
-from mmseg.models.decode_heads.decode_head import BaseDecodeHead
from mmseg.registry import MODELS
from mmseg.utils import register_all_modules
-@MODELS.register_module(name='InferExampleHead')
-class ExampleDecodeHead(BaseDecodeHead):
-
- def __init__(self, num_classes=19, out_channels=None):
- super().__init__(
- 3, 3, num_classes=num_classes, out_channels=out_channels)
-
- def forward(self, inputs):
- return self.cls_seg(inputs[0])
-
-
-@MODELS.register_module(name='InferExampleBackbone')
-class ExampleBackbone(nn.Module):
-
- def __init__(self):
- super().__init__()
- self.conv = nn.Conv2d(3, 3, 3)
-
- def init_weights(self, pretrained=None):
- pass
-
- def forward(self, x):
- return [self.conv(x)]
-
-
-@MODELS.register_module(name='InferExampleModel')
-class ExampleModel(EncoderDecoder):
-
- def __init__(self, **kwargs):
- super().__init__(**kwargs)
-
-
-class ExampleDataset(Dataset):
-
- def __init__(self) -> None:
- super().__init__()
- self.pipeline = [
- dict(type='LoadImageFromFile'),
- dict(type='LoadAnnotations'),
- dict(type='PackSegInputs')
- ]
-
- def __getitem__(self, idx):
- return dict(img=torch.tensor([1]), img_metas=dict())
-
- def __len__(self):
- return 1
-
-
def test_inferencer():
register_all_modules()
- test_dataset = ExampleDataset()
- data_loader = DataLoader(
- test_dataset,
- batch_size=1,
- sampler=None,
- num_workers=0,
- shuffle=False,
- )
visualizer = dict(
type='SegLocalVisualizer',
@@ -87,7 +27,14 @@ def test_inferencer():
decode_head=dict(type='InferExampleHead'),
test_cfg=dict(mode='whole')),
visualizer=visualizer,
- test_dataloader=data_loader)
+ test_dataloader=dict(
+ dataset=dict(
+ type='ExampleDataset',
+ pipeline=[
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+ ]), ))
cfg = ConfigDict(cfg_dict)
model = MODELS.build(cfg.model)
diff --git a/tests/test_apis/test_rs_inferencer.py b/tests/test_apis/test_rs_inferencer.py
new file mode 100644
index 00000000000..03423d9680e
--- /dev/null
+++ b/tests/test_apis/test_rs_inferencer.py
@@ -0,0 +1,73 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import os.path as osp
+from unittest import TestCase
+
+import numpy as np
+from mmengine import ConfigDict, init_default_scope
+from utils import * # noqa: F401, F403
+
+from mmseg.apis import RSImage, RSInferencer
+from mmseg.registry import MODELS
+
+
+class TestRSImage(TestCase):
+
+ def test_read_whole_image(self):
+ init_default_scope('mmseg')
+ img_path = osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_loveda_dataset/img_dir/0.png')
+ rs_image = RSImage(img_path)
+ window_size = (16, 16)
+ rs_image.create_grids(window_size)
+ image_data = rs_image.read(rs_image.grids[0])
+ self.assertIsNotNone(image_data)
+
+ def test_write_image_data(self):
+ init_default_scope('mmseg')
+ img_path = osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_loveda_dataset/img_dir/0.png')
+ rs_image = RSImage(img_path)
+ window_size = (16, 16)
+ rs_image.create_grids(window_size)
+ data = np.random.random((16, 16)).astype(np.int8)
+ rs_image.write(data, rs_image.grids[0])
+
+
+class TestRSInferencer(TestCase):
+
+ def test_read_and_inference(self):
+ init_default_scope('mmseg')
+ cfg_dict = dict(
+ model=dict(
+ type='InferExampleModel',
+ data_preprocessor=dict(type='SegDataPreProcessor'),
+ backbone=dict(type='InferExampleBackbone'),
+ decode_head=dict(type='InferExampleHead'),
+ test_cfg=dict(mode='whole')),
+ test_dataloader=dict(
+ dataset=dict(
+ type='ExampleDataset',
+ pipeline=[
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+ ])),
+ test_pipeline=[
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+ ])
+ cfg = ConfigDict(cfg_dict)
+ model = MODELS.build(cfg.model)
+ model.cfg = cfg
+ inferencer = RSInferencer.from_model(model)
+
+ img_path = osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_loveda_dataset/img_dir/0.png')
+ rs_image = RSImage(img_path)
+ window_size = (16, 16)
+ stride = (16, 16)
+ inferencer.run(rs_image, window_size, stride)
diff --git a/tests/test_apis/utils.py b/tests/test_apis/utils.py
new file mode 100644
index 00000000000..0a9928fccf1
--- /dev/null
+++ b/tests/test_apis/utils.py
@@ -0,0 +1,38 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch.nn as nn
+
+from mmseg.models import EncoderDecoder
+from mmseg.models.decode_heads.decode_head import BaseDecodeHead
+from mmseg.registry import MODELS
+
+
+@MODELS.register_module(name='InferExampleHead')
+class ExampleDecodeHead(BaseDecodeHead):
+
+ def __init__(self, num_classes=19, out_channels=None):
+ super().__init__(
+ 3, 3, num_classes=num_classes, out_channels=out_channels)
+
+ def forward(self, inputs):
+ return self.cls_seg(inputs[0])
+
+
+@MODELS.register_module(name='InferExampleBackbone')
+class ExampleBackbone(nn.Module):
+
+ def __init__(self):
+ super().__init__()
+ self.conv = nn.Conv2d(3, 3, 3)
+
+ def init_weights(self, pretrained=None):
+ pass
+
+ def forward(self, x):
+ return [self.conv(x)]
+
+
+@MODELS.register_module(name='InferExampleModel')
+class ExampleModel(EncoderDecoder):
+
+ def __init__(self, **kwargs):
+ super().__init__(**kwargs)
diff --git a/tests/test_config.py b/tests/test_config.py
index 13de460181c..cdd85ff57cd 100644
--- a/tests/test_config.py
+++ b/tests/test_config.py
@@ -30,6 +30,7 @@ def _get_config_directory():
def test_config_build_segmentor():
"""Test that all segmentation models defined in the configs can be
initialized."""
+ init_default_scope('mmseg')
config_dpath = _get_config_directory()
print(f'Found config_dpath = {config_dpath!r}')
@@ -92,7 +93,12 @@ def test_config_data_pipeline():
del config_mod.train_pipeline[0]
del config_mod.test_pipeline[0]
# remove loading annotation in test pipeline
- del config_mod.test_pipeline[-2]
+ load_anno_idx = -1
+ for i in range(len(config_mod.test_pipeline)):
+ if config_mod.test_pipeline[i].type in ('LoadAnnotations',
+ 'LoadDepthAnnotation'):
+ load_anno_idx = i
+ del config_mod.test_pipeline[load_anno_idx]
train_pipeline = Compose(config_mod.train_pipeline)
test_pipeline = Compose(config_mod.test_pipeline)
@@ -101,6 +107,7 @@ def test_config_data_pipeline():
if to_float32:
img = img.astype(np.float32)
seg = np.random.randint(0, 255, size=(1024, 2048, 1), dtype=np.uint8)
+ depth = np.random.rand(1024, 2048).astype(np.float32)
results = dict(
filename='test_img.png',
@@ -108,24 +115,30 @@ def test_config_data_pipeline():
img=img,
img_shape=img.shape,
ori_shape=img.shape,
- gt_seg_map=seg)
+ gt_seg_map=seg,
+ gt_depth_map=depth)
results['seg_fields'] = ['gt_seg_map']
-
+ _check_concat_cd_input(config_mod, results)
print(f'Test training data pipeline: \n{train_pipeline!r}')
output_results = train_pipeline(results)
assert output_results is not None
- results = dict(
- filename='test_img.png',
- ori_filename='test_img.png',
- img=img,
- img_shape=img.shape,
- ori_shape=img.shape)
+ _check_concat_cd_input(config_mod, results)
print(f'Test testing data pipeline: \n{test_pipeline!r}')
output_results = test_pipeline(results)
assert output_results is not None
+def _check_concat_cd_input(config_mod: Config, results: dict):
+ keys = []
+ pipeline = config_mod.train_pipeline.copy()
+ pipeline.extend(config_mod.test_pipeline)
+ for t in pipeline:
+ keys.append(t.type)
+ if 'ConcatCDInput' in keys:
+ results.update({'img2': results['img']})
+
+
def _check_decode_head(decode_head_cfg, decode_head):
if isinstance(decode_head_cfg, list):
assert isinstance(decode_head, nn.ModuleList)
@@ -149,14 +162,14 @@ def _check_decode_head(decode_head_cfg, decode_head):
elif input_transform == 'resize_concat':
assert sum(in_channels) == decode_head.in_channels
else:
- assert isinstance(in_channels, int)
assert in_channels == decode_head.in_channels
- assert isinstance(decode_head.in_index, int)
if decode_head_cfg['type'] == 'PointHead':
assert decode_head_cfg.channels+decode_head_cfg.num_classes == \
decode_head.fc_seg.in_channels
assert decode_head.fc_seg.out_channels == decode_head_cfg.num_classes
+ elif decode_head_cfg['type'] == 'VPDDepthHead':
+ assert decode_head.out_channels == 1
else:
assert decode_head_cfg.channels == decode_head.conv_seg.in_channels
assert decode_head.conv_seg.out_channels == decode_head_cfg.num_classes
diff --git a/tests/test_datasets/test_dataset.py b/tests/test_datasets/test_dataset.py
index db4a7799069..2904e09cedd 100644
--- a/tests/test_datasets/test_dataset.py
+++ b/tests/test_datasets/test_dataset.py
@@ -5,15 +5,21 @@
import pytest
-from mmseg.datasets import (ADE20KDataset, BaseSegDataset, CityscapesDataset,
- COCOStuffDataset, DecathlonDataset, ISPRSDataset,
+from mmseg.datasets import (ADE20KDataset, BaseSegDataset, BDD100KDataset,
+ CityscapesDataset, COCOStuffDataset,
+ DecathlonDataset, DSDLSegDataset, ISPRSDataset,
LIPDataset, LoveDADataset, MapillaryDataset_v1,
- MapillaryDataset_v2, PascalVOCDataset,
+ MapillaryDataset_v2, NYUDataset, PascalVOCDataset,
PotsdamDataset, REFUGEDataset, SynapseDataset,
iSAIDDataset)
from mmseg.registry import DATASETS
from mmseg.utils import get_classes, get_palette
+try:
+ from dsdl.dataset import DSDLDataset
+except ImportError:
+ DSDLDataset = None
+
def test_classes():
assert list(
@@ -32,7 +38,7 @@ def test_classes():
MapillaryDataset_v1.METAINFO['classes']) == get_classes('mapillary_v1')
assert list(
MapillaryDataset_v2.METAINFO['classes']) == get_classes('mapillary_v2')
-
+ assert list(BDD100KDataset.METAINFO['classes']) == get_classes('bdd100k')
with pytest.raises(ValueError):
get_classes('unsupported')
@@ -89,7 +95,7 @@ def test_palette():
MapillaryDataset_v1.METAINFO['palette']) == get_palette('mapillary_v1')
assert list(
MapillaryDataset_v2.METAINFO['palette']) == get_palette('mapillary_v2')
-
+ assert list(BDD100KDataset.METAINFO['palette']) == get_palette('bdd100k')
with pytest.raises(ValueError):
get_palette('unsupported')
@@ -326,6 +332,19 @@ def test_mapillary():
assert len(test_dataset) == 1
+def test_bdd100k():
+ test_dataset = BDD100KDataset(
+ pipeline=[],
+ data_prefix=dict(
+ img_path=osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_bdd100k_dataset/images/10k/val'),
+ seg_map_path=osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val')))
+ assert len(test_dataset) == 3
+
+
@pytest.mark.parametrize('dataset, classes', [
('ADE20KDataset', ('wall', 'building')),
('CityscapesDataset', ('road', 'sidewalk')),
@@ -433,3 +452,24 @@ def test_custom_dataset_custom_palette():
ann_file=tempfile.mkdtemp(),
metainfo=dict(classes=('bus', 'car'), palette=[[200, 200, 200]]),
lazy_init=True)
+
+
+def test_dsdlseg_dataset():
+ if DSDLDataset is not None:
+ dataset = DSDLSegDataset(
+ data_root='tests/data/dsdl_seg', ann_file='set-train/train.yaml')
+ assert len(dataset) == 3
+ assert len(dataset.metainfo['classes']) == 21
+ else:
+ ImportWarning('Package `dsdl` is not installed.')
+
+
+def test_nyu_dataset():
+ dataset = NYUDataset(
+ data_root='tests/data/pseudo_nyu_dataset',
+ data_prefix=dict(img_path='images', depth_map_path='annotations'),
+ )
+ assert len(dataset) == 1
+ data = dataset[0]
+ assert data.get('depth_map_path', None) is not None
+ assert data.get('category_id', -1) == 26
diff --git a/tests/test_datasets/test_loading.py b/tests/test_datasets/test_loading.py
index 5ce624bff6a..3eea6e3f9dd 100644
--- a/tests/test_datasets/test_loading.py
+++ b/tests/test_datasets/test_loading.py
@@ -7,10 +7,11 @@
import numpy as np
from mmcv.transforms import LoadImageFromFile
-from mmseg.datasets.transforms import (LoadAnnotations,
- LoadBiomedicalAnnotation,
+from mmseg.datasets.transforms import LoadAnnotations # noqa
+from mmseg.datasets.transforms import (LoadBiomedicalAnnotation,
LoadBiomedicalData,
LoadBiomedicalImageFromFile,
+ LoadDepthAnnotation,
LoadImageFromNDArray)
@@ -276,3 +277,19 @@ def test_load_biomedical_data(self):
"decode_backend='numpy', "
'to_xyz=False, '
'backend_args=None)')
+
+ def test_load_depth_annotation(self):
+ input_results = dict(
+ img_path='tests/data/pseudo_nyu_dataset/images/'
+ 'bookstore_0001d_00001.jpg',
+ depth_map_path='tests/data/pseudo_nyu_dataset/'
+ 'annotations/bookstore_0001d_00001.png',
+ category_id=-1,
+ seg_fields=[])
+ transform = LoadDepthAnnotation(depth_rescale_factor=0.001)
+ results = transform(input_results)
+ assert 'gt_depth_map' in results
+ assert results['gt_depth_map'].shape[:2] == mmcv.imread(
+ input_results['depth_map_path']).shape[:2]
+ assert results['gt_depth_map'].dtype == np.float32
+ assert 'gt_depth_map' in results['seg_fields']
diff --git a/tests/test_datasets/test_transform.py b/tests/test_datasets/test_transform.py
index 92d6c6106d2..e73e558ee8e 100644
--- a/tests/test_datasets/test_transform.py
+++ b/tests/test_datasets/test_transform.py
@@ -1,6 +1,7 @@
# Copyright (c) OpenMMLab. All rights reserved.
import copy
import os.path as osp
+from unittest import TestCase
import mmcv
import numpy as np
@@ -11,7 +12,8 @@
from mmseg.datasets.transforms import * # noqa
from mmseg.datasets.transforms import (LoadBiomedicalData,
LoadBiomedicalImageFromFile,
- PhotoMetricDistortion, RandomCrop)
+ PhotoMetricDistortion, RandomCrop,
+ RandomDepthMix)
from mmseg.registry import TRANSFORMS
init_default_scope('mmseg')
@@ -184,6 +186,14 @@ def test_flip():
assert np.equal(original_img, results['img']).all()
assert np.equal(original_seg, results['gt_semantic_seg']).all()
+ results['gt_depth_map'] = seg
+ results['seg_fields'] = ['gt_depth_map']
+ results = flip_module(results)
+ flip_module = TRANSFORMS.build(transform)
+ results = flip_module(results)
+ assert np.equal(original_img, results['img']).all()
+ assert np.equal(original_seg, results['gt_depth_map']).all()
+
def test_random_rotate_flip():
with pytest.raises(AssertionError):
@@ -1160,3 +1170,104 @@ def test_biomedical_3d_flip():
results = transform(results)
assert np.equal(original_img, results['img']).all()
assert np.equal(original_seg, results['gt_seg_map']).all()
+
+
+def test_albu_transform():
+ results = dict(
+ img_path=osp.join(osp.dirname(__file__), '../data/color.jpg'))
+
+ # Define simple pipeline
+ load = dict(type='LoadImageFromFile')
+ load = TRANSFORMS.build(load)
+
+ albu_transform = dict(
+ type='Albu', transforms=[dict(type='ChannelShuffle', p=1)])
+ albu_transform = TRANSFORMS.build(albu_transform)
+
+ normalize = dict(type='Normalize', mean=[0] * 3, std=[0] * 3, to_rgb=True)
+ normalize = TRANSFORMS.build(normalize)
+
+ # Execute transforms
+ results = load(results)
+ results = albu_transform(results)
+ results = normalize(results)
+
+ assert results['img'].dtype == np.float32
+
+
+def test_albu_channel_order():
+ results = dict(
+ img_path=osp.join(osp.dirname(__file__), '../data/color.jpg'))
+
+ # Define simple pipeline
+ load = dict(type='LoadImageFromFile')
+ load = TRANSFORMS.build(load)
+
+ # Transform is modifying B channel
+ albu_transform = dict(
+ type='Albu',
+ transforms=[
+ dict(
+ type='RGBShift',
+ r_shift_limit=0,
+ g_shift_limit=0,
+ b_shift_limit=200,
+ p=1)
+ ])
+ albu_transform = TRANSFORMS.build(albu_transform)
+
+ # Execute transforms
+ results_load = load(results)
+ results_albu = albu_transform(results_load)
+
+ # assert only Green and Red channel are not modified
+ np.testing.assert_array_equal(results_albu['img'][..., 1:],
+ results_load['img'][..., 1:])
+
+ # assert Blue channel is modified
+ with pytest.raises(AssertionError):
+ np.testing.assert_array_equal(results_albu['img'][..., 0],
+ results_load['img'][..., 0])
+
+
+class TestRandomDepthMix(TestCase):
+
+ def setUp(self):
+ self.transform = RandomDepthMix(prob=1.0)
+
+ def test_transform_shape(self):
+ # Create a dummy result dict
+ results = {
+ 'img_shape': (10, 10),
+ 'img': np.random.rand(10, 10, 3),
+ 'gt_depth_map': np.random.rand(10, 10)
+ }
+ transformed = self.transform.transform(results)
+
+ # Check if the shape remains the same
+ self.assertEqual(results['img'].shape, transformed['img'].shape)
+
+ def test_transform_values(self):
+ # Create a dummy result dict
+ results = {
+ 'img_shape': (10, 10),
+ 'img': np.zeros((10, 10, 3)),
+ 'gt_depth_map': np.ones((10, 10))
+ }
+ transformed = self.transform.transform(results)
+
+ # Assuming the transformation modifies a portion of the image,
+ # it shouldn't remain all zeros
+ self.assertFalse(np.all(transformed['img'] == 0))
+
+ def test_invalid_image_dimension(self):
+ # Create a dummy result dict with invalid image dimension
+ results = {
+ 'img_shape': (10, 10),
+ 'img': np.random.rand(10, 10, 3, 3),
+ 'gt_depth_map': np.random.rand(10, 10)
+ }
+
+ # Check if a ValueError is raised for invalid dimension
+ with self.assertRaises(ValueError):
+ self.transform.transform(results)
diff --git a/tests/test_evaluation/test_metrics/test_depth_metric.py b/tests/test_evaluation/test_metrics/test_depth_metric.py
new file mode 100644
index 00000000000..a172db8fa20
--- /dev/null
+++ b/tests/test_evaluation/test_metrics/test_depth_metric.py
@@ -0,0 +1,85 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import os.path as osp
+import shutil
+from unittest import TestCase
+
+import torch
+from mmengine.structures import PixelData
+
+from mmseg.evaluation import DepthMetric
+from mmseg.structures import SegDataSample
+
+
+class TestDepthMetric(TestCase):
+
+ def _demo_mm_inputs(self,
+ batch_size=2,
+ image_shapes=(3, 64, 64),
+ num_classes=5):
+ """Create a superset of inputs needed to run test or train batches.
+
+ Args:
+ batch_size (int): batch size. Default to 2.
+ image_shapes (List[tuple], Optional): image shape.
+ Default to (3, 64, 64)
+ num_classes (int): number of different classes.
+ Default to 5.
+ """
+ if isinstance(image_shapes, list):
+ assert len(image_shapes) == batch_size
+ else:
+ image_shapes = [image_shapes] * batch_size
+
+ data_samples = []
+ for idx in range(batch_size):
+ image_shape = image_shapes[idx]
+ _, h, w = image_shape
+
+ data_sample = SegDataSample()
+ gt_depth_map = torch.rand((1, h, w)) * 10
+ data_sample.gt_depth_map = PixelData(data=gt_depth_map)
+
+ data_samples.append(data_sample.to_dict())
+
+ return data_samples
+
+ def _demo_mm_model_output(self,
+ data_samples,
+ batch_size=2,
+ image_shapes=(3, 64, 64),
+ num_classes=5):
+
+ _, h, w = image_shapes
+
+ for data_sample in data_samples:
+ data_sample['pred_depth_map'] = dict(data=torch.randn(1, h, w))
+
+ data_sample[
+ 'img_path'] = 'tests/data/pseudo_dataset/imgs/00000_img.jpg'
+ return data_samples
+
+ def test_evaluate(self):
+ """Test using the metric in the same way as Evalutor."""
+
+ data_samples = self._demo_mm_inputs()
+ data_samples = self._demo_mm_model_output(data_samples)
+
+ depth_metric = DepthMetric()
+ depth_metric.process([0] * len(data_samples), data_samples)
+ res = depth_metric.compute_metrics(depth_metric.results)
+ self.assertIsInstance(res, dict)
+
+ # test save depth map file in output_dir
+ depth_metric = DepthMetric(output_dir='tmp')
+ depth_metric.process([0] * len(data_samples), data_samples)
+ assert osp.exists('tmp')
+ assert osp.isfile('tmp/00000_img.png')
+ shutil.rmtree('tmp')
+
+ # test format_only
+ depth_metric = DepthMetric(output_dir='tmp', format_only=True)
+ depth_metric.process([0] * len(data_samples), data_samples)
+ assert depth_metric.results == []
+ assert osp.exists('tmp')
+ assert osp.isfile('tmp/00000_img.png')
+ shutil.rmtree('tmp')
diff --git a/tests/test_models/test_assigners/test_hungarian_assigner.py b/tests/test_models/test_assigners/test_hungarian_assigner.py
new file mode 100644
index 00000000000..2cdb1de839d
--- /dev/null
+++ b/tests/test_models/test_assigners/test_hungarian_assigner.py
@@ -0,0 +1,77 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from unittest import TestCase
+
+import torch
+from mmengine.structures import InstanceData
+
+from mmseg.models.assigners import HungarianAssigner
+
+
+class TestHungarianAssigner(TestCase):
+
+ def test_init(self):
+ with self.assertRaises(AssertionError):
+ HungarianAssigner([])
+
+ def test_hungarian_match_assigner(self):
+ assigner = HungarianAssigner([
+ dict(type='ClassificationCost', weight=2.0),
+ dict(type='CrossEntropyLossCost', weight=5.0, use_sigmoid=True),
+ dict(type='DiceCost', weight=5.0, pred_act=True, eps=1.0)
+ ])
+ num_classes = 3
+ num_masks = 10
+ num_points = 20
+ gt_instances = InstanceData()
+ gt_instances.labels = torch.randint(0, num_classes, (num_classes, ))
+ gt_instances.masks = torch.randint(0, 2, (num_classes, num_points))
+ pred_instances = InstanceData()
+ pred_instances.scores = torch.rand((num_masks, num_classes))
+ pred_instances.masks = torch.rand((num_masks, num_points))
+
+ matched_quiery_inds, matched_label_inds = \
+ assigner.assign(pred_instances, gt_instances)
+ unique_quiery_inds = torch.unique(matched_quiery_inds)
+ unique_label_inds = torch.unique(matched_label_inds)
+ self.assertTrue(len(unique_quiery_inds) == len(matched_quiery_inds))
+ self.assertTrue(
+ torch.equal(unique_label_inds, torch.arange(0, num_classes)))
+
+ def test_cls_match_cost(self):
+ num_classes = 3
+ num_masks = 10
+ gt_instances = InstanceData()
+ gt_instances.labels = torch.randint(0, num_classes, (num_classes, ))
+ pred_instances = InstanceData()
+ pred_instances.scores = torch.rand((num_masks, num_classes))
+
+ # test ClassificationCost
+ assigner = HungarianAssigner(dict(type='ClassificationCost'))
+ matched_quiery_inds, matched_label_inds = \
+ assigner.assign(pred_instances, gt_instances)
+ unique_quiery_inds = torch.unique(matched_quiery_inds)
+ unique_label_inds = torch.unique(matched_label_inds)
+ self.assertTrue(len(unique_quiery_inds) == len(matched_quiery_inds))
+ self.assertTrue(
+ torch.equal(unique_label_inds, torch.arange(0, num_classes)))
+
+ def test_mask_match_cost(self):
+ num_classes = 3
+ num_masks = 10
+ num_points = 20
+ gt_instances = InstanceData()
+ gt_instances.masks = torch.randint(0, 2, (num_classes, num_points))
+ pred_instances = InstanceData()
+ pred_instances.masks = torch.rand((num_masks, num_points))
+
+ # test DiceCost
+ assigner = HungarianAssigner(
+ dict(type='DiceCost', pred_act=True, eps=1.0))
+ assign_result = assigner.assign(pred_instances, gt_instances)
+ self.assertTrue(len(assign_result[0]) == len(assign_result[1]))
+
+ # test CrossEntropyLossCost
+ assigner = HungarianAssigner(
+ dict(type='CrossEntropyLossCost', use_sigmoid=True))
+ assign_result = assigner.assign(pred_instances, gt_instances)
+ self.assertTrue(len(assign_result[0]) == len(assign_result[1]))
diff --git a/tests/test_models/test_backbones/test_clip_text_encoder.py b/tests/test_models/test_backbones/test_clip_text_encoder.py
new file mode 100644
index 00000000000..ea06c5b5b3f
--- /dev/null
+++ b/tests/test_models/test_backbones/test_clip_text_encoder.py
@@ -0,0 +1,43 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch
+from mmengine import Config
+from mmengine.registry import init_default_scope
+
+from mmseg.models.text_encoder import CLIPTextEncoder
+from mmseg.utils import get_classes
+
+
+def test_clip_text_encoder():
+ init_default_scope('mmseg')
+ # test vocabulary
+ output_dims = 8
+ embed_dims = 32
+ vocabulary = ['cat', 'dog', 'bird', 'car', 'bike']
+ cfg = dict(
+ vocabulary=vocabulary,
+ templates=['a photo of a {}.'],
+ embed_dims=embed_dims,
+ output_dims=output_dims)
+ cfg = Config(cfg)
+
+ text_encoder = CLIPTextEncoder(**cfg)
+ if torch.cuda.is_available():
+ text_encoder = text_encoder.cuda()
+
+ with torch.no_grad():
+ class_embeds = text_encoder()
+ assert class_embeds.shape == (len(vocabulary) + 1, output_dims)
+
+ # test dataset name
+ cfg = dict(
+ dataset_name='vaihingen',
+ templates=['a photo of a {}.'],
+ embed_dims=embed_dims,
+ output_dims=output_dims)
+ cfg = Config(cfg)
+
+ text_encoder = CLIPTextEncoder(**cfg)
+ with torch.no_grad():
+ class_embeds = text_encoder()
+ class_nums = len(get_classes('vaihingen'))
+ assert class_embeds.shape == (class_nums + 1, output_dims)
diff --git a/tests/test_models/test_backbones/test_vpd.py b/tests/test_models/test_backbones/test_vpd.py
new file mode 100644
index 00000000000..a268159155e
--- /dev/null
+++ b/tests/test_models/test_backbones/test_vpd.py
@@ -0,0 +1,51 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from os.path import dirname, join
+from unittest import TestCase
+
+import torch
+from mmengine import Config
+
+import mmseg
+from mmseg.models.backbones import VPD
+
+
+class TestVPD(TestCase):
+
+ def setUp(self) -> None:
+
+ repo_dpath = dirname(dirname(mmseg.__file__))
+ config_dpath = join(repo_dpath, 'configs/_base_/models/vpd_sd.py')
+ vpd_cfg = Config.fromfile(config_dpath).stable_diffusion_cfg
+ vpd_cfg.pop('checkpoint')
+
+ self.vpd_model = VPD(
+ diffusion_cfg=vpd_cfg,
+ class_embed_path='https://download.openmmlab.com/mmsegmentation/'
+ 'v0.5/vpd/nyu_class_embeddings.pth',
+ class_embed_select=True,
+ pad_shape=64,
+ unet_cfg=dict(use_attn=False),
+ )
+
+ def test_forward(self):
+ # test forward without class_id
+ x = torch.randn(1, 3, 60, 60)
+ with torch.no_grad():
+ out = self.vpd_model(x)
+
+ self.assertEqual(len(out), 4)
+ self.assertListEqual(list(out[0].shape), [1, 320, 8, 8])
+ self.assertListEqual(list(out[1].shape), [1, 640, 4, 4])
+ self.assertListEqual(list(out[2].shape), [1, 1280, 2, 2])
+ self.assertListEqual(list(out[3].shape), [1, 1280, 1, 1])
+
+ # test forward with class_id
+ x = torch.randn(1, 3, 60, 60)
+ with torch.no_grad():
+ out = self.vpd_model((x, torch.tensor([2])))
+
+ self.assertEqual(len(out), 4)
+ self.assertListEqual(list(out[0].shape), [1, 320, 8, 8])
+ self.assertListEqual(list(out[1].shape), [1, 640, 4, 4])
+ self.assertListEqual(list(out[2].shape), [1, 1280, 2, 2])
+ self.assertListEqual(list(out[3].shape), [1, 1280, 1, 1])
diff --git a/tests/test_models/test_heads/test_san_head.py b/tests/test_models/test_heads/test_san_head.py
new file mode 100644
index 00000000000..af85a6e2ca0
--- /dev/null
+++ b/tests/test_models/test_heads/test_san_head.py
@@ -0,0 +1,126 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch
+from mmengine import Config
+from mmengine.structures import PixelData
+
+from mmseg.models.decode_heads import SideAdapterCLIPHead
+from mmseg.structures import SegDataSample
+from .utils import list_to_cuda
+
+
+def test_san_head():
+ H, W = (64, 64)
+ clip_channels = 64
+ img_channels = 4
+ num_queries = 40
+ out_dims = 64
+ num_classes = 19
+ cfg = dict(
+ num_classes=num_classes,
+ deep_supervision_idxs=[4],
+ san_cfg=dict(
+ in_channels=img_channels,
+ embed_dims=128,
+ clip_channels=clip_channels,
+ num_queries=num_queries,
+ cfg_encoder=dict(num_encode_layer=4, mlp_ratio=2, num_heads=2),
+ cfg_decoder=dict(
+ num_heads=4,
+ num_layers=1,
+ embed_channels=32,
+ mlp_channels=32,
+ num_mlp=2,
+ rescale=True)),
+ maskgen_cfg=dict(
+ sos_token_num=num_queries,
+ embed_dims=clip_channels,
+ out_dims=out_dims,
+ num_heads=4,
+ mlp_ratio=2),
+ train_cfg=dict(
+ num_points=100,
+ oversample_ratio=3.0,
+ importance_sample_ratio=0.75,
+ assigner=dict(
+ type='HungarianAssigner',
+ match_costs=[
+ dict(type='ClassificationCost', weight=2.0),
+ dict(
+ type='CrossEntropyLossCost',
+ weight=5.0,
+ use_sigmoid=True),
+ dict(type='DiceCost', weight=5.0, pred_act=True, eps=1.0)
+ ])),
+ loss_decode=[
+ dict(
+ type='CrossEntropyLoss',
+ loss_name='loss_cls_ce',
+ loss_weight=2.0,
+ class_weight=[1.0] * num_classes + [0.1]),
+ dict(
+ type='CrossEntropyLoss',
+ use_sigmoid=True,
+ loss_name='loss_mask_ce',
+ loss_weight=5.0),
+ dict(
+ type='DiceLoss',
+ ignore_index=None,
+ naive_dice=True,
+ eps=1,
+ loss_name='loss_mask_dice',
+ loss_weight=5.0)
+ ])
+
+ cfg = Config(cfg)
+ head = SideAdapterCLIPHead(**cfg)
+
+ inputs = torch.rand((2, img_channels, H, W))
+ clip_feature = [[
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ],
+ [
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ],
+ [
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ],
+ [
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ]]
+ class_embed = torch.rand((num_classes + 1, out_dims))
+
+ data_samples = []
+ for i in range(2):
+ data_sample = SegDataSample()
+ img_meta = {}
+ img_meta['img_shape'] = (H, W)
+ img_meta['ori_shape'] = (H, W)
+ data_sample.gt_sem_seg = PixelData(
+ data=torch.randint(0, num_classes, (1, H, W)))
+ data_sample.set_metainfo(img_meta)
+ data_samples.append(data_sample)
+
+ batch_img_metas = []
+ for data_sample in data_samples:
+ batch_img_metas.append(data_sample.metainfo)
+
+ if torch.cuda.is_available():
+ head = head.cuda()
+ data = list_to_cuda([inputs, clip_feature, class_embed])
+ for data_sample in data_samples:
+ data_sample.gt_sem_seg.data = data_sample.gt_sem_seg.data.cuda()
+ else:
+ data = [inputs, clip_feature, class_embed]
+
+ # loss test
+ loss_dict = head.loss(data, data_samples, None)
+ assert isinstance(loss_dict, dict)
+
+ # prediction test
+ with torch.no_grad():
+ seg_logits = head.predict(data, batch_img_metas, None)
+ assert seg_logits.shape == torch.Size((2, num_classes, H, W))
diff --git a/tests/test_models/test_heads/test_vpd_depth_head.py b/tests/test_models/test_heads/test_vpd_depth_head.py
new file mode 100644
index 00000000000..e3a4f7558ec
--- /dev/null
+++ b/tests/test_models/test_heads/test_vpd_depth_head.py
@@ -0,0 +1,50 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from unittest import TestCase
+
+import torch
+from mmengine.structures import PixelData
+
+from mmseg.models.decode_heads import VPDDepthHead
+from mmseg.structures import SegDataSample
+
+
+class TestVPDDepthHead(TestCase):
+
+ def setUp(self):
+ """Set up common resources."""
+ self.in_channels = [320, 640, 1280, 1280]
+ self.max_depth = 10.0
+ self.loss_decode = dict(
+ type='SiLogLoss'
+ ) # Replace with your actual loss type and parameters
+ self.vpd_depth_head = VPDDepthHead(
+ max_depth=self.max_depth,
+ in_channels=self.in_channels,
+ loss_decode=self.loss_decode)
+
+ def test_forward(self):
+ """Test the forward method."""
+ # Create a mock input tensor. Replace shape as per your needs.
+ x = [
+ torch.randn(1, 320, 32, 32),
+ torch.randn(1, 640, 16, 16),
+ torch.randn(1, 1280, 8, 8),
+ torch.randn(1, 1280, 4, 4)
+ ]
+
+ output = self.vpd_depth_head.forward(x)
+ print(output.shape)
+
+ self.assertEqual(output.shape, (1, 1, 256, 256))
+
+ def test_loss_by_feat(self):
+ """Test the loss_by_feat method."""
+ # Create mock data for `pred_depth_map` and `batch_data_samples`.
+ pred_depth_map = torch.randn(1, 1, 32, 32)
+ gt_depth_map = PixelData(data=torch.rand(1, 32, 32))
+ batch_data_samples = [SegDataSample(gt_depth_map=gt_depth_map)]
+
+ loss = self.vpd_depth_head.loss_by_feat(pred_depth_map,
+ batch_data_samples)
+
+ self.assertIsNotNone(loss)
diff --git a/tests/test_models/test_heads/utils.py b/tests/test_models/test_heads/utils.py
index 335e261a5e5..72823401552 100644
--- a/tests/test_models/test_heads/utils.py
+++ b/tests/test_models/test_heads/utils.py
@@ -20,3 +20,12 @@ def to_cuda(module, data):
for i in range(len(data)):
data[i] = data[i].cuda()
return module, data
+
+
+def list_to_cuda(data):
+ if isinstance(data, list):
+ for i in range(len(data)):
+ data[i] = list_to_cuda(data[i])
+ return data
+ else:
+ return data.cuda()
diff --git a/tests/test_models/test_losses/test_dice_loss.py b/tests/test_models/test_losses/test_dice_loss.py
new file mode 100644
index 00000000000..34253dae12e
--- /dev/null
+++ b/tests/test_models/test_losses/test_dice_loss.py
@@ -0,0 +1,96 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import pytest
+import torch
+
+from mmseg.models.losses import DiceLoss
+
+
+@pytest.mark.parametrize('naive_dice', [True, False])
+def test_dice_loss(naive_dice):
+ loss_class = DiceLoss
+ pred = torch.rand((1, 10, 4, 4))
+ target = torch.randint(0, 10, (1, 4, 4))
+ weight = torch.rand(1)
+ # Test loss forward
+ loss = loss_class(naive_dice=naive_dice)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with weight
+ loss = loss_class(naive_dice=naive_dice)(pred, target, weight)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with reduction_override
+ loss = loss_class(naive_dice=naive_dice)(
+ pred, target, reduction_override='mean')
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with avg_factor
+ loss = loss_class(naive_dice=naive_dice)(pred, target, avg_factor=10)
+ assert isinstance(loss, torch.Tensor)
+
+ with pytest.raises(ValueError):
+ # loss can evaluate with avg_factor only if
+ # reduction is None, 'none' or 'mean'.
+ reduction_override = 'sum'
+ loss_class(naive_dice=naive_dice)(
+ pred, target, avg_factor=10, reduction_override=reduction_override)
+
+ # Test loss forward with avg_factor and reduction
+ for reduction_override in [None, 'none', 'mean']:
+ loss_class(naive_dice=naive_dice)(
+ pred, target, avg_factor=10, reduction_override=reduction_override)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with has_acted=False and use_sigmoid=False
+ for use_sigmoid in [True, False]:
+ loss_class(
+ use_sigmoid=use_sigmoid, activate=True,
+ naive_dice=naive_dice)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with weight.ndim != loss.ndim
+ with pytest.raises(AssertionError):
+ weight = torch.rand((2, 8))
+ loss_class(naive_dice=naive_dice)(pred, target, weight)
+
+ # Test loss forward with len(weight) != len(pred)
+ with pytest.raises(AssertionError):
+ weight = torch.rand(8)
+ loss_class(naive_dice=naive_dice)(pred, target, weight)
+
+ # Test _expand_onehot_labels_dice
+ pred = torch.tensor([[[[1, 1], [1, 0]], [[0, 1], [1, 1]]]]).float()
+ target = torch.tensor([[[0, 0], [0, 1]]])
+ target_onehot = torch.tensor([[[[1, 1], [1, 0]], [[0, 0], [0, 1]]]])
+ weight = torch.rand(1)
+ loss = loss_class(naive_dice=naive_dice)(pred, target, weight)
+ loss_onehot = loss_class(naive_dice=naive_dice)(pred, target_onehot,
+ weight)
+ assert torch.equal(loss, loss_onehot)
+
+ # Test Whether Loss is 0 when pred == target, eps == 0 and naive_dice=False
+ target = torch.randint(0, 2, (1, 10, 4, 4))
+ pred = target.float()
+ target = target.sigmoid()
+ weight = torch.rand(1)
+ loss = loss_class(
+ naive_dice=False, use_sigmoid=True, eps=0)(pred, target, weight)
+ assert loss.item() == 0
+
+ # Test ignore_index when ignore_index is the only class
+ with pytest.raises(AssertionError):
+ pred = torch.ones((1, 1, 4, 4))
+ target = torch.randint(0, 1, (1, 4, 4))
+ weight = torch.rand(1)
+ loss = loss_class(
+ naive_dice=naive_dice, use_sigmoid=False, ignore_index=0,
+ eps=0)(pred, target, weight)
+
+ # Test ignore_index with naive_dice = False
+ pred = torch.tensor([[[[1, 1], [1, 0]], [[0, 1], [1, 1]]]]).float()
+ target = torch.tensor([[[[1, 1], [1, 0]], [[1, 0], [0, 1]]]]).sigmoid()
+ weight = torch.rand(1)
+ loss = loss_class(
+ naive_dice=False, use_sigmoid=True, ignore_index=1,
+ eps=0)(pred, target, weight)
+ assert loss.item() == 0
diff --git a/tests/test_models/test_losses/test_huasdorff_distance_loss.py b/tests/test_models/test_losses/test_huasdorff_distance_loss.py
new file mode 100644
index 00000000000..29c2732d3f1
--- /dev/null
+++ b/tests/test_models/test_losses/test_huasdorff_distance_loss.py
@@ -0,0 +1,29 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import pytest
+import torch
+
+from mmseg.models.losses import HuasdorffDisstanceLoss
+
+
+def test_huasdorff_distance_loss():
+ loss_class = HuasdorffDisstanceLoss
+ pred = torch.rand((10, 8, 6, 6))
+ target = torch.rand((10, 6, 6))
+ class_weight = torch.rand(8)
+
+ # Test loss forward
+ loss = loss_class()(pred, target)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with avg_factor
+ loss = loss_class()(pred, target, avg_factor=10)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with avg_factor and reduction is None, 'sum' and 'mean'
+ for reduction in [None, 'sum', 'mean']:
+ loss = loss_class()(pred, target, avg_factor=10, reduction=reduction)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with class_weight
+ with pytest.raises(AssertionError):
+ loss_class(class_weight=class_weight)(pred, target)
diff --git a/tests/test_models/test_losses/test_kldiv_loss.py b/tests/test_models/test_losses/test_kldiv_loss.py
new file mode 100644
index 00000000000..48bcc4bfd9f
--- /dev/null
+++ b/tests/test_models/test_losses/test_kldiv_loss.py
@@ -0,0 +1,40 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch
+
+from mmseg.models.losses.kldiv_loss import KLDivLoss
+
+
+def test_kldiv_loss_with_none_reduction():
+ loss_class = KLDivLoss
+ pred = torch.rand((8, 5, 5))
+ target = torch.rand((8, 5, 5))
+ reduction = 'none'
+
+ # Test loss forward
+ loss = loss_class(reduction=reduction)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+ assert loss.shape == (8, 5, 5), f'{loss.shape}'
+
+
+def test_kldiv_loss_with_mean_reduction():
+ loss_class = KLDivLoss
+ pred = torch.rand((8, 5, 5))
+ target = torch.rand((8, 5, 5))
+ reduction = 'mean'
+
+ # Test loss forward
+ loss = loss_class(reduction=reduction)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+ assert loss.shape == (8, ), f'{loss.shape}'
+
+
+def test_kldiv_loss_with_sum_reduction():
+ loss_class = KLDivLoss
+ pred = torch.rand((8, 5, 5))
+ target = torch.rand((8, 5, 5))
+ reduction = 'sum'
+
+ # Test loss forward
+ loss = loss_class(reduction=reduction)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+ assert loss.shape == (8, ), f'{loss.shape}'
diff --git a/tests/test_models/test_losses/test_silog_loss.py b/tests/test_models/test_losses/test_silog_loss.py
new file mode 100644
index 00000000000..022434bcc14
--- /dev/null
+++ b/tests/test_models/test_losses/test_silog_loss.py
@@ -0,0 +1,20 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from unittest import TestCase
+
+import torch
+
+from mmseg.models.losses import SiLogLoss
+
+
+class TestSiLogLoss(TestCase):
+
+ def test_SiLogLoss_forward(self):
+ pred = torch.tensor([[1.0, 2.0], [3.5, 4.0]], dtype=torch.float32)
+ target = torch.tensor([[0.0, 2.0], [3.0, 4.0]], dtype=torch.float32)
+ weight = torch.tensor([1.0, 0.5], dtype=torch.float32)
+
+ loss_module = SiLogLoss()
+ loss = loss_module.forward(pred, target, weight)
+
+ expected_loss = 0.02
+ self.assertAlmostEqual(loss.item(), expected_loss, places=2)
diff --git a/tests/test_models/test_segmentors/test_depth_estimator.py b/tests/test_models/test_segmentors/test_depth_estimator.py
new file mode 100644
index 00000000000..e819c9e7633
--- /dev/null
+++ b/tests/test_models/test_segmentors/test_depth_estimator.py
@@ -0,0 +1,64 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from copy import deepcopy
+from os.path import dirname, join
+from unittest import TestCase
+
+import torch
+from mmengine import Config, ConfigDict
+from mmengine.structures import PixelData
+
+import mmseg
+from mmseg.models.segmentors import DepthEstimator
+from mmseg.structures import SegDataSample
+
+
+class TestDepthEstimator(TestCase):
+
+ def setUp(self) -> None:
+ repo_dpath = dirname(dirname(mmseg.__file__))
+ config_dpath = join(repo_dpath, 'configs/_base_/models/vpd_sd.py')
+ vpd_cfg = Config.fromfile(config_dpath).stable_diffusion_cfg
+ vpd_cfg.pop('checkpoint')
+
+ backbone_cfg = dict(
+ type='VPD',
+ diffusion_cfg=vpd_cfg,
+ class_embed_path='https://download.openmmlab.com/mmsegmentation/'
+ 'v0.5/vpd/nyu_class_embeddings.pth',
+ class_embed_select=True,
+ pad_shape=64,
+ unet_cfg=dict(use_attn=False),
+ )
+
+ head_cfg = dict(
+ type='VPDDepthHead',
+ max_depth=10,
+ )
+
+ self.model = DepthEstimator(
+ backbone=backbone_cfg, decode_head=head_cfg)
+
+ inputs = torch.randn(1, 3, 64, 80)
+ data_sample = SegDataSample()
+ data_sample.gt_depth_map = PixelData(data=torch.rand(1, 64, 80))
+ data_sample.set_metainfo(dict(img_shape=(64, 80), ori_shape=(64, 80)))
+ self.data = dict(inputs=inputs, data_samples=[data_sample])
+
+ def test_slide_flip_inference(self):
+
+ self.model.test_cfg = ConfigDict(
+ dict(mode='slide_flip', crop_size=(64, 64), stride=(16, 16)))
+
+ with torch.no_grad():
+ out = self.model.predict(**deepcopy(self.data))
+
+ self.assertEqual(len(out), 1)
+ self.assertIn('pred_depth_map', out[0].keys())
+ self.assertListEqual(list(out[0].pred_depth_map.shape), [64, 80])
+
+ def test__forward(self):
+ data = deepcopy(self.data)
+ data['inputs'] = data['inputs'][:, :, :64, :64]
+ with torch.no_grad():
+ out = self.model._forward(**data)
+ self.assertListEqual(list(out.shape), [1, 1, 64, 64])
diff --git a/tests/test_models/test_segmentors/test_multimodal_encoder_decoder.py b/tests/test_models/test_segmentors/test_multimodal_encoder_decoder.py
new file mode 100644
index 00000000000..75258d89a7d
--- /dev/null
+++ b/tests/test_models/test_segmentors/test_multimodal_encoder_decoder.py
@@ -0,0 +1,24 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from mmengine import ConfigDict
+
+from mmseg.models import build_segmentor
+from tests.test_models.test_segmentors.utils import \
+ _segmentor_forward_train_test
+
+
+def test_multimodal_encoder_decoder():
+
+ cfg = ConfigDict(
+ type='MultimodalEncoderDecoder',
+ asymetric_input=False,
+ image_encoder=dict(type='ExampleBackbone', out_indices=[1, 2, 3, 4]),
+ text_encoder=dict(
+ type='ExampleTextEncoder',
+ vocabulary=['A', 'B', 'C'],
+ output_dims=3),
+ decode_head=dict(
+ type='ExampleDecodeHead', out_channels=1, num_classes=2),
+ train_cfg=None,
+ test_cfg=dict(mode='whole'))
+ segmentor = build_segmentor(cfg)
+ _segmentor_forward_train_test(segmentor)
diff --git a/tests/test_models/test_segmentors/test_seg_tta_model.py b/tests/test_models/test_segmentors/test_seg_tta_model.py
index 3c9699e8df4..1e152ed0565 100644
--- a/tests/test_models/test_segmentors/test_seg_tta_model.py
+++ b/tests/test_models/test_segmentors/test_seg_tta_model.py
@@ -1,4 +1,6 @@
# Copyright (c) OpenMMLab. All rights reserved.
+import tempfile
+
import torch
from mmengine import ConfigDict
from mmengine.model import BaseTTAModel
@@ -37,7 +39,8 @@ def test_encoder_decoder_tta():
ori_shape=(10, 10),
img_shape=(10 + i, 10 + i),
flip=(i % 2 == 0),
- flip_direction=flip_direction),
+ flip_direction=flip_direction,
+ img_path=tempfile.mktemp()),
gt_sem_seg=PixelData(data=torch.randint(0, 19, (1, 10, 10))))
])
diff --git a/tests/test_models/test_segmentors/utils.py b/tests/test_models/test_segmentors/utils.py
index 6b440df906d..ac31e2b2774 100644
--- a/tests/test_models/test_segmentors/utils.py
+++ b/tests/test_models/test_segmentors/utils.py
@@ -52,15 +52,22 @@ def _demo_mm_inputs(input_shape=(1, 3, 8, 16), num_classes=10):
@MODELS.register_module()
class ExampleBackbone(nn.Module):
- def __init__(self):
+ def __init__(self, out_indices=None):
super().__init__()
self.conv = nn.Conv2d(3, 3, 3)
+ self.out_indices = out_indices
def init_weights(self, pretrained=None):
pass
def forward(self, x):
- return [self.conv(x)]
+ if self.out_indices is None:
+ return [self.conv(x)]
+ else:
+ outs = []
+ for i in self.out_indices:
+ outs.append(self.conv(x))
+ return outs
@MODELS.register_module()
@@ -74,6 +81,18 @@ def forward(self, inputs):
return self.cls_seg(inputs[0])
+@MODELS.register_module()
+class ExampleTextEncoder(nn.Module):
+
+ def __init__(self, vocabulary=None, output_dims=None):
+ super().__init__()
+ self.vocabulary = vocabulary
+ self.output_dims = output_dims
+
+ def forward(self):
+ return torch.randn((len(self.vocabulary), self.output_dims))
+
+
@MODELS.register_module()
class ExampleCascadeDecodeHead(BaseCascadeDecodeHead):
@@ -132,3 +151,32 @@ def _segmentor_forward_train_test(segmentor):
data_batch = dict(inputs=imgs, data_samples=data_samples)
results = segmentor.forward(imgs, data_samples, mode='tensor')
assert isinstance(results, torch.Tensor)
+
+
+def _segmentor_predict(segmentor):
+ if isinstance(segmentor.decode_head, nn.ModuleList):
+ num_classes = segmentor.decode_head[-1].num_classes
+ else:
+ num_classes = segmentor.decode_head.num_classes
+ # batch_size=2 for BatchNorm
+ mm_inputs = _demo_mm_inputs(num_classes=num_classes)
+
+ # convert to cuda Tensor if applicable
+ if torch.cuda.is_available():
+ segmentor = segmentor.cuda()
+
+ # check data preprocessor
+ if not hasattr(segmentor,
+ 'data_preprocessor') or segmentor.data_preprocessor is None:
+ segmentor.data_preprocessor = SegDataPreProcessor()
+
+ mm_inputs = segmentor.data_preprocessor(mm_inputs, True)
+ imgs = mm_inputs.pop('imgs')
+ data_samples = mm_inputs.pop('data_samples')
+
+ # Test predict
+ with torch.no_grad():
+ segmentor.eval()
+ data_batch = dict(inputs=imgs, data_samples=data_samples)
+ outputs = segmentor.predict(**data_batch)
+ assert isinstance(outputs, list)
diff --git a/tests/test_visualization/test_local_visualizer.py b/tests/test_visualization/test_local_visualizer.py
index b60a9b87507..e3b2a88cfb7 100644
--- a/tests/test_visualization/test_local_visualizer.py
+++ b/tests/test_visualization/test_local_visualizer.py
@@ -155,3 +155,59 @@ def _assert_image_and_shape(self, out_file, out_shape):
assert os.path.exists(out_file)
drawn_img = cv2.imread(out_file)
assert drawn_img.shape == out_shape
+
+ def test_add_datasample_depth(self):
+ h = 10
+ w = 12
+ out_file = 'out_file'
+
+ image = np.random.randint(0, 256, size=(h, w, 3)).astype('uint8')
+
+ # test gt_depth_map
+ gt_depth_map = PixelData(data=torch.rand(1, h, w))
+
+ def test_add_datasample_forward_depth(gt_depth_map):
+ data_sample = SegDataSample()
+ data_sample.gt_depth_map = gt_depth_map
+
+ with tempfile.TemporaryDirectory() as tmp_dir:
+ seg_local_visualizer = SegLocalVisualizer(
+ vis_backends=[dict(type='LocalVisBackend')],
+ save_dir=tmp_dir)
+ seg_local_visualizer.dataset_meta = dict(
+ classes=('background', 'foreground'),
+ palette=[[120, 120, 120], [6, 230, 230]])
+
+ # test out_file
+ seg_local_visualizer.add_datasample(out_file, image,
+ data_sample)
+
+ assert os.path.exists(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'))
+ drawn_img = cv2.imread(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'))
+ assert drawn_img.shape == (h * 2, w, 3)
+
+ # test gt_instances and pred_instances
+
+ pred_depth_map = PixelData(data=torch.rand(1, h, w))
+
+ data_sample.pred_depth_map = pred_depth_map
+
+ seg_local_visualizer.add_datasample(out_file, image,
+ data_sample)
+ self._assert_image_and_shape(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'), (h * 2, w * 2, 3))
+
+ seg_local_visualizer.add_datasample(
+ out_file, image, data_sample, draw_gt=False)
+ self._assert_image_and_shape(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'), (h * 2, w, 3))
+
+ if torch.cuda.is_available():
+ test_add_datasample_forward_depth(gt_depth_map.cuda())
+ test_add_datasample_forward_depth(gt_depth_map)
diff --git a/tools/analysis_tools/confusion_matrix.py b/tools/analysis_tools/confusion_matrix.py
index 9a87bc14c9b..39756cdfdd2 100644
--- a/tools/analysis_tools/confusion_matrix.py
+++ b/tools/analysis_tools/confusion_matrix.py
@@ -5,10 +5,14 @@
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.ticker import MultipleLocator
-from mmengine import Config, DictAction
-from mmengine.utils import ProgressBar, load
+from mmengine.config import Config, DictAction
+from mmengine.registry import init_default_scope
+from mmengine.utils import mkdir_or_exist, progressbar
+from PIL import Image
-from mmseg.datasets import build_dataset
+from mmseg.registry import DATASETS
+
+init_default_scope('mmseg')
def parse_args():
@@ -16,7 +20,7 @@ def parse_args():
description='Generate confusion matrix from segmentation results')
parser.add_argument('config', help='test config file path')
parser.add_argument(
- 'prediction_path', help='prediction path where test .pkl result')
+ 'prediction_path', help='prediction path where test folder result')
parser.add_argument(
'save_dir', help='directory where confusion matrix will be saved')
parser.add_argument(
@@ -50,15 +54,23 @@ def calculate_confusion_matrix(dataset, results):
dataset (Dataset): Test or val dataset.
results (list[ndarray]): A list of segmentation results in each image.
"""
- n = len(dataset.CLASSES)
+ n = len(dataset.METAINFO['classes'])
confusion_matrix = np.zeros(shape=[n, n])
assert len(dataset) == len(results)
- prog_bar = ProgressBar(len(results))
+ ignore_index = dataset.ignore_index
+ reduce_zero_label = dataset.reduce_zero_label
+ prog_bar = progressbar.ProgressBar(len(results))
for idx, per_img_res in enumerate(results):
res_segm = per_img_res
- gt_segm = dataset.get_gt_seg_map_by_idx(idx)
+ gt_segm = dataset[idx]['data_samples'] \
+ .gt_sem_seg.data.squeeze().numpy().astype(np.uint8)
+ gt_segm, res_segm = gt_segm.flatten(), res_segm.flatten()
+ if reduce_zero_label:
+ gt_segm = gt_segm - 1
+ to_ignore = gt_segm == ignore_index
+
+ gt_segm, res_segm = gt_segm[~to_ignore], res_segm[~to_ignore]
inds = n * gt_segm + res_segm
- inds = inds.flatten()
mat = np.bincount(inds, minlength=n**2).reshape(n, n)
confusion_matrix += mat
prog_bar.update()
@@ -70,7 +82,7 @@ def plot_confusion_matrix(confusion_matrix,
save_dir=None,
show=True,
title='Normalized Confusion Matrix',
- color_theme='winter'):
+ color_theme='OrRd'):
"""Draw confusion matrix with matplotlib.
Args:
@@ -89,14 +101,15 @@ def plot_confusion_matrix(confusion_matrix,
num_classes = len(labels)
fig, ax = plt.subplots(
- figsize=(2 * num_classes, 2 * num_classes * 0.8), dpi=180)
+ figsize=(2 * num_classes, 2 * num_classes * 0.8), dpi=300)
cmap = plt.get_cmap(color_theme)
im = ax.imshow(confusion_matrix, cmap=cmap)
- plt.colorbar(mappable=im, ax=ax)
+ colorbar = plt.colorbar(mappable=im, ax=ax)
+ colorbar.ax.tick_params(labelsize=20) # 设置 colorbar 标签的字体大小
- title_font = {'weight': 'bold', 'size': 12}
+ title_font = {'weight': 'bold', 'size': 20}
ax.set_title(title, fontdict=title_font)
- label_font = {'size': 10}
+ label_font = {'size': 40}
plt.ylabel('Ground Truth Label', fontdict=label_font)
plt.xlabel('Prediction Label', fontdict=label_font)
@@ -116,8 +129,8 @@ def plot_confusion_matrix(confusion_matrix,
# draw label
ax.set_xticks(np.arange(num_classes))
ax.set_yticks(np.arange(num_classes))
- ax.set_xticklabels(labels)
- ax.set_yticklabels(labels)
+ ax.set_xticklabels(labels, fontsize=20)
+ ax.set_yticklabels(labels, fontsize=20)
ax.tick_params(
axis='x', bottom=False, top=True, labelbottom=False, labeltop=True)
@@ -135,13 +148,14 @@ def plot_confusion_matrix(confusion_matrix,
) if not np.isnan(confusion_matrix[i, j]) else -1),
ha='center',
va='center',
- color='w',
- size=7)
+ color='k',
+ size=20)
ax.set_ylim(len(confusion_matrix) - 0.5, -0.5) # matplotlib>3.1.1
fig.tight_layout()
if save_dir is not None:
+ mkdir_or_exist(save_dir)
plt.savefig(
os.path.join(save_dir, 'confusion_matrix.png'), format='png')
if show:
@@ -155,7 +169,12 @@ def main():
if args.cfg_options is not None:
cfg.merge_from_dict(args.cfg_options)
- results = load(args.prediction_path)
+ results = []
+ for img in sorted(os.listdir(args.prediction_path)):
+ img = os.path.join(args.prediction_path, img)
+ image = Image.open(img)
+ image = np.copy(image)
+ results.append(image)
assert isinstance(results, list)
if isinstance(results[0], np.ndarray):
@@ -163,17 +182,11 @@ def main():
else:
raise TypeError('invalid type of prediction results')
- if isinstance(cfg.data.test, dict):
- cfg.data.test.test_mode = True
- elif isinstance(cfg.data.test, list):
- for ds_cfg in cfg.data.test:
- ds_cfg.test_mode = True
-
- dataset = build_dataset(cfg.data.test)
+ dataset = DATASETS.build(cfg.test_dataloader.dataset)
confusion_matrix = calculate_confusion_matrix(dataset, results)
plot_confusion_matrix(
confusion_matrix,
- dataset.CLASSES,
+ dataset.METAINFO['classes'],
save_dir=args.save_dir,
show=args.show,
title=args.title,
diff --git a/tools/analysis_tools/visualization_cam.py b/tools/analysis_tools/visualization_cam.py
new file mode 100644
index 00000000000..00cdb3e04ab
--- /dev/null
+++ b/tools/analysis_tools/visualization_cam.py
@@ -0,0 +1,127 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+"""Use the pytorch-grad-cam tool to visualize Class Activation Maps (CAM).
+
+requirement: pip install grad-cam
+"""
+
+from argparse import ArgumentParser
+
+import numpy as np
+import torch
+import torch.nn.functional as F
+from mmengine import Config
+from mmengine.model import revert_sync_batchnorm
+from PIL import Image
+from pytorch_grad_cam import GradCAM
+from pytorch_grad_cam.utils.image import preprocess_image, show_cam_on_image
+
+from mmseg.apis import inference_model, init_model, show_result_pyplot
+from mmseg.utils import register_all_modules
+
+
+class SemanticSegmentationTarget:
+ """wrap the model.
+
+ requirement: pip install grad-cam
+
+ Args:
+ category (int): Visualization class.
+ mask (ndarray): Mask of class.
+ size (tuple): Image size.
+ """
+
+ def __init__(self, category, mask, size):
+ self.category = category
+ self.mask = torch.from_numpy(mask)
+ self.size = size
+ if torch.cuda.is_available():
+ self.mask = self.mask.cuda()
+
+ def __call__(self, model_output):
+ model_output = torch.unsqueeze(model_output, dim=0)
+ model_output = F.interpolate(
+ model_output, size=self.size, mode='bilinear')
+ model_output = torch.squeeze(model_output, dim=0)
+
+ return (model_output[self.category, :, :] * self.mask).sum()
+
+
+def main():
+ parser = ArgumentParser()
+ parser.add_argument('img', help='Image file')
+ parser.add_argument('config', help='Config file')
+ parser.add_argument('checkpoint', help='Checkpoint file')
+ parser.add_argument(
+ '--out-file',
+ default='prediction.png',
+ help='Path to output prediction file')
+ parser.add_argument(
+ '--cam-file', default='vis_cam.png', help='Path to output cam file')
+ parser.add_argument(
+ '--target-layers',
+ default='backbone.layer4[2]',
+ help='Target layers to visualize CAM')
+ parser.add_argument(
+ '--category-index', default='7', help='Category to visualize CAM')
+ parser.add_argument(
+ '--device', default='cuda:0', help='Device used for inference')
+ args = parser.parse_args()
+
+ # build the model from a config file and a checkpoint file
+ register_all_modules()
+ model = init_model(args.config, args.checkpoint, device=args.device)
+ if args.device == 'cpu':
+ model = revert_sync_batchnorm(model)
+
+ # test a single image
+ result = inference_model(model, args.img)
+
+ # show the results
+ show_result_pyplot(
+ model,
+ args.img,
+ result,
+ draw_gt=False,
+ show=False if args.out_file is not None else True,
+ out_file=args.out_file)
+
+ # result data conversion
+ prediction_data = result.pred_sem_seg.data
+ pre_np_data = prediction_data.cpu().numpy().squeeze(0)
+
+ target_layers = args.target_layers
+ target_layers = [eval(f'model.{target_layers}')]
+
+ category = int(args.category_index)
+ mask_float = np.float32(pre_np_data == category)
+
+ # data processing
+ image = np.array(Image.open(args.img).convert('RGB'))
+ height, width = image.shape[0], image.shape[1]
+ rgb_img = np.float32(image) / 255
+ config = Config.fromfile(args.config)
+ image_mean = config.data_preprocessor['mean']
+ image_std = config.data_preprocessor['std']
+ input_tensor = preprocess_image(
+ rgb_img,
+ mean=[x / 255 for x in image_mean],
+ std=[x / 255 for x in image_std])
+
+ # Grad CAM(Class Activation Maps)
+ # Can also be LayerCAM, XGradCAM, GradCAMPlusPlus, EigenCAM, EigenGradCAM
+ targets = [
+ SemanticSegmentationTarget(category, mask_float, (height, width))
+ ]
+ with GradCAM(
+ model=model,
+ target_layers=target_layers,
+ use_cuda=torch.cuda.is_available()) as cam:
+ grayscale_cam = cam(input_tensor=input_tensor, targets=targets)[0, :]
+ cam_image = show_cam_on_image(rgb_img, grayscale_cam, use_rgb=True)
+
+ # save cam file
+ Image.fromarray(cam_image).save(args.cam_file)
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/dataset_converters/isaid.py b/tools/dataset_converters/isaid.py
index 1da264d975e..1d5ccd9c776 100644
--- a/tools/dataset_converters/isaid.py
+++ b/tools/dataset_converters/isaid.py
@@ -91,7 +91,7 @@ def slide_crop_image(src_path, out_dir, mode, patch_H, patch_W, overlap):
x_end) + '.png'
# print(image)
save_path_image = osp.join(out_dir, 'img_dir', mode, str(image))
- img_patch.save(save_path_image)
+ img_patch.save(save_path_image, format='BMP')
def slide_crop_label(src_path, out_dir, mode, patch_H, patch_W, overlap):
diff --git a/tools/dataset_converters/levircd.py b/tools/dataset_converters/levircd.py
new file mode 100644
index 00000000000..8717f3e856b
--- /dev/null
+++ b/tools/dataset_converters/levircd.py
@@ -0,0 +1,99 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import glob
+import math
+import os
+import os.path as osp
+
+import mmcv
+import numpy as np
+from mmengine.utils import ProgressBar
+
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='Convert levir-cd dataset to mmsegmentation format')
+ parser.add_argument('--dataset_path', help='potsdam folder path')
+ parser.add_argument('-o', '--out_dir', help='output path')
+ parser.add_argument(
+ '--clip_size',
+ type=int,
+ help='clipped size of image after preparation',
+ default=256)
+ parser.add_argument(
+ '--stride_size',
+ type=int,
+ help='stride of clipping original images',
+ default=256)
+ args = parser.parse_args()
+ return args
+
+
+def main():
+ args = parse_args()
+ input_folder = args.dataset_path
+ png_files = glob.glob(
+ os.path.join(input_folder, '**/*.png'), recursive=True)
+ output_folder = args.out_dir
+ prog_bar = ProgressBar(len(png_files))
+ for png_file in png_files:
+ new_path = os.path.join(
+ output_folder,
+ os.path.relpath(os.path.dirname(png_file), input_folder))
+ os.makedirs(os.path.dirname(new_path), exist_ok=True)
+ label = False
+ if 'label' in png_file:
+ label = True
+ clip_big_image(png_file, new_path, args, label)
+ prog_bar.update()
+
+
+def clip_big_image(image_path, clip_save_dir, args, to_label=False):
+ image = mmcv.imread(image_path)
+
+ h, w, c = image.shape
+ clip_size = args.clip_size
+ stride_size = args.stride_size
+
+ num_rows = math.ceil((h - clip_size) / stride_size) if math.ceil(
+ (h - clip_size) /
+ stride_size) * stride_size + clip_size >= h else math.ceil(
+ (h - clip_size) / stride_size) + 1
+ num_cols = math.ceil((w - clip_size) / stride_size) if math.ceil(
+ (w - clip_size) /
+ stride_size) * stride_size + clip_size >= w else math.ceil(
+ (w - clip_size) / stride_size) + 1
+
+ x, y = np.meshgrid(np.arange(num_cols + 1), np.arange(num_rows + 1))
+ xmin = x * clip_size
+ ymin = y * clip_size
+
+ xmin = xmin.ravel()
+ ymin = ymin.ravel()
+ xmin_offset = np.where(xmin + clip_size > w, w - xmin - clip_size,
+ np.zeros_like(xmin))
+ ymin_offset = np.where(ymin + clip_size > h, h - ymin - clip_size,
+ np.zeros_like(ymin))
+ boxes = np.stack([
+ xmin + xmin_offset, ymin + ymin_offset,
+ np.minimum(xmin + clip_size, w),
+ np.minimum(ymin + clip_size, h)
+ ],
+ axis=1)
+
+ if to_label:
+ image[image == 255] = 1
+ image = image[:, :, 0]
+ for box in boxes:
+ start_x, start_y, end_x, end_y = box
+ clipped_image = image[start_y:end_y, start_x:end_x] \
+ if to_label else image[start_y:end_y, start_x:end_x, :]
+ idx = osp.basename(image_path).split('.')[0]
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(clip_save_dir,
+ f'{idx}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/dataset_converters/nyu.py b/tools/dataset_converters/nyu.py
new file mode 100644
index 00000000000..49e09e7af68
--- /dev/null
+++ b/tools/dataset_converters/nyu.py
@@ -0,0 +1,89 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import os.path as osp
+import shutil
+import tempfile
+import zipfile
+
+from mmengine.utils import mkdir_or_exist
+
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='Convert NYU Depth dataset to mmsegmentation format')
+ parser.add_argument('raw_data', help='the path of raw data')
+ parser.add_argument(
+ '-o', '--out_dir', help='output path', default='./data/nyu')
+ args = parser.parse_args()
+ return args
+
+
+def reorganize(raw_data_dir: str, out_dir: str):
+ """Reorganize NYU Depth dataset files into the required directory
+ structure.
+
+ Args:
+ raw_data_dir (str): Path to the raw data directory.
+ out_dir (str): Output directory for the organized dataset.
+ """
+
+ def move_data(data_list, dst_prefix, fname_func):
+ """Move data files from source to destination directory.
+
+ Args:
+ data_list (list): List of data file paths.
+ dst_prefix (str): Prefix to be added to destination paths.
+ fname_func (callable): Function to process file names
+ """
+ for data_item in data_list:
+ data_item = data_item.strip().strip('/')
+ new_item = fname_func(data_item)
+ shutil.move(
+ osp.join(raw_data_dir, data_item),
+ osp.join(out_dir, dst_prefix, new_item))
+
+ def process_phase(phase):
+ """Process a dataset phase (e.g., 'train' or 'test')."""
+ with open(osp.join(raw_data_dir, f'nyu_{phase}.txt')) as f:
+ data = filter(lambda x: len(x.strip()) > 0, f.readlines())
+ data = map(lambda x: x.split()[:2], data)
+ images, annos = zip(*data)
+
+ move_data(images, f'images/{phase}',
+ lambda x: x.replace('/rgb', ''))
+ move_data(annos, f'annotations/{phase}',
+ lambda x: x.replace('/sync_depth', ''))
+
+ process_phase('train')
+ process_phase('test')
+
+
+def main():
+ args = parse_args()
+
+ print('Making directories...')
+ mkdir_or_exist(args.out_dir)
+ for subdir in [
+ 'images/train', 'images/test', 'annotations/train',
+ 'annotations/test'
+ ]:
+ mkdir_or_exist(osp.join(args.out_dir, subdir))
+
+ print('Generating images and annotations...')
+
+ if args.raw_data.endswith('.zip'):
+ with tempfile.TemporaryDirectory() as tmp_dir:
+ zip_file = zipfile.ZipFile(args.raw_data)
+ zip_file.extractall(tmp_dir)
+ reorganize(osp.join(tmp_dir, 'nyu'), args.out_dir)
+ else:
+ assert osp.isdir(
+ args.raw_data
+ ), 'the argument --raw-data should be either a zip file or directory.'
+ reorganize(args.raw_data, args.out_dir)
+
+ print('Done!')
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/misc/publish_model.py b/tools/misc/publish_model.py
index c1bbc9ac1ad..e035ad90e85 100644
--- a/tools/misc/publish_model.py
+++ b/tools/misc/publish_model.py
@@ -1,9 +1,12 @@
# Copyright (c) OpenMMLab. All rights reserved.
import argparse
import subprocess
+from hashlib import sha256
import torch
+BLOCK_SIZE = 128 * 1024
+
def parse_args():
parser = argparse.ArgumentParser(
@@ -14,6 +17,17 @@ def parse_args():
return args
+def sha256sum(filename: str) -> str:
+ """Compute SHA256 message digest from a file."""
+ hash_func = sha256()
+ byte_array = bytearray(BLOCK_SIZE)
+ memory_view = memoryview(byte_array)
+ with open(filename, 'rb', buffering=0) as file:
+ for block in iter(lambda: file.readinto(memory_view), 0):
+ hash_func.update(memory_view[:block])
+ return hash_func.hexdigest()
+
+
def process_checkpoint(in_file, out_file):
checkpoint = torch.load(in_file, map_location='cpu')
# remove optimizer for smaller file size
@@ -22,7 +36,7 @@ def process_checkpoint(in_file, out_file):
# if it is necessary to remove some sensitive data in checkpoint['meta'],
# add the code here.
torch.save(checkpoint, out_file)
- sha = subprocess.check_output(['sha256sum', out_file]).decode()
+ sha = sha256sum(in_file)
final_file = out_file.rstrip('.pth') + f'-{sha[:8]}.pth'
subprocess.Popen(['mv', out_file, final_file])
diff --git a/tools/model_converters/clip2mmseg.py b/tools/model_converters/clip2mmseg.py
new file mode 100644
index 00000000000..9a97e4b04ab
--- /dev/null
+++ b/tools/model_converters/clip2mmseg.py
@@ -0,0 +1,163 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import os.path as osp
+from collections import OrderedDict
+
+import mmengine
+import torch
+from mmengine.runner import CheckpointLoader
+
+
+def convert_vitlayer(paras):
+ new_para_name = ''
+ if paras[0] == 'ln_1':
+ new_para_name = '.'.join(['ln1'] + paras[1:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(['attn.attn'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['ln2'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffn.layers.0.0'] + paras[-1:])
+ else:
+ new_para_name = '.'.join(['ffn.layers.1'] + paras[-1:])
+ else:
+ print(f'Wrong for {paras}')
+ return new_para_name
+
+
+def convert_translayer(paras):
+ new_para_name = ''
+ if paras[0] == 'attn':
+ new_para_name = '.'.join(['attentions.0.attn'] + paras[1:])
+ elif paras[0] == 'ln_1':
+ new_para_name = '.'.join(['norms.0'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['norms.1'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffns.0.layers.0.0'] + paras[2:])
+ elif paras[1] == 'c_proj':
+ new_para_name = '.'.join(['ffns.0.layers.1'] + paras[2:])
+ else:
+ print(f'Wrong for {paras}')
+ else:
+ print(f'Wrong for {paras}')
+ return new_para_name
+
+
+def convert_key_name(ckpt, visual_split):
+ new_ckpt = OrderedDict()
+ for k, v in ckpt.items():
+ key_list = k.split('.')
+ if key_list[0] == 'visual':
+ new_transform_name = 'image_encoder'
+ if key_list[1] == 'class_embedding':
+ new_name = '.'.join([new_transform_name, 'cls_token'])
+ elif key_list[1] == 'positional_embedding':
+ new_name = '.'.join([new_transform_name, 'pos_embed'])
+ elif key_list[1] == 'conv1':
+ new_name = '.'.join([
+ new_transform_name, 'patch_embed.projection', key_list[2]
+ ])
+ elif key_list[1] == 'ln_pre':
+ new_name = '.'.join(
+ [new_transform_name, key_list[1], key_list[2]])
+ elif key_list[1] == 'transformer':
+ new_layer_name = 'layers'
+ layer_index = key_list[3]
+ paras = key_list[4:]
+ if int(layer_index) < visual_split:
+ new_para_name = convert_vitlayer(paras)
+ new_name = '.'.join([
+ new_transform_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ else:
+ new_para_name = convert_translayer(paras)
+ new_transform_name = 'decode_head.rec_with_attnbias'
+ new_layer_name = 'layers'
+ layer_index = str(int(layer_index) - visual_split)
+ new_name = '.'.join([
+ new_transform_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[1] == 'proj':
+ new_name = 'decode_head.rec_with_attnbias.proj.weight'
+ elif key_list[1] == 'ln_post':
+ new_name = k.replace('visual', 'decode_head.rec_with_attnbias')
+ else:
+ print(f'pop parameter: {k}')
+ continue
+ else:
+ text_encoder_name = 'text_encoder'
+ if key_list[0] == 'transformer':
+ layer_name = 'transformer'
+ layer_index = key_list[2]
+ paras = key_list[3:]
+ new_para_name = convert_translayer(paras)
+ new_name = '.'.join([
+ text_encoder_name, layer_name, layer_index, new_para_name
+ ])
+ elif key_list[0] in [
+ 'positional_embedding', 'text_projection', 'bg_embed',
+ 'attn_mask', 'logit_scale', 'token_embedding', 'ln_final'
+ ]:
+ new_name = 'text_encoder.' + k
+ else:
+ print(f'pop parameter: {k}')
+ continue
+ new_ckpt[new_name] = v
+
+ return new_ckpt
+
+
+def convert_tensor(ckpt):
+ cls_token = ckpt['image_encoder.cls_token']
+ new_cls_token = cls_token.unsqueeze(0).unsqueeze(0)
+ ckpt['image_encoder.cls_token'] = new_cls_token
+ pos_embed = ckpt['image_encoder.pos_embed']
+ new_pos_embed = pos_embed.unsqueeze(0)
+ ckpt['image_encoder.pos_embed'] = new_pos_embed
+ proj_weight = ckpt['decode_head.rec_with_attnbias.proj.weight']
+ new_proj_weight = proj_weight.transpose(1, 0)
+ ckpt['decode_head.rec_with_attnbias.proj.weight'] = new_proj_weight
+ return ckpt
+
+
+def main():
+ parser = argparse.ArgumentParser(
+ description='Convert keys in timm pretrained vit models to '
+ 'MMSegmentation style.')
+ parser.add_argument('src', help='src model path or url')
+ # The dst path must be a full path of the new checkpoint.
+ parser.add_argument('dst', help='save path')
+ args = parser.parse_args()
+
+ if any([s in args.src for s in ['B-16', 'b16', 'base_patch16']]):
+ visual_split = 9
+ elif any([s in args.src for s in ['L-14', 'l14', 'large_patch14']]):
+ visual_split = 18
+ else:
+ print('Make sure the clip model is ViT-B/16 or ViT-L/14!')
+ visual_split = -1
+ checkpoint = CheckpointLoader.load_checkpoint(args.src, map_location='cpu')
+ if isinstance(checkpoint, torch.jit.RecursiveScriptModule):
+ state_dict = checkpoint.state_dict()
+ else:
+ if 'state_dict' in checkpoint:
+ # timm checkpoint
+ state_dict = checkpoint['state_dict']
+ elif 'model' in checkpoint:
+ # deit checkpoint
+ state_dict = checkpoint['model']
+ else:
+ state_dict = checkpoint
+ weight = convert_key_name(state_dict, visual_split)
+ weight = convert_tensor(weight)
+ mmengine.mkdir_or_exist(osp.dirname(args.dst))
+ torch.save(weight, args.dst)
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/model_converters/san2mmseg.py b/tools/model_converters/san2mmseg.py
new file mode 100644
index 00000000000..301a46608e0
--- /dev/null
+++ b/tools/model_converters/san2mmseg.py
@@ -0,0 +1,220 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import os.path as osp
+from collections import OrderedDict
+
+import mmengine
+import torch
+from mmengine.runner import CheckpointLoader
+
+
+def convert_key_name(ckpt):
+ new_ckpt = OrderedDict()
+
+ for k, v in ckpt.items():
+ key_list = k.split('.')
+ if key_list[0] == 'clip_visual_extractor':
+ new_transform_name = 'image_encoder'
+ if key_list[1] == 'class_embedding':
+ new_name = '.'.join([new_transform_name, 'cls_token'])
+ elif key_list[1] == 'positional_embedding':
+ new_name = '.'.join([new_transform_name, 'pos_embed'])
+ elif key_list[1] == 'conv1':
+ new_name = '.'.join([
+ new_transform_name, 'patch_embed.projection', key_list[2]
+ ])
+ elif key_list[1] == 'ln_pre':
+ new_name = '.'.join(
+ [new_transform_name, key_list[1], key_list[2]])
+ elif key_list[1] == 'resblocks':
+ new_layer_name = 'layers'
+ layer_index = key_list[2]
+ paras = key_list[3:]
+ if paras[0] == 'ln_1':
+ new_para_name = '.'.join(['ln1'] + key_list[4:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(['attn.attn'] + key_list[4:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['ln2'] + key_list[4:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffn.layers.0.0'] +
+ key_list[-1:])
+ else:
+ new_para_name = '.'.join(['ffn.layers.1'] +
+ key_list[-1:])
+ new_name = '.'.join([
+ new_transform_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[0] == 'side_adapter_network':
+ decode_head_name = 'decode_head'
+ module_name = 'side_adapter_network'
+ if key_list[1] == 'vit_model':
+ if key_list[2] == 'blocks':
+ layer_name = 'encode_layers'
+ layer_index = key_list[3]
+ paras = key_list[4:]
+ if paras[0] == 'norm1':
+ new_para_name = '.'.join(['ln1'] + key_list[5:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(key_list[4:])
+ new_para_name = new_para_name.replace(
+ 'attn.qkv.', 'attn.attn.in_proj_')
+ new_para_name = new_para_name.replace(
+ 'attn.proj', 'attn.attn.out_proj')
+ elif paras[0] == 'norm2':
+ new_para_name = '.'.join(['ln2'] + key_list[5:])
+ elif paras[0] == 'mlp':
+ new_para_name = '.'.join(['ffn'] + key_list[5:])
+ new_para_name = new_para_name.replace(
+ 'fc1', 'layers.0.0')
+ new_para_name = new_para_name.replace(
+ 'fc2', 'layers.1')
+ else:
+ print(f'Wrong for {k}')
+ new_name = '.'.join([
+ decode_head_name, module_name, layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[2] == 'pos_embed':
+ new_name = '.'.join(
+ [decode_head_name, module_name, 'pos_embed'])
+ elif key_list[2] == 'patch_embed':
+ new_name = '.'.join([
+ decode_head_name, module_name, 'patch_embed',
+ 'projection', key_list[4]
+ ])
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[1] == 'query_embed' or key_list[
+ 1] == 'query_pos_embed':
+ new_name = '.'.join(
+ [decode_head_name, module_name, key_list[1]])
+ elif key_list[1] == 'fusion_layers':
+ layer_name = 'conv_clips'
+ layer_index = key_list[2][-1]
+ paras = '.'.join(key_list[3:])
+ new_para_name = paras.replace('input_proj.0', '0')
+ new_para_name = new_para_name.replace('input_proj.1', '1.conv')
+ new_name = '.'.join([
+ decode_head_name, module_name, layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[1] == 'mask_decoder':
+ new_name = 'decode_head.' + k
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[0] == 'clip_rec_head':
+ module_name = 'rec_with_attnbias'
+ if key_list[1] == 'proj':
+ new_name = '.'.join(
+ [decode_head_name, module_name, 'proj.weight'])
+ elif key_list[1] == 'ln_post':
+ new_name = '.'.join(
+ [decode_head_name, module_name, 'ln_post', key_list[2]])
+ elif key_list[1] == 'resblocks':
+ new_layer_name = 'layers'
+ layer_index = key_list[2]
+ paras = key_list[3:]
+ if paras[0] == 'ln_1':
+ new_para_name = '.'.join(['norms.0'] + paras[1:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(['attentions.0.attn'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['norms.1'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffns.0.layers.0.0'] +
+ paras[2:])
+ elif paras[1] == 'c_proj':
+ new_para_name = '.'.join(['ffns.0.layers.1'] +
+ paras[2:])
+ else:
+ print(f'Wrong for {k}')
+ new_name = '.'.join([
+ decode_head_name, module_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[0] == 'ov_classifier':
+ text_encoder_name = 'text_encoder'
+ if key_list[1] == 'transformer':
+ layer_name = 'transformer'
+ layer_index = key_list[3]
+ paras = key_list[4:]
+ if paras[0] == 'attn':
+ new_para_name = '.'.join(['attentions.0.attn'] + paras[1:])
+ elif paras[0] == 'ln_1':
+ new_para_name = '.'.join(['norms.0'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['norms.1'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffns.0.layers.0.0'] +
+ paras[2:])
+ elif paras[1] == 'c_proj':
+ new_para_name = '.'.join(['ffns.0.layers.1'] +
+ paras[2:])
+ else:
+ print(f'Wrong for {k}')
+ else:
+ print(f'Wrong for {k}')
+ new_name = '.'.join([
+ text_encoder_name, layer_name, layer_index, new_para_name
+ ])
+ elif key_list[1] in [
+ 'positional_embedding', 'text_projection', 'bg_embed',
+ 'attn_mask', 'logit_scale', 'token_embedding', 'ln_final'
+ ]:
+ new_name = k.replace('ov_classifier', 'text_encoder')
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[0] == 'criterion':
+ new_name = k
+ else:
+ print(f'Wrong for {k}')
+ new_ckpt[new_name] = v
+ return new_ckpt
+
+
+def convert_tensor(ckpt):
+ cls_token = ckpt['image_encoder.cls_token']
+ new_cls_token = cls_token.unsqueeze(0).unsqueeze(0)
+ ckpt['image_encoder.cls_token'] = new_cls_token
+ pos_embed = ckpt['image_encoder.pos_embed']
+ new_pos_embed = pos_embed.unsqueeze(0)
+ ckpt['image_encoder.pos_embed'] = new_pos_embed
+ proj_weight = ckpt['decode_head.rec_with_attnbias.proj.weight']
+ new_proj_weight = proj_weight.transpose(1, 0)
+ ckpt['decode_head.rec_with_attnbias.proj.weight'] = new_proj_weight
+ return ckpt
+
+
+def main():
+ parser = argparse.ArgumentParser(
+ description='Convert keys in timm pretrained vit models to '
+ 'MMSegmentation style.')
+ parser.add_argument('src', help='src model path or url')
+ # The dst path must be a full path of the new checkpoint.
+ parser.add_argument('dst', help='save path')
+ args = parser.parse_args()
+
+ checkpoint = CheckpointLoader.load_checkpoint(args.src, map_location='cpu')
+ if 'state_dict' in checkpoint:
+ # timm checkpoint
+ state_dict = checkpoint['state_dict']
+ elif 'model' in checkpoint:
+ # deit checkpoint
+ state_dict = checkpoint['model']
+ else:
+ state_dict = checkpoint
+ weight = convert_key_name(state_dict)
+ weight = convert_tensor(weight)
+ mmengine.mkdir_or_exist(osp.dirname(args.dst))
+ torch.save(weight, args.dst)
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/test.py b/tools/test.py
index 64da2bc2616..0d7f39b3a8b 100644
--- a/tools/test.py
+++ b/tools/test.py
@@ -47,7 +47,10 @@ def parse_args():
help='job launcher')
parser.add_argument(
'--tta', action='store_true', help='Test time augmentation')
- parser.add_argument('--local_rank', type=int, default=0)
+ # When using PyTorch version >= 2.0.0, the `torch.distributed.launch`
+ # will pass the `--local-rank` parameter to `tools/train.py` instead
+ # of `--local_rank`.
+ parser.add_argument('--local_rank', '--local-rank', type=int, default=0)
args = parser.parse_args()
if 'LOCAL_RANK' not in os.environ:
os.environ['LOCAL_RANK'] = str(args.local_rank)
@@ -65,8 +68,8 @@ def trigger_visualization_hook(cfg, args):
visualization_hook['show'] = True
visualization_hook['wait_time'] = args.wait_time
if args.show_dir:
- visulizer = cfg.visualizer
- visulizer['save_dir'] = args.show_dir
+ visualizer = cfg.visualizer
+ visualizer['save_dir'] = args.show_dir
else:
raise RuntimeError(
'VisualizationHook must be included in default_hooks.'
diff --git a/tools/train.py b/tools/train.py
index 17213066648..10fdaa1874b 100644
--- a/tools/train.py
+++ b/tools/train.py
@@ -40,7 +40,10 @@ def parse_args():
choices=['none', 'pytorch', 'slurm', 'mpi'],
default='none',
help='job launcher')
- parser.add_argument('--local_rank', type=int, default=0)
+ # When using PyTorch version >= 2.0.0, the `torch.distributed.launch`
+ # will pass the `--local-rank` parameter to `tools/train.py` instead
+ # of `--local_rank`.
+ parser.add_argument('--local_rank', '--local-rank', type=int, default=0)
args = parser.parse_args()
if 'LOCAL_RANK' not in os.environ:
os.environ['LOCAL_RANK'] = str(args.local_rank)
Performance
+
+### ADE20K
+
+| Model | Backbone | Training Iters | Batchsize | Train Resolution | mIoU(%) | latency(ms)\* | params(M) | config | Links |
+| ----------------- | ----------------- | -------------- | --------- | ---------------- | ------- | ------------- | --------- | ------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| PP-MobileSeg-Base | StrideFormer-Base | 80000 | 32 | 512x512 | 41.57% | 265.5 | 5.62 | [config](https://github.com/Yang-Changhui/mmsegmentation/tree/add_ppmobileseg/projects/pp_mobileseg/configs/pp_mobileseg) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_2xb16_3rdparty-base_512x512-ade20k-f12b44f3.pth)\|[log](https://bj.bcebos.com/paddleseg/dygraph/ade20k/pp_mobileseg_base/train.log) |
+| PP-MobileSeg-Tiny | StrideFormer-Tiny | 80000 | 32 | 512x512 | 36.39% | 215.3 | 1.61 | [config](https://github.com/Yang-Changhui/mmsegmentation/tree/add_ppmobileseg/projects/pp_mobileseg/configs/pp_mobileseg) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_2xb16_3rdparty-tiny_512x512-ade20k-a351ebf5.pth)\|[log](https://bj.bcebos.com/paddleseg/dygraph/ade20k/pp_mobileseg_tiny/train.log) |
+
+## Usage
+
+Same as other models in MMsegmentation, you can run the following command to test the model at ${MMSEG_ROOT}:
+
+```shell
+./tools/dist_test.sh projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py checkpoints/pp_mobileseg_mobilenetv3_2xb16_3rdparty-base_512x512-ade20k-f12b44f3.pth 8
+```
+
+## Inference with ONNXRuntime
+
+### Prerequisites
+
+**1. Install onnxruntime inference engine.**
+
+Choose one of the following ways to install onnxruntime.
+
+- CPU version
+
+```shell
+pip install onnxruntime==1.15.1
+wget https://github.com/microsoft/onnxruntime/releases/download/v1.15.1/onnxruntime-linux-x64-1.15.1.tgz
+tar -zxvf onnxruntime-linux-x64-1.15.1.tgz
+export ONNXRUNTIME_DIR=$(pwd)/onnxruntime-linux-x64-1.15.1
+export LD_LIBRARY_PATH=$ONNXRUNTIME_DIR/lib:$LD_LIBRARY_PATH
+```
+
+**2. Convert model to onnx file**
+
+- Install `mim` and `mmdeploy`.
+
+```shell
+pip install openmim
+mim install mmdeploy
+git clone https://github.com/open-mmlab/mmdeploy.git
+```
+
+- Download pp_mobileseg model.
+
+```shell
+wget https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_2xb16_3rdparty-tiny_512x512-ade20k-a351ebf5.pth
+```
+
+- Convert model to onnx files.
+
+```shell
+python mmdeploy/tools/deploy.py mmdeploy/configs/mmseg/segmentation_onnxruntime_dynamic.py \
+ configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py \
+ pp_mobileseg_mobilenetv3_2xb16_3rdparty-tiny_512x512-ade20k-a351ebf5.pth \
+ ../../demo/demo.png \
+ --work-dir mmdeploy_model/mmseg/ort \
+ --show
+```
+
+**3. Run demo**
+
+```shell
+python inference_onnx.py ${ONNX_FILE_PATH} ${IMAGE_PATH} [${MODEL_INPUT_SIZE} ${DEVICE} ${OUTPUT_IMAGE_PATH}]
+```
+
+Example:
+
+```shell
+python inference_onnx.py mmdeploy_model/mmseg/ort/end2end.onnx ../../demo/demo.png
+```
+
+## Citation
+
+If you find our project useful in your research, please consider citing:
+
+```
+@misc{liu2021paddleseg,
+ title={PaddleSeg: A High-Efficient Development Toolkit for Image Segmentation},
+ author={Yi Liu and Lutao Chu and Guowei Chen and Zewu Wu and Zeyu Chen and Baohua Lai and Yuying Hao},
+ year={2021},
+ eprint={2101.06175},
+ archivePrefix={arXiv},
+ primaryClass={cs.CV}
+}
+
+@misc{paddleseg2019,
+ title={PaddleSeg, End-to-end image segmentation kit based on PaddlePaddle},
+ author={PaddlePaddle Contributors},
+ howpublished = {\url{https://github.com/PaddlePaddle/PaddleSeg}},
+ year={2019}
+}
+```
diff --git a/projects/pp_mobileseg/backbones/__init__.py b/projects/pp_mobileseg/backbones/__init__.py
new file mode 100644
index 00000000000..244b33d37a8
--- /dev/null
+++ b/projects/pp_mobileseg/backbones/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .strideformer import StrideFormer
+
+__all__ = ['StrideFormer']
diff --git a/projects/pp_mobileseg/backbones/strideformer.py b/projects/pp_mobileseg/backbones/strideformer.py
new file mode 100644
index 00000000000..3f09be5225d
--- /dev/null
+++ b/projects/pp_mobileseg/backbones/strideformer.py
@@ -0,0 +1,958 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import math
+import warnings
+
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import ConvModule, build_activation_layer
+from mmcv.cnn.bricks.transformer import build_dropout
+from mmengine.logging import print_log
+from mmengine.model import BaseModule
+from mmengine.runner.checkpoint import CheckpointLoader, load_state_dict
+
+from mmseg.registry import MODELS
+
+
+@MODELS.register_module()
+class StrideFormer(BaseModule):
+ """The StrideFormer implementation based on torch.
+
+ The original article refers to:https://arxiv.org/abs/2304.05152
+ Args:
+ mobileV3_cfg(list): Each sublist describe the config for a
+ MobileNetV3 block.
+ channels(list): The input channels for each MobileNetV3 block.
+ embed_dims(list): The channels of the features input to the sea
+ attention block.
+ key_dims(list, optional): The embeding dims for each head in
+ attention.
+ depths(list, optional): describes the depth of the attention block.
+ i,e: M,N.
+ num_heads(int, optional): The number of heads of the attention
+ blocks.
+ attn_ratios(int, optional): The expand ratio of V.
+ mlp_ratios(list, optional): The ratio of mlp blocks.
+ drop_path_rate(float, optional): The drop path rate in attention
+ block.
+ act_cfg(dict, optional): The activation layer of AAM:
+ Aggregate Attention Module.
+ inj_type(string, optional): The type of injection/AAM.
+ out_channels(int, optional): The output channels of the AAM.
+ dims(list, optional): The dimension of the fusion block.
+ out_feat_chs(list, optional): The input channels of the AAM.
+ stride_attention(bool, optional): whether to stride attention in
+ each attention layer.
+ pretrained(str, optional): the path of pretrained model.
+ """
+
+ def __init__(
+ self,
+ mobileV3_cfg,
+ channels,
+ embed_dims,
+ key_dims=[16, 24],
+ depths=[2, 2],
+ num_heads=8,
+ attn_ratios=2,
+ mlp_ratios=[2, 4],
+ drop_path_rate=0.1,
+ act_cfg=dict(type='ReLU'),
+ inj_type='AAM',
+ out_channels=256,
+ dims=(128, 160),
+ out_feat_chs=None,
+ stride_attention=True,
+ pretrained=None,
+ init_cfg=None,
+ ):
+ super().__init__(init_cfg=init_cfg)
+ assert not (init_cfg and pretrained
+ ), 'init_cfg and pretrained cannot be set at the same time'
+ if isinstance(pretrained, str):
+ warnings.warn('DeprecationWarning: pretrained is deprecated, '
+ 'please use "init_cfg" instead')
+ self.init_cfg = dict(type='Pretrained', checkpoint=pretrained)
+ elif pretrained is not None:
+ raise TypeError('pretrained must be a str or None')
+
+ self.depths = depths
+ self.cfgs = mobileV3_cfg
+ self.dims = dims
+ for i in range(len(self.cfgs)):
+ smb = StackedMV3Block(
+ cfgs=self.cfgs[i],
+ stem=True if i == 0 else False,
+ in_channels=channels[i],
+ )
+ setattr(self, f'smb{i + 1}', smb)
+ for i in range(len(depths)):
+ dpr = [
+ x.item() for x in torch.linspace(0, drop_path_rate, depths[i])
+ ]
+ trans = BasicLayer(
+ block_num=depths[i],
+ embedding_dim=embed_dims[i],
+ key_dim=key_dims[i],
+ num_heads=num_heads,
+ mlp_ratio=mlp_ratios[i],
+ attn_ratio=attn_ratios,
+ drop=0,
+ attn_drop=0.0,
+ drop_path=dpr,
+ act_cfg=act_cfg,
+ stride_attention=stride_attention,
+ )
+ setattr(self, f'trans{i + 1}', trans)
+
+ self.inj_type = inj_type
+ if self.inj_type == 'AAM':
+ self.inj_module = InjectionMultiSumallmultiallsum(
+ in_channels=out_feat_chs, out_channels=out_channels)
+ self.feat_channels = [
+ out_channels,
+ ]
+ elif self.inj_type == 'AAMSx8':
+ self.inj_module = InjectionMultiSumallmultiallsumSimpx8(
+ in_channels=out_feat_chs, out_channels=out_channels)
+ self.feat_channels = [
+ out_channels,
+ ]
+ elif self.inj_type == 'origin':
+ for i in range(len(dims)):
+ fuse = FusionBlock(
+ out_feat_chs[0] if i == 0 else dims[i - 1],
+ out_feat_chs[i + 1],
+ embed_dim=dims[i],
+ act_cfg=None,
+ )
+ setattr(self, f'fuse{i + 1}', fuse)
+ self.feat_channels = [
+ dims[i],
+ ]
+ else:
+ raise NotImplementedError(self.inj_module + ' is not implemented')
+
+ self.pretrained = pretrained
+ # self.init_weights()
+
+ def init_weights(self):
+ if (isinstance(self.init_cfg, dict)
+ and self.init_cfg.get('type') == 'Pretrained'):
+ checkpoint = CheckpointLoader.load_checkpoint(
+ self.init_cfg['checkpoint'], logger=None, map_location='cpu')
+
+ if 'state_dict' in checkpoint:
+ state_dict = checkpoint['state_dict']
+ else:
+ state_dict = checkpoint
+
+ if 'pos_embed' in state_dict.keys():
+ if self.pos_embed.shape != state_dict['pos_embed'].shape:
+ print_log(msg=f'Resize the pos_embed shape from '
+ f'{state_dict["pos_embed"].shape} to '
+ f'{self.pos_embed.shape}')
+ h, w = self.img_size
+ pos_size = int(
+ math.sqrt(state_dict['pos_embed'].shape[1] - 1))
+ state_dict['pos_embed'] = self.resize_pos_embed(
+ state_dict['pos_embed'],
+ (h // self.patch_size, w // self.patch_size),
+ (pos_size, pos_size),
+ self.interpolate_mode,
+ )
+
+ load_state_dict(self, state_dict, strict=False, logger=None)
+
+ def forward(self, x):
+ x_hw = x.shape[2:]
+ outputs = []
+ num_smb_stage = len(self.cfgs)
+ num_trans_stage = len(self.depths)
+
+ for i in range(num_smb_stage):
+ smb = getattr(self, f'smb{i + 1}')
+ x = smb(x)
+
+ # 1/8 shared feat
+ if i == 1:
+ outputs.append(x)
+ if num_trans_stage + i >= num_smb_stage:
+ trans = getattr(
+ self, f'trans{i + num_trans_stage - num_smb_stage + 1}')
+ x = trans(x)
+ outputs.append(x)
+ if self.inj_type == 'origin':
+ x_detail = outputs[0]
+ for i in range(len(self.dims)):
+ fuse = getattr(self, f'fuse{i + 1}')
+
+ x_detail = fuse(x_detail, outputs[i + 1])
+ output = x_detail
+ else:
+ output = self.inj_module(outputs)
+
+ return [output, x_hw]
+
+
+class StackedMV3Block(nn.Module):
+ """The MobileNetV3 block.
+
+ Args:
+ cfgs (list): The MobileNetV3 config list of a stage.
+ stem (bool): Whether is the first stage or not.
+ in_channels (int, optional): The channels of input image. Default: 3.
+ scale: float=1.0.
+ The coefficient that controls the size of network parameters.
+
+ Returns:
+ model: nn.Module.
+ A stage of specific MobileNetV3 model depends on args.
+ """
+
+ def __init__(self,
+ cfgs,
+ stem,
+ in_channels,
+ scale=1.0,
+ norm_cfg=dict(type='BN')):
+ super().__init__()
+
+ self.scale = scale
+ self.stem = stem
+
+ if self.stem:
+ self.conv = ConvModule(
+ in_channels=3,
+ out_channels=_make_divisible(in_channels * self.scale),
+ kernel_size=3,
+ stride=2,
+ padding=1,
+ groups=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=dict(type='HSwish'),
+ )
+
+ self.blocks = nn.ModuleList()
+ for i, (k, exp, c, se, act, s) in enumerate(cfgs):
+ self.blocks.append(
+ ResidualUnit(
+ in_channel=_make_divisible(in_channels * self.scale),
+ mid_channel=_make_divisible(self.scale * exp),
+ out_channel=_make_divisible(self.scale * c),
+ kernel_size=k,
+ stride=s,
+ use_se=se,
+ act=act,
+ dilation=1,
+ ))
+ in_channels = _make_divisible(self.scale * c)
+
+ def forward(self, x):
+ if self.stem:
+ x = self.conv(x)
+ for i, block in enumerate(self.blocks):
+ x = block(x)
+
+ return x
+
+
+class ResidualUnit(nn.Module):
+ """The Residual module.
+
+ Args:
+ in_channel (int, optional): The channels of input feature.
+ mid_channel (int, optional): The channels of middle process.
+ out_channel (int, optional): The channels of output feature.
+ kernel_size (int, optional): The size of the convolving kernel.
+ stride (int, optional): The stride size.
+ use_se (bool, optional): if to use the SEModule.
+ act (string, optional): activation layer.
+ dilation (int, optional): The dilation size.
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN', requires_grad=True).
+ """
+
+ def __init__(
+ self,
+ in_channel,
+ mid_channel,
+ out_channel,
+ kernel_size,
+ stride,
+ use_se,
+ act=None,
+ dilation=1,
+ norm_cfg=dict(type='BN'),
+ ):
+ super().__init__()
+ self.if_shortcut = stride == 1 and in_channel == out_channel
+ self.if_se = use_se
+ self.expand_conv = ConvModule(
+ in_channels=in_channel,
+ out_channels=mid_channel,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=dict(type=act) if act is not None else None,
+ )
+ self.bottleneck_conv = ConvModule(
+ in_channels=mid_channel,
+ out_channels=mid_channel,
+ kernel_size=kernel_size,
+ stride=stride,
+ padding=int((kernel_size - 1) // 2) * dilation,
+ bias=False,
+ groups=mid_channel,
+ dilation=dilation,
+ norm_cfg=norm_cfg,
+ act_cfg=dict(type=act) if act is not None else None,
+ )
+ if self.if_se:
+ self.mid_se = SEModule(mid_channel)
+ self.linear_conv = ConvModule(
+ in_channels=mid_channel,
+ out_channels=out_channel,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+
+ def forward(self, x):
+ identity = x
+ x = self.expand_conv(x)
+ x = self.bottleneck_conv(x)
+ if self.if_se:
+ x = self.mid_se(x)
+ x = self.linear_conv(x)
+ if self.if_shortcut:
+ x = torch.add(identity, x)
+ return x
+
+
+class SEModule(nn.Module):
+ """SE Module.
+
+ Args:
+ channel (int, optional): The channels of input feature.
+ reduction (int, optional): The channel reduction rate.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ """
+
+ def __init__(self, channel, reduction=4, act_cfg=dict(type='ReLU')):
+ super().__init__()
+ self.avg_pool = nn.AdaptiveAvgPool2d(1)
+ self.conv_act1 = ConvModule(
+ in_channels=channel,
+ out_channels=channel // reduction,
+ kernel_size=1,
+ norm_cfg=None,
+ act_cfg=act_cfg,
+ )
+
+ self.conv_act2 = ConvModule(
+ in_channels=channel // reduction,
+ out_channels=channel,
+ kernel_size=1,
+ norm_cfg=None,
+ act_cfg=dict(type='Hardsigmoid', slope=0.2, offset=0.5),
+ )
+
+ def forward(self, x):
+ identity = x
+ x = self.avg_pool(x)
+ x = self.conv_act1(x)
+ x = self.conv_act2(x)
+ return torch.mul(identity, x)
+
+
+class BasicLayer(nn.Module):
+ """The transformer basic layer.
+
+ Args:
+ block_num (int): the block nums of the transformer basic layer.
+ embedding_dim (int): The feature dimension.
+ key_dim (int): the key dim.
+ num_heads (int): Parallel attention heads.
+ mlp_ratio (float): the mlp ratio.
+ attn_ratio (float): the attention ratio.
+ drop (float): Probability of an element to be zeroed
+ after the feed forward layer.Default: 0.0.
+ attn_drop (float): The drop out rate for attention layer.
+ Default: 0.0.
+ drop_path (float): stochastic depth rate. Default 0.0.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ stride_attention (bool, optional): whether to stride attention in
+ each attention layer.
+ """
+
+ def __init__(
+ self,
+ block_num,
+ embedding_dim,
+ key_dim,
+ num_heads,
+ mlp_ratio=4.0,
+ attn_ratio=2.0,
+ drop=0.0,
+ attn_drop=0.0,
+ drop_path=None,
+ act_cfg=None,
+ stride_attention=None,
+ ):
+ super().__init__()
+ self.block_num = block_num
+
+ self.transformer_blocks = nn.ModuleList()
+ for i in range(self.block_num):
+ self.transformer_blocks.append(
+ Block(
+ embedding_dim,
+ key_dim=key_dim,
+ num_heads=num_heads,
+ mlp_ratio=mlp_ratio,
+ attn_ratio=attn_ratio,
+ drop=drop,
+ drop_path=drop_path[i]
+ if isinstance(drop_path, list) else drop_path,
+ act_cfg=act_cfg,
+ stride_attention=stride_attention,
+ ))
+
+ def forward(self, x):
+ for i in range(self.block_num):
+ x = self.transformer_blocks[i](x)
+ return x
+
+
+class Block(nn.Module):
+ """the block of the transformer basic layer.
+
+ Args:
+ dim (int): The feature dimension.
+ key_dim (int): The key dimension.
+ num_heads (int): Parallel attention heads.
+ mlp_ratio (float): the mlp ratio.
+ attn_ratio (float): the attention ratio.
+ drop (float): Probability of an element to be zeroed
+ after the feed forward layer.Default: 0.0.
+ drop_path (float): stochastic depth rate. Default 0.0.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ stride_attention (bool, optional): whether to stride attention in
+ each attention layer.
+ """
+
+ def __init__(
+ self,
+ dim,
+ key_dim,
+ num_heads,
+ mlp_ratio=4.0,
+ attn_ratio=2.0,
+ drop=0.0,
+ drop_path=0.0,
+ act_cfg=None,
+ stride_attention=None,
+ ):
+ super().__init__()
+ self.dim = dim
+ self.num_heads = num_heads
+ self.mlp_ratio = mlp_ratio
+ self.attn = SeaAttention(
+ dim,
+ key_dim=key_dim,
+ num_heads=num_heads,
+ attn_ratio=attn_ratio,
+ act_cfg=act_cfg,
+ stride_attention=stride_attention,
+ )
+ self.drop_path = (
+ build_dropout(dict(type='DropPath', drop_prob=drop_path))
+ if drop_path > 0.0 else nn.Identity())
+ mlp_hidden_dim = int(dim * mlp_ratio)
+ self.mlp = MLP(
+ in_features=dim,
+ hidden_features=mlp_hidden_dim,
+ act_cfg=act_cfg,
+ drop=drop,
+ )
+
+ def forward(self, x1):
+ x1 = x1 + self.drop_path(self.attn(x1))
+ x1 = x1 + self.drop_path(self.mlp(x1))
+
+ return x1
+
+
+class SqueezeAxialPositionalEmbedding(nn.Module):
+ """the Squeeze Axial Positional Embedding.
+
+ Args:
+ dim (int): The feature dimension.
+ shape (int): The patch size.
+ """
+
+ def __init__(self, dim, shape):
+ super().__init__()
+ self.pos_embed = nn.init.normal_(
+ nn.Parameter(torch.zeros(1, dim, shape)))
+
+ def forward(self, x):
+ B, C, N = x.shape
+ x = x + F.interpolate(
+ self.pos_embed, size=(N, ), mode='linear', align_corners=False)
+ return x
+
+
+class SeaAttention(nn.Module):
+ """The sea attention.
+
+ Args:
+ dim (int): The feature dimension.
+ key_dim (int): The key dimension.
+ num_heads (int): number of attention heads.
+ attn_ratio (float): the attention ratio.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='LN')
+ stride_attention (bool, optional): whether to stride attention in
+ each attention layer.
+ """
+
+ def __init__(
+ self,
+ dim,
+ key_dim,
+ num_heads,
+ attn_ratio=4.0,
+ act_cfg=None,
+ norm_cfg=dict(type='BN'),
+ stride_attention=False,
+ ):
+
+ super().__init__()
+ self.num_heads = num_heads
+ self.scale = key_dim**-0.5
+ self.nh_kd = nh_kd = key_dim * num_heads
+ self.d = int(attn_ratio * key_dim)
+ self.dh = int(attn_ratio * key_dim) * num_heads
+ self.attn_ratio = attn_ratio
+
+ self.to_q = ConvModule(
+ dim, nh_kd, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+ self.to_k = ConvModule(
+ dim, nh_kd, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+
+ self.to_v = ConvModule(
+ dim, self.dh, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+ self.stride_attention = stride_attention
+ if self.stride_attention:
+ self.stride_conv = ConvModule(
+ dim,
+ dim,
+ kernel_size=3,
+ stride=2,
+ padding=1,
+ bias=True,
+ groups=dim,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+
+ self.proj = ConvModule(
+ self.dh,
+ dim,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ order=('act', 'conv', 'norm'),
+ )
+ self.proj_encode_row = ConvModule(
+ self.dh,
+ self.dh,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ order=('act', 'conv', 'norm'),
+ )
+ self.pos_emb_rowq = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.pos_emb_rowk = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.proj_encode_column = ConvModule(
+ self.dh,
+ self.dh,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ order=('act', 'conv', 'norm'),
+ )
+ self.pos_emb_columnq = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.pos_emb_columnk = SqueezeAxialPositionalEmbedding(nh_kd, 16)
+ self.dwconv = ConvModule(
+ 2 * self.dh,
+ 2 * self.dh,
+ 3,
+ padding=1,
+ groups=2 * self.dh,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ )
+ self.pwconv = ConvModule(
+ 2 * self.dh, dim, 1, bias=False, norm_cfg=norm_cfg, act_cfg=None)
+ self.sigmoid = build_activation_layer(dict(type='HSigmoid'))
+
+ def forward(self, x):
+ B, C, H_ori, W_ori = x.shape
+ if self.stride_attention:
+ x = self.stride_conv(x)
+ B, C, H, W = x.shape
+
+ q = self.to_q(x) # [B, nhead*dim, H, W]
+ k = self.to_k(x)
+ v = self.to_v(x)
+
+ qkv = torch.cat([q, k, v], dim=1)
+ qkv = self.dwconv(qkv)
+ qkv = self.pwconv(qkv)
+
+ qrow = (self.pos_emb_rowq(q.mean(-1)).reshape(
+ [B, self.num_heads, -1, H]).permute(
+ (0, 1, 3, 2))) # [B, nhead, H, dim]
+ krow = self.pos_emb_rowk(k.mean(-1)).reshape(
+ [B, self.num_heads, -1, H]) # [B, nhead, dim, H]
+ vrow = (v.mean(-1).reshape([B, self.num_heads, -1,
+ H]).permute([0, 1, 3, 2])
+ ) # [B, nhead, H, dim*attn_ratio]
+
+ attn_row = torch.matmul(qrow, krow) * self.scale # [B, nhead, H, H]
+ attn_row = nn.functional.softmax(attn_row, dim=-1)
+
+ xx_row = torch.matmul(attn_row, vrow) # [B, nhead, H, dim*attn_ratio]
+ xx_row = self.proj_encode_row(
+ xx_row.permute([0, 1, 3, 2]).reshape([B, self.dh, H, 1]))
+
+ # squeeze column
+ qcolumn = (
+ self.pos_emb_columnq(q.mean(-2)).reshape(
+ [B, self.num_heads, -1, W]).permute([0, 1, 3, 2]))
+ kcolumn = self.pos_emb_columnk(k.mean(-2)).reshape(
+ [B, self.num_heads, -1, W])
+ vcolumn = (
+ torch.mean(v, -2).reshape([B, self.num_heads, -1,
+ W]).permute([0, 1, 3, 2]))
+
+ attn_column = torch.matmul(qcolumn, kcolumn) * self.scale
+ attn_column = nn.functional.softmax(attn_column, dim=-1)
+
+ xx_column = torch.matmul(attn_column, vcolumn) # B nH W C
+ xx_column = self.proj_encode_column(
+ xx_column.permute([0, 1, 3, 2]).reshape([B, self.dh, 1, W]))
+
+ xx = torch.add(xx_row, xx_column) # [B, self.dh, H, W]
+ xx = torch.add(v, xx)
+
+ xx = self.proj(xx)
+ xx = self.sigmoid(xx) * qkv
+ if self.stride_attention:
+ xx = F.interpolate(xx, size=(H_ori, W_ori), mode='bilinear')
+
+ return xx
+
+
+class MLP(nn.Module):
+ """the Multilayer Perceptron.
+
+ Args:
+ in_features (int): the input feature.
+ hidden_features (int): the hidden feature.
+ out_features (int): the output feature.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='PReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ drop (float): Probability of an element to be zeroed.
+ Default 0.0
+ """
+
+ def __init__(
+ self,
+ in_features,
+ hidden_features=None,
+ out_features=None,
+ act_cfg=None,
+ norm_cfg=dict(type='BN'),
+ drop=0.0,
+ ):
+ super().__init__()
+ out_features = out_features or in_features
+ hidden_features = hidden_features or in_features
+ self.fc1 = ConvModule(
+ in_features,
+ hidden_features,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+ self.dwconv = ConvModule(
+ hidden_features,
+ hidden_features,
+ kernel_size=3,
+ padding=1,
+ groups=hidden_features,
+ norm_cfg=None,
+ act_cfg=act_cfg,
+ )
+
+ self.fc2 = ConvModule(
+ hidden_features,
+ out_features,
+ 1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+ self.drop = build_dropout(dict(type='Dropout', drop_prob=drop))
+
+ def forward(self, x):
+ x = self.fc1(x)
+ x = self.dwconv(x)
+ x = self.drop(x)
+ x = self.fc2(x)
+ x = self.drop(x)
+ return x
+
+
+class FusionBlock(nn.Module):
+ """The feature fusion block.
+
+ Args:
+ in_channel (int): the input channel.
+ out_channel (int): the output channel.
+ embed_dim (int): embedding dimension.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ """
+
+ def __init__(
+ self,
+ in_channel,
+ out_channel,
+ embed_dim,
+ norm_cfg=dict(type='BN'),
+ act_cfg=dict(type='ReLU'),
+ ) -> None:
+ super().__init__()
+ self.local_embedding = ConvModule(
+ in_channels=in_channel,
+ out_channels=embed_dim,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ )
+
+ self.global_act = ConvModule(
+ in_channels=out_channel,
+ out_channels=embed_dim,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg if act_cfg is not None else None,
+ )
+
+ def forward(self, x_l, x_g):
+ """
+ x_g: global features
+ x_l: local features
+ """
+ B, C, H, W = x_l.shape
+
+ local_feat = self.local_embedding(x_l)
+ global_act = self.global_act(x_g)
+ sig_act = F.interpolate(
+ global_act, size=(H, W), mode='bilinear', align_corners=False)
+
+ out = local_feat * sig_act
+
+ return out
+
+
+class InjectionMultiSumallmultiallsum(nn.Module):
+ """the Aggregate Attention Module.
+
+ Args:
+ in_channels (tuple): the input channel.
+ out_channels (int): the output channel.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ """
+
+ def __init__(
+ self,
+ in_channels=(64, 128, 256, 384),
+ out_channels=256,
+ act_cfg=dict(type='Sigmoid'),
+ norm_cfg=dict(type='BN'),
+ ):
+ super().__init__()
+ self.embedding_list = nn.ModuleList()
+ self.act_embedding_list = nn.ModuleList()
+ self.act_list = nn.ModuleList()
+ for i in range(len(in_channels)):
+ self.embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ ))
+ self.act_embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ))
+
+ def forward(self, inputs): # x_x8, x_x16, x_x32, x_x64
+ low_feat1 = F.interpolate(inputs[0], scale_factor=0.5, mode='bilinear')
+ low_feat1_act = self.act_embedding_list[0](low_feat1)
+ low_feat1 = self.embedding_list[0](low_feat1)
+
+ low_feat2 = F.interpolate(
+ inputs[1], size=low_feat1.shape[-2:], mode='bilinear')
+ low_feat2_act = self.act_embedding_list[1](low_feat2) # x16
+ low_feat2 = self.embedding_list[1](low_feat2)
+
+ high_feat_act = F.interpolate(
+ self.act_embedding_list[2](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear',
+ )
+ high_feat = F.interpolate(
+ self.embedding_list[2](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear')
+
+ res = (
+ low_feat1_act * low_feat2_act * high_feat_act *
+ (low_feat1 + low_feat2) + high_feat)
+
+ return res
+
+
+class InjectionMultiSumallmultiallsumSimpx8(nn.Module):
+ """the Aggregate Attention Module.
+
+ Args:
+ in_channels (tuple): the input channel.
+ out_channels (int): the output channel.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN')
+ """
+
+ def __init__(
+ self,
+ in_channels=(64, 128, 256, 384),
+ out_channels=256,
+ act_cfg=dict(type='Sigmoid'),
+ norm_cfg=dict(type='BN'),
+ ):
+ super().__init__()
+ self.embedding_list = nn.ModuleList()
+ self.act_embedding_list = nn.ModuleList()
+ self.act_list = nn.ModuleList()
+ for i in range(len(in_channels)):
+ if i != 1:
+ self.embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=None,
+ ))
+ if i != 0:
+ self.act_embedding_list.append(
+ ConvModule(
+ in_channels=in_channels[i],
+ out_channels=out_channels,
+ kernel_size=1,
+ bias=False,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg,
+ ))
+
+ def forward(self, inputs):
+ # x_x8, x_x16, x_x32
+ low_feat1 = self.embedding_list[0](inputs[0])
+
+ low_feat2 = F.interpolate(
+ inputs[1], size=low_feat1.shape[-2:], mode='bilinear')
+ low_feat2_act = self.act_embedding_list[0](low_feat2)
+
+ high_feat_act = F.interpolate(
+ self.act_embedding_list[1](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear',
+ )
+ high_feat = F.interpolate(
+ self.embedding_list[1](inputs[2]),
+ size=low_feat2.shape[2:],
+ mode='bilinear')
+
+ res = low_feat2_act * high_feat_act * low_feat1 + high_feat
+
+ return res
+
+
+def _make_divisible(v, divisor=8, min_value=None):
+ if min_value is None:
+ min_value = divisor
+ new_v = max(min_value, int(v + divisor / 2) // divisor * divisor)
+ if new_v < 0.9 * v:
+ new_v += divisor
+ return new_v
+
+
+@MODELS.register_module()
+class Hardsigmoid(nn.Module):
+ """the hardsigmoid activation.
+
+ Args:
+ slope (float, optional): The slope of hardsigmoid function.
+ Default is 0.1666667.
+ offset (float, optional): The offset of hardsigmoid function.
+ Default is 0.5.
+ inplace (bool): can optionally do the operation in-place.
+ Default: ``False``
+ """
+
+ def __init__(self, slope=0.1666667, offset=0.5, inplace=False):
+ super().__init__()
+ self.slope = slope
+ self.offset = offset
+
+ def forward(self, x):
+ return (x * self.slope + self.offset).clamp(0, 1)
diff --git a/projects/pp_mobileseg/configs/_base_/datasets/ade20k.py b/projects/pp_mobileseg/configs/_base_/datasets/ade20k.py
new file mode 100644
index 00000000000..48340d11eea
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/datasets/ade20k.py
@@ -0,0 +1,68 @@
+# dataset settings
+dataset_type = 'ADE20KDataset'
+data_root = 'data/ade/ADEChallengeData2016'
+crop_size = (512, 512)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations', reduce_zero_label=True),
+ dict(
+ type='RandomResize',
+ scale=(2048, 512),
+ ratio_range=(0.5, 2.0),
+ keep_ratio=True),
+ dict(type='RandomCrop', crop_size=crop_size, cat_max_ratio=0.75),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 512), keep_ratio=True),
+ # add loading annotation after ``Resize`` because ground truth
+ # does not need to do resize data transform
+ dict(type='LoadAnnotations', reduce_zero_label=True),
+ dict(type='PackSegInputs')
+]
+img_ratios = [0.5, 0.75, 1.0, 1.25, 1.5, 1.75]
+tta_pipeline = [
+ dict(type='LoadImageFromFile', backend_args=None),
+ dict(
+ type='TestTimeAug',
+ transforms=[
+ [
+ dict(type='Resize', scale_factor=r, keep_ratio=True)
+ for r in img_ratios
+ ],
+ [
+ dict(type='RandomFlip', prob=0., direction='horizontal'),
+ dict(type='RandomFlip', prob=1., direction='horizontal')
+ ], [dict(type='LoadAnnotations')], [dict(type='PackSegInputs')]
+ ])
+]
+train_dataloader = dict(
+ batch_size=4,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/training', seg_map_path='annotations/training'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/validation',
+ seg_map_path='annotations/validation'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU'])
+test_evaluator = val_evaluator
diff --git a/projects/pp_mobileseg/configs/_base_/default_runtime.py b/projects/pp_mobileseg/configs/_base_/default_runtime.py
new file mode 100644
index 00000000000..272b4d24679
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/default_runtime.py
@@ -0,0 +1,15 @@
+default_scope = 'mmseg'
+env_cfg = dict(
+ cudnn_benchmark=True,
+ mp_cfg=dict(mp_start_method='fork', opencv_num_threads=0),
+ dist_cfg=dict(backend='nccl'),
+)
+vis_backends = [dict(type='LocalVisBackend')]
+visualizer = dict(
+ type='SegLocalVisualizer', vis_backends=vis_backends, name='visualizer')
+log_processor = dict(by_epoch=False)
+log_level = 'INFO'
+load_from = None
+resume = False
+
+tta_model = dict(type='SegTTAModel')
diff --git a/projects/pp_mobileseg/configs/_base_/models/pp_mobile.py b/projects/pp_mobileseg/configs/_base_/models/pp_mobile.py
new file mode 100644
index 00000000000..0c7695636f6
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/models/pp_mobile.py
@@ -0,0 +1,47 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255)
+
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ # pretrained='open-mmlab://resnet50_v1c',
+ backbone=dict(
+ type='StrideFormer',
+ mobileV3_cfg=[
+ # k t c, s
+ [[3, 16, 16, True, 'ReLU', 1], [3, 64, 32, False, 'ReLU', 2],
+ [3, 96, 32, False, 'ReLU', 1]], # cfg1
+ [[5, 128, 64, True, 'HSwish', 2], [5, 240, 64, True, 'HSwish',
+ 1]], # cfg2
+ [[5, 384, 128, True, 'HSwish', 2],
+ [5, 384, 128, True, 'HSwish', 1]], # cfg3
+ [[5, 768, 192, True, 'HSwish', 2],
+ [5, 768, 192, True, 'HSwish', 1]], # cfg4
+ ],
+ channels=[16, 32, 64, 128, 192],
+ depths=[3, 3],
+ embed_dims=[128, 192],
+ num_heads=8,
+ inj_type='AAMSx8',
+ out_feat_chs=[64, 128, 192],
+ act_cfg=dict(type='ReLU6'),
+ ),
+ decode_head=dict(
+ type='PPMobileSegHead',
+ num_classes=150,
+ in_channels=256,
+ dropout_ratio=0.1,
+ use_dw=True,
+ act_cfg=dict(type='ReLU'),
+ align_corners=False),
+
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/pp_mobileseg/configs/_base_/schedules/schedule_80k.py b/projects/pp_mobileseg/configs/_base_/schedules/schedule_80k.py
new file mode 100644
index 00000000000..0dcd6c4d1bc
--- /dev/null
+++ b/projects/pp_mobileseg/configs/_base_/schedules/schedule_80k.py
@@ -0,0 +1,24 @@
+# optimizer
+optimizer = dict(type='SGD', lr=0.01, momentum=0.9, weight_decay=0.0005)
+optim_wrapper = dict(type='OptimWrapper', optimizer=optimizer, clip_grad=None)
+# learning policy
+param_scheduler = [
+ dict(
+ type='PolyLR',
+ eta_min=1e-4,
+ power=0.9,
+ begin=0,
+ end=80000,
+ by_epoch=False)
+]
+# training schedule for 80k
+train_cfg = dict(type='IterBasedTrainLoop', max_iters=80000, val_interval=8000)
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
+default_hooks = dict(
+ timer=dict(type='IterTimerHook'),
+ logger=dict(type='LoggerHook', interval=50, log_metric_by_epoch=False),
+ param_scheduler=dict(type='ParamSchedulerHook'),
+ checkpoint=dict(type='CheckpointHook', by_epoch=False, interval=8000),
+ sampler_seed=dict(type='DistSamplerSeedHook'),
+ visualization=dict(type='SegVisualizationHook'))
diff --git a/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py
new file mode 100644
index 00000000000..4b68a927e20
--- /dev/null
+++ b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_base.py
@@ -0,0 +1,18 @@
+_base_ = [
+ '../_base_/models/pp_mobile.py', '../_base_/datasets/ade20k.py',
+ '../_base_/default_runtime.py', '../_base_/schedules/schedule_80k.py'
+]
+# the custom import path is determined by your workspace path (i.e., where you run the command from) # noqa
+custom_imports = dict(
+ imports=[
+ 'projects.pp_mobileseg.backbones', 'projects.pp_mobileseg.decode_head'
+ ],
+ allow_failed_imports=False)
+checkpoint = 'https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_3rdparty-base-ed0be681.pth' # noqa
+crop_size = (512, 512)
+data_preprocessor = dict(size=crop_size, test_cfg=dict(size_divisor=32))
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(init_cfg=dict(type='Pretrained', checkpoint=checkpoint)),
+ decode_head=dict(num_classes=150, upsample='intepolate'))
diff --git a/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py
new file mode 100644
index 00000000000..b78869e517a
--- /dev/null
+++ b/projects/pp_mobileseg/configs/pp_mobileseg/pp_mobileseg_mobilenetv3_2x16_80k_ade20k_512x512_tiny.py
@@ -0,0 +1,45 @@
+_base_ = [
+ '../_base_/models/pp_mobile.py', '../_base_/datasets/ade20k.py',
+ '../_base_/default_runtime.py', '../_base_/schedules/schedule_80k.py'
+]
+# the custom import path is determined by your workspace path (i.e., where you run the command from) # noqa
+custom_imports = dict(
+ imports=[
+ 'projects.pp_mobileseg.backbones', 'projects.pp_mobileseg.decode_head'
+ ],
+ allow_failed_imports=False)
+checkpoint = 'https://download.openmmlab.com/mmsegmentation/v0.5/pp_mobileseg/pp_mobileseg_mobilenetv3_3rdparty-tiny-e4b35e96.pth' # noqa
+crop_size = (512, 512)
+data_preprocessor = dict(size=crop_size, test_cfg=dict(size_divisor=32))
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ init_cfg=dict(type='Pretrained', checkpoint=checkpoint),
+ type='StrideFormer',
+ mobileV3_cfg=[
+ # k t c, s
+ [[3, 16, 16, True, 'ReLU', 1], [3, 64, 32, False, 'ReLU', 2],
+ [3, 48, 24, False, 'ReLU', 1]], # cfg1
+ [[5, 96, 32, True, 'HSwish', 2], [5, 96, 32, True, 'HSwish',
+ 1]], # cfg2
+ [[5, 160, 64, True, 'HSwish', 2], [5, 160, 64, True, 'HSwish',
+ 1]], # cfg3
+ [[3, 384, 128, True, 'HSwish', 2],
+ [3, 384, 128, True, 'HSwish', 1]], # cfg4
+ ],
+ channels=[16, 24, 32, 64, 128],
+ depths=[2, 2],
+ embed_dims=[64, 128],
+ num_heads=4,
+ inj_type='AAM',
+ out_feat_chs=[32, 64, 128],
+ act_cfg=dict(type='ReLU6'),
+ ),
+ decode_head=dict(
+ num_classes=150,
+ in_channels=256,
+ use_dw=True,
+ act_cfg=dict(type='ReLU'),
+ upsample='intepolate'),
+)
diff --git a/projects/pp_mobileseg/decode_head/__init__.py b/projects/pp_mobileseg/decode_head/__init__.py
new file mode 100644
index 00000000000..6f71b784e15
--- /dev/null
+++ b/projects/pp_mobileseg/decode_head/__init__.py
@@ -0,0 +1,6 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .pp_mobileseg_head import PPMobileSegHead
+
+__all__ = [
+ 'PPMobileSegHead',
+]
diff --git a/projects/pp_mobileseg/decode_head/pp_mobileseg_head.py b/projects/pp_mobileseg/decode_head/pp_mobileseg_head.py
new file mode 100644
index 00000000000..243f0263729
--- /dev/null
+++ b/projects/pp_mobileseg/decode_head/pp_mobileseg_head.py
@@ -0,0 +1,94 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from typing import List
+
+import torch
+import torch.nn as nn
+import torch.nn.functional as F
+from mmcv.cnn import ConvModule, build_conv_layer
+from torch import Tensor
+
+from mmseg.registry import MODELS
+
+
+@MODELS.register_module()
+class PPMobileSegHead(nn.Module):
+ """the segmentation head.
+
+ Args:
+ num_classes (int): the classes num.
+ in_channels (int): the input channels.
+ use_dw (bool): if to use deepwith convolution.
+ dropout_ratio (float): Probability of an element to be zeroed.
+ Default 0.0。
+ align_corners (bool, optional): Geometrically, we consider the pixels
+ of the input and output as squares rather than points.
+ upsample (str): the upsample method.
+ out_channels (int): the output channel.
+ conv_cfg (dict): Config dict for convolution layer.
+ act_cfg (dict): Config dict for activation layer.
+ Default: dict(type='ReLU').
+ norm_cfg (dict): Config dict for normalization layer.
+ Default: dict(type='BN').
+ """
+
+ def __init__(self,
+ num_classes,
+ in_channels,
+ use_dw=True,
+ dropout_ratio=0.1,
+ align_corners=False,
+ upsample='intepolate',
+ out_channels=None,
+ conv_cfg=dict(type='Conv'),
+ act_cfg=dict(type='ReLU'),
+ norm_cfg=dict(type='BN')):
+
+ super().__init__()
+ self.align_corners = align_corners
+ self.last_channels = in_channels
+ self.upsample = upsample
+ self.num_classes = num_classes
+ self.out_channels = out_channels
+ self.linear_fuse = ConvModule(
+ in_channels=self.last_channels,
+ out_channels=self.last_channels,
+ kernel_size=1,
+ bias=False,
+ groups=self.last_channels if use_dw else 1,
+ norm_cfg=norm_cfg,
+ act_cfg=act_cfg)
+ self.dropout = nn.Dropout2d(dropout_ratio)
+ self.conv_seg = build_conv_layer(
+ conv_cfg, self.last_channels, self.num_classes, kernel_size=1)
+
+ def forward(self, x):
+ x, x_hw = x[0], x[1]
+ x = self.linear_fuse(x)
+ x = self.dropout(x)
+ x = self.conv_seg(x)
+ if self.upsample == 'intepolate' or self.training or \
+ self.num_classes < 30:
+ x = F.interpolate(
+ x, x_hw, mode='bilinear', align_corners=self.align_corners)
+ elif self.upsample == 'vim':
+ labelset = torch.unique(torch.argmax(x, 1))
+ x = torch.gather(x, 1, labelset)
+ x = F.interpolate(
+ x, x_hw, mode='bilinear', align_corners=self.align_corners)
+
+ pred = torch.argmax(x, 1)
+ pred_retrieve = torch.zeros(pred.shape, dtype=torch.int32)
+ for i, val in enumerate(labelset):
+ pred_retrieve[pred == i] = labelset[i].cast('int32')
+
+ x = pred_retrieve
+ else:
+ raise NotImplementedError(self.upsample, ' is not implemented')
+
+ return [x]
+
+ def predict(self, inputs, batch_img_metas: List[dict], test_cfg,
+ **kwargs) -> List[Tensor]:
+ """Forward function for testing, only ``pam_cam`` is used."""
+ seg_logits = self.forward(inputs)[0]
+ return seg_logits
diff --git a/projects/pp_mobileseg/inference_onnx.py b/projects/pp_mobileseg/inference_onnx.py
new file mode 100644
index 00000000000..139d1b13243
--- /dev/null
+++ b/projects/pp_mobileseg/inference_onnx.py
@@ -0,0 +1,203 @@
+import argparse
+import time
+from typing import List, Tuple
+
+import cv2
+import loguru
+import numpy as np
+import onnxruntime as ort
+
+logger = loguru.logger
+
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='PP_Mobileseg ONNX inference demo.')
+ parser.add_argument('onnx_file', help='ONNX file path')
+ parser.add_argument('image_file', help='Input image file path')
+ parser.add_argument(
+ '--input-size',
+ type=int,
+ nargs='+',
+ default=[512, 512],
+ help='input image size')
+ parser.add_argument(
+ '--device', help='device type for inference', default='cpu')
+ parser.add_argument(
+ '--save-path',
+ help='path to save the output image',
+ default='output.jpg')
+ args = parser.parse_args()
+ return args
+
+
+def preprocess(
+ img: np.ndarray, input_size: Tuple[int, int] = (512, 512)
+) -> Tuple[np.ndarray, np.ndarray]:
+ """Preprocess image for inference."""
+ img_shape = img.shape[:2]
+ # Resize
+ resized_img = cv2.resize(img, input_size)
+
+ # Normalize
+ mean = np.array([123.575, 116.28, 103.53], dtype=np.float32)
+ std = np.array([58.395, 57.12, 57.375], dtype=np.float32)
+ resized_img = (resized_img - mean) / std
+
+ return resized_img, img_shape
+
+
+def build_session(onnx_file: str, device: str = 'cpu') -> ort.InferenceSession:
+ """Build onnxruntime session.
+
+ Args:
+ onnx_file (str): ONNX file path.
+ device (str): Device type for inference.
+
+ Returns:
+ sess (ort.InferenceSession): ONNXRuntime session.
+ """
+ providers = ['CPUExecutionProvider'
+ ] if device == 'cpu' else ['CUDAExecutionProvider']
+ sess = ort.InferenceSession(path_or_bytes=onnx_file, providers=providers)
+
+ return sess
+
+
+def inference(sess: ort.InferenceSession, img: np.ndarray) -> np.ndarray:
+ """Inference RTMPose model.
+
+ Args:
+ sess (ort.InferenceSession): ONNXRuntime session.
+ img (np.ndarray): Input image in shape.
+
+ Returns:
+ outputs (np.ndarray): Output of RTMPose model.
+ """
+ # build input
+ input_img = [img.transpose(2, 0, 1).astype(np.float32)]
+
+ # build output
+ sess_input = {sess.get_inputs()[0].name: input_img}
+ sess_output = []
+ for out in sess.get_outputs():
+ sess_output.append(out.name)
+
+ # inference
+ outputs = sess.run(output_names=sess_output, input_feed=sess_input)
+
+ return outputs
+
+
+def postprocess(outputs: List[np.ndarray],
+ origin_shape: Tuple[int, int]) -> np.ndarray:
+ """Postprocess outputs of PP_Mobileseg model.
+
+ Args:
+ outputs (List[np.ndarray]): Outputs of PP_Mobileseg model.
+ origin_shape (Tuple[int, int]): Input size of PP_Mobileseg model.
+
+ Returns:
+ seg_map (np.ndarray): Segmentation map.
+ """
+ seg_map = outputs[0][0][0]
+ seg_map = cv2.resize(seg_map.astype(np.float32), origin_shape)
+ return seg_map
+
+
+def visualize(img: np.ndarray,
+ seg_map: np.ndarray,
+ filename: str = 'output.jpg',
+ opacity: float = 0.8) -> np.ndarray:
+ assert 0.0 <= opacity <= 1.0, 'opacity should be in range [0, 1]'
+ palette = np.array(PALETTE)
+ color_seg = np.zeros((seg_map.shape[0], seg_map.shape[1], 3),
+ dtype=np.uint8)
+ for label, color in enumerate(palette):
+ color_seg[seg_map == label, :] = color
+ # convert to BGR
+ color_seg = color_seg[..., ::-1]
+
+ img = img * (1 - opacity) + color_seg * opacity
+ cv2.imwrite(filename, img)
+
+ return img
+
+
+def main():
+ args = parse_args()
+ logger.info('Start running model inference...')
+
+ # read image from file
+ logger.info(f'1. Read image from file {args.image_file}...')
+ img = cv2.imread(args.image_file)
+
+ # build onnx model
+ logger.info(f'2. Build onnx model from {args.onnx_file}...')
+ sess = build_session(args.onnx_file, args.device)
+
+ # preprocess
+ logger.info('3. Preprocess image...')
+ model_input_size = tuple(args.input_size)
+ assert len(model_input_size) == 2
+ resized_img, origin_shape = preprocess(img, model_input_size)
+
+ # inference
+ logger.info('4. Inference...')
+ start = time.time()
+ outputs = inference(sess, resized_img)
+ logger.info(f'Inference time: {time.time() - start:.4f}s')
+
+ # postprocess
+ logger.info('5. Postprocess...')
+ h, w = origin_shape
+ seg_map = postprocess(outputs, (w, h))
+
+ # visualize
+ logger.info('6. Visualize...')
+ visualize(img, seg_map, args.save_path)
+
+ logger.info('Done...')
+
+
+PALETTE = [[120, 120, 120], [180, 120, 120], [6, 230, 230], [80, 50, 50],
+ [4, 200, 3], [120, 120, 80], [140, 140, 140], [204, 5, 255],
+ [230, 230, 230], [4, 250, 7], [224, 5, 255], [235, 255, 7],
+ [150, 5, 61], [120, 120, 70], [8, 255, 51], [255, 6, 82],
+ [143, 255, 140], [204, 255, 4], [255, 51, 7], [204, 70, 3],
+ [0, 102, 200], [61, 230, 250], [255, 6, 51], [11, 102, 255],
+ [255, 7, 71], [255, 9, 224], [9, 7, 230], [220, 220, 220],
+ [255, 9, 92], [112, 9, 255], [8, 255, 214], [7, 255, 224],
+ [255, 184, 6], [10, 255, 71], [255, 41, 10], [7, 255, 255],
+ [224, 255, 8], [102, 8, 255], [255, 61, 6], [255, 194, 7],
+ [255, 122, 8], [0, 255, 20], [255, 8, 41], [255, 5, 153],
+ [6, 51, 255], [235, 12, 255], [160, 150, 20], [0, 163, 255],
+ [140, 140, 140], [250, 10, 15], [20, 255, 0], [31, 255, 0],
+ [255, 31, 0], [255, 224, 0], [153, 255, 0], [0, 0, 255],
+ [255, 71, 0], [0, 235, 255], [0, 173, 255], [31, 0, 255],
+ [11, 200, 200], [255, 82, 0], [0, 255, 245], [0, 61, 255],
+ [0, 255, 112], [0, 255, 133], [255, 0, 0], [255, 163, 0],
+ [255, 102, 0], [194, 255, 0], [0, 143, 255], [51, 255, 0],
+ [0, 82, 255], [0, 255, 41], [0, 255, 173], [10, 0, 255],
+ [173, 255, 0], [0, 255, 153], [255, 92, 0], [255, 0, 255],
+ [255, 0, 245], [255, 0, 102], [255, 173, 0], [255, 0, 20],
+ [255, 184, 184], [0, 31, 255], [0, 255, 61], [0, 71, 255],
+ [255, 0, 204], [0, 255, 194], [0, 255, 82], [0, 10, 255],
+ [0, 112, 255], [51, 0, 255], [0, 194, 255], [0, 122, 255],
+ [0, 255, 163], [255, 153, 0], [0, 255, 10], [255, 112, 0],
+ [143, 255, 0], [82, 0, 255], [163, 255, 0], [255, 235, 0],
+ [8, 184, 170], [133, 0, 255], [0, 255, 92], [184, 0, 255],
+ [255, 0, 31], [0, 184, 255], [0, 214, 255], [255, 0, 112],
+ [92, 255, 0], [0, 224, 255], [112, 224, 255], [70, 184, 160],
+ [163, 0, 255], [153, 0, 255], [71, 255, 0], [255, 0, 163],
+ [255, 204, 0], [255, 0, 143], [0, 255, 235], [133, 255, 0],
+ [255, 0, 235], [245, 0, 255], [255, 0, 122], [255, 245, 0],
+ [10, 190, 212], [214, 255, 0], [0, 204, 255], [20, 0, 255],
+ [255, 255, 0], [0, 153, 255], [0, 41, 255], [0, 255, 204],
+ [41, 0, 255], [41, 255, 0], [173, 0, 255], [0, 245, 255],
+ [71, 0, 255], [122, 0, 255], [0, 255, 184], [0, 92, 255],
+ [184, 255, 0], [0, 133, 255], [255, 214, 0], [25, 194, 194],
+ [102, 255, 0], [92, 0, 255]]
+
+if __name__ == '__main__':
+ main()
diff --git a/projects/sam_inference_demo/README.md b/projects/sam_inference_demo/README.md
new file mode 100644
index 00000000000..f8077b8729f
--- /dev/null
+++ b/projects/sam_inference_demo/README.md
@@ -0,0 +1,40 @@
+# Introducing the Segment Anything Model (SAM) Inference Demo!
+
+Welcome to the Segment Anything (SA) Inference Demo, a user-friendly implementation based on the original Segment Anything project. Our demo allows you to experience the power and versatility of the Segment Anything Model (SAM) through an easy-to-use API.
+
+With this inference demo, you can explore the capabilities of the Segment Anything Model and witness its effectiveness in various tasks and image distributions. For more information on the original project, dataset, and model, please visit the official website at https://segment-anything.com.
+
+### Prerequisites
+
+- Python 3.10
+- PyTorch 1.13
+- MMEngine >= v0.7.2
+- MMCV >= v2.0.0
+
+### Installation
+
+We assume that you have already installed PyTorch. If not, please follow the instructions on the [PyTorch website](https://pytorch.org/).
+
+**1. Install MMEngine & MMCV**
+
+```shell
+pip install openmim
+mim install mmengine
+mim install 'mmcv>=2.0.0'
+```
+
+**2. Install MMPretrain**
+
+```shell
+pip install git+https://github.com/open-mmlab/mmpretrain.git@dev
+```
+
+**3. Install MMSegmentation**
+
+```shell
+pip install mmsegmentation
+```
+
+### Usage
+
+Open the `sam_image_demo.ipynb` notebook and follow the instructions to run the demo.
diff --git a/projects/sam_inference_demo/sam/__init__.py b/projects/sam_inference_demo/sam/__init__.py
new file mode 100644
index 00000000000..82b6b78469c
--- /dev/null
+++ b/projects/sam_inference_demo/sam/__init__.py
@@ -0,0 +1,2 @@
+from .modeling import * # noqa
+from .utils import * # noqa
diff --git a/projects/sam_inference_demo/sam/modeling/__init__.py b/projects/sam_inference_demo/sam/modeling/__init__.py
new file mode 100644
index 00000000000..9892a6b085d
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/__init__.py
@@ -0,0 +1,12 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+from .mask_decoder import MaskDecoder
+from .prompt_encoder import PromptEncoder
+from .sam import SAM
+from .transformer import TwoWayTransformer
+
+__all__ = ['SAM', 'MaskDecoder', 'PromptEncoder', 'TwoWayTransformer']
diff --git a/projects/sam_inference_demo/sam/modeling/common.py b/projects/sam_inference_demo/sam/modeling/common.py
new file mode 100644
index 00000000000..d2892761122
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/common.py
@@ -0,0 +1,45 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+from typing import Type
+
+import torch
+import torch.nn as nn
+
+
+class MLPBlock(nn.Module):
+
+ def __init__(
+ self,
+ embedding_dim: int,
+ mlp_dim: int,
+ act: Type[nn.Module] = nn.GELU,
+ ) -> None:
+ super().__init__()
+ self.lin1 = nn.Linear(embedding_dim, mlp_dim)
+ self.lin2 = nn.Linear(mlp_dim, embedding_dim)
+ self.act = act()
+
+ def forward(self, x: torch.Tensor) -> torch.Tensor:
+ return self.lin2(self.act(self.lin1(x)))
+
+
+# From https://github.com/facebookresearch/detectron2/blob/main/detectron2/layers/batch_norm.py # noqa
+# Itself from https://github.com/facebookresearch/ConvNeXt/blob/d1fa8f6fef0a165b27399986cc2bdacc92777e40/models/convnext.py#L119 # noqa
+class LayerNorm2d(nn.Module):
+
+ def __init__(self, num_channels: int, eps: float = 1e-6) -> None:
+ super().__init__()
+ self.weight = nn.Parameter(torch.ones(num_channels))
+ self.bias = nn.Parameter(torch.zeros(num_channels))
+ self.eps = eps
+
+ def forward(self, x: torch.Tensor) -> torch.Tensor:
+ u = x.mean(1, keepdim=True)
+ s = (x - u).pow(2).mean(1, keepdim=True)
+ x = (x - u) / torch.sqrt(s + self.eps)
+ x = self.weight[:, None, None] * x + self.bias[:, None, None]
+ return x
diff --git a/projects/sam_inference_demo/sam/modeling/mask_decoder.py b/projects/sam_inference_demo/sam/modeling/mask_decoder.py
new file mode 100644
index 00000000000..9ad616b5899
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/mask_decoder.py
@@ -0,0 +1,196 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# Borrowed from https://github.com/facebookresearch/segment-anything
+
+from typing import List, Tuple
+
+import torch
+from torch import Tensor, nn
+from torch.nn import functional as F
+
+from mmseg.registry import MODELS
+from .common import LayerNorm2d
+
+
+@MODELS.register_module()
+class MaskDecoder(nn.Module):
+
+ def __init__(
+ self,
+ *,
+ transformer_dim: int,
+ transformer: dict,
+ num_multimask_outputs: int = 3,
+ act_cfg: dict = dict(type='GELU'),
+ iou_head_depth: int = 3,
+ iou_head_hidden_dim: int = 256,
+ ) -> None:
+ """Predicts masks given an image and prompt embeddings, using a
+ tranformer architecture.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ transformer_dim (int): the channel dimension of the transformer
+ transformer (nn.Module): the transformer used to predict masks
+ num_multimask_outputs (int): the number of masks to predict
+ when disambiguating masks
+ activation (nn.Module): the type of activation to use when
+ upscaling masks
+ iou_head_depth (int): the depth of the MLP used to predict
+ mask quality
+ iou_head_hidden_dim (int): the hidden dimension of the MLP
+ used to predict mask quality
+ """
+ super().__init__()
+ self.transformer_dim = transformer_dim
+ self.transformer = MODELS.build(transformer)
+
+ self.num_multimask_outputs = num_multimask_outputs
+
+ self.iou_token = nn.Embedding(1, transformer_dim)
+ self.num_mask_tokens = num_multimask_outputs + 1
+ self.mask_tokens = nn.Embedding(self.num_mask_tokens, transformer_dim)
+
+ activation = MODELS.build(act_cfg)
+ self.output_upscaling = nn.Sequential(
+ nn.ConvTranspose2d(
+ transformer_dim, transformer_dim // 4, kernel_size=2,
+ stride=2),
+ LayerNorm2d(transformer_dim // 4),
+ activation,
+ nn.ConvTranspose2d(
+ transformer_dim // 4,
+ transformer_dim // 8,
+ kernel_size=2,
+ stride=2),
+ activation,
+ )
+ self.output_hypernetworks_mlps = nn.ModuleList([
+ MLP(transformer_dim, transformer_dim, transformer_dim // 8, 3)
+ for i in range(self.num_mask_tokens)
+ ])
+
+ self.iou_prediction_head = MLP(transformer_dim, iou_head_hidden_dim,
+ self.num_mask_tokens, iou_head_depth)
+
+ def forward(
+ self,
+ image_embeddings: Tensor,
+ image_pe: Tensor,
+ sparse_prompt_embeddings: Tensor,
+ dense_prompt_embeddings: Tensor,
+ multimask_output: bool,
+ ) -> Tuple[Tensor, Tensor]:
+ """Predict masks given image and prompt embeddings.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ image_embeddings (Tensor): the embeddings from the image encoder
+ image_pe (Tensor): positional encoding with the shape of
+ image_embeddings
+ sparse_prompt_embeddings (Tensor): the embeddings of
+ the points and boxes
+ dense_prompt_embeddings (Tensor): the embeddings of the mask inputs
+ multimask_output (bool): Whether to return multiple masks or a single
+ mask.
+
+ Returns:
+ Tensor: batched predicted masks
+ Tensor: batched predictions of mask quality
+ """
+ masks, iou_pred = self.predict_masks(
+ image_embeddings=image_embeddings,
+ image_pe=image_pe,
+ sparse_prompt_embeddings=sparse_prompt_embeddings,
+ dense_prompt_embeddings=dense_prompt_embeddings,
+ )
+
+ # Select the correct mask or masks for output
+ if multimask_output:
+ mask_slice = slice(1, None)
+ else:
+ mask_slice = slice(0, 1)
+ masks = masks[:, mask_slice, :, :]
+ iou_pred = iou_pred[:, mask_slice]
+
+ # Prepare output
+ return masks, iou_pred
+
+ def predict_masks(
+ self,
+ image_embeddings: Tensor,
+ image_pe: Tensor,
+ sparse_prompt_embeddings: Tensor,
+ dense_prompt_embeddings: Tensor,
+ ) -> Tuple[Tensor, Tensor]:
+ """Predicts masks.
+
+ See 'forward' for more details.
+ """
+ # Concatenate output tokens
+ output_tokens = torch.cat(
+ [self.iou_token.weight, self.mask_tokens.weight], dim=0)
+ output_tokens = output_tokens.unsqueeze(0).expand(
+ sparse_prompt_embeddings.size(0), -1, -1)
+ tokens = torch.cat((output_tokens, sparse_prompt_embeddings), dim=1)
+
+ # Expand per-image data in batch direction to be per-mask
+ src = torch.repeat_interleave(image_embeddings, tokens.shape[0], dim=0)
+ src = src + dense_prompt_embeddings
+ pos_src = torch.repeat_interleave(image_pe, tokens.shape[0], dim=0)
+ b, c, h, w = src.shape
+
+ # Run the transformer
+ hs, src = self.transformer(src, pos_src, tokens)
+ iou_token_out = hs[:, 0, :]
+ mask_tokens_out = hs[:, 1:(1 + self.num_mask_tokens), :]
+
+ # Upscale mask embeddings and predict masks using the mask tokens
+ src = src.transpose(1, 2).view(b, c, h, w)
+ upscaled_embedding = self.output_upscaling(src)
+ hyper_in_list: List[Tensor] = []
+ for i in range(self.num_mask_tokens):
+ hyper_in_list.append(self.output_hypernetworks_mlps[i](
+ mask_tokens_out[:, i, :]))
+ hyper_in = torch.stack(hyper_in_list, dim=1)
+ b, c, h, w = upscaled_embedding.shape
+ masks = (hyper_in @ upscaled_embedding.view(b, c, h * w)).view(
+ b, -1, h, w)
+
+ # Generate mask quality predictions
+ iou_pred = self.iou_prediction_head(iou_token_out)
+
+ return masks, iou_pred
+
+
+# Lightly adapted from
+# https://github.com/facebookresearch/MaskFormer/blob/main/mask_former/modeling/transformer/transformer_predictor.py # noqa
+class MLP(nn.Module):
+
+ def __init__(
+ self,
+ input_dim: int,
+ hidden_dim: int,
+ output_dim: int,
+ num_layers: int,
+ sigmoid_output: bool = False,
+ ) -> None:
+ super().__init__()
+ self.num_layers = num_layers
+ h = [hidden_dim] * (num_layers - 1)
+ self.layers = nn.ModuleList(
+ nn.Linear(n, k) for n, k in zip([input_dim] + h, h + [output_dim]))
+ self.sigmoid_output = sigmoid_output
+
+ def forward(self, x):
+ for i, layer in enumerate(self.layers):
+ x = F.relu(layer(x)) if i < self.num_layers - 1 else layer(x)
+ if self.sigmoid_output:
+ x = F.sigmoid(x)
+ return x
diff --git a/projects/sam_inference_demo/sam/modeling/prompt_encoder.py b/projects/sam_inference_demo/sam/modeling/prompt_encoder.py
new file mode 100644
index 00000000000..6b7c0833871
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/prompt_encoder.py
@@ -0,0 +1,227 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# Borrowed from https://github.com/facebookresearch/segment-anything
+
+from typing import Any, Optional, Tuple, Type
+
+import numpy as np
+import torch
+from torch import nn
+
+from mmseg.registry import MODELS
+from .common import LayerNorm2d
+
+
+@MODELS.register_module()
+class PromptEncoder(nn.Module):
+
+ def __init__(
+ self,
+ embed_dim: int,
+ image_embedding_size: Tuple[int, int],
+ input_image_size: Tuple[int, int],
+ mask_in_chans: int,
+ activation: Type[nn.Module] = nn.GELU,
+ ) -> None:
+ """Encodes prompts for input to SAM's mask decoder.
+
+ Arguments:
+ embed_dim (int): The prompts' embedding dimension
+ image_embedding_size (tuple(int, int)): The spatial size of the
+ image embedding, as (H, W).
+ input_image_size (int): The padded size of the image as input
+ to the image encoder, as (H, W).
+ mask_in_chans (int): The number of hidden channels used for
+ encoding input masks.
+ activation (nn.Module): The activation to use when encoding
+ input masks.
+ """
+ super().__init__()
+ self.embed_dim = embed_dim
+ self.input_image_size = input_image_size
+ self.image_embedding_size = image_embedding_size
+ self.pe_layer = PositionEmbeddingRandom(embed_dim // 2)
+
+ self.num_point_embeddings: int = 4 # pos/neg point + 2 box corners
+ point_embeddings = [
+ nn.Embedding(1, embed_dim)
+ for i in range(self.num_point_embeddings)
+ ]
+ self.point_embeddings = nn.ModuleList(point_embeddings)
+ self.not_a_point_embed = nn.Embedding(1, embed_dim)
+
+ self.mask_input_size = (4 * image_embedding_size[0],
+ 4 * image_embedding_size[1])
+ self.mask_downscaling = nn.Sequential(
+ nn.Conv2d(1, mask_in_chans // 4, kernel_size=2, stride=2),
+ LayerNorm2d(mask_in_chans // 4),
+ activation(),
+ nn.Conv2d(
+ mask_in_chans // 4, mask_in_chans, kernel_size=2, stride=2),
+ LayerNorm2d(mask_in_chans),
+ activation(),
+ nn.Conv2d(mask_in_chans, embed_dim, kernel_size=1),
+ )
+ self.no_mask_embed = nn.Embedding(1, embed_dim)
+
+ def get_dense_pe(self) -> torch.Tensor:
+ """Returns the positional encoding used to encode point prompts,
+ applied to a dense set of points the shape of the image encoding.
+
+ Returns:
+ torch.Tensor: Positional encoding with shape
+ 1x(embed_dim)x(embedding_h)x(embedding_w)
+ """
+ return self.pe_layer(self.image_embedding_size).unsqueeze(0)
+
+ def _embed_points(
+ self,
+ points: torch.Tensor,
+ labels: torch.Tensor,
+ pad: bool,
+ ) -> torch.Tensor:
+ """Embeds point prompts."""
+ points = points + 0.5 # Shift to center of pixel
+ if pad:
+ padding_point = torch.zeros((points.shape[0], 1, 2),
+ device=points.device)
+ padding_label = -torch.ones(
+ (labels.shape[0], 1), device=labels.device)
+ points = torch.cat([points, padding_point], dim=1)
+ labels = torch.cat([labels, padding_label], dim=1)
+ point_embedding = self.pe_layer.forward_with_coords(
+ points, self.input_image_size)
+ point_embedding[labels == -1] = 0.0
+ point_embedding[labels == -1] += self.not_a_point_embed.weight
+ point_embedding[labels == 0] += self.point_embeddings[0].weight
+ point_embedding[labels == 1] += self.point_embeddings[1].weight
+ return point_embedding
+
+ def _embed_boxes(self, boxes: torch.Tensor) -> torch.Tensor:
+ """Embeds box prompts."""
+ boxes = boxes + 0.5 # Shift to center of pixel
+ coords = boxes.reshape(-1, 2, 2)
+ corner_embedding = self.pe_layer.forward_with_coords(
+ coords, self.input_image_size)
+ corner_embedding[:, 0, :] += self.point_embeddings[2].weight
+ corner_embedding[:, 1, :] += self.point_embeddings[3].weight
+ return corner_embedding
+
+ def _embed_masks(self, masks: torch.Tensor) -> torch.Tensor:
+ """Embeds mask inputs."""
+ mask_embedding = self.mask_downscaling(masks)
+ return mask_embedding
+
+ def _get_batch_size(
+ self,
+ points: Optional[Tuple[torch.Tensor, torch.Tensor]],
+ boxes: Optional[torch.Tensor],
+ masks: Optional[torch.Tensor],
+ ) -> int:
+ """Gets the batch size of the output given the batch size of the input
+ prompts."""
+ if points is not None:
+ return points[0].shape[0]
+ elif boxes is not None:
+ return boxes.shape[0]
+ elif masks is not None:
+ return masks.shape[0]
+ else:
+ return 1
+
+ def _get_device(self) -> torch.device:
+ return self.point_embeddings[0].weight.device
+
+ def forward(
+ self,
+ points: Optional[Tuple[torch.Tensor, torch.Tensor]],
+ boxes: Optional[torch.Tensor],
+ masks: Optional[torch.Tensor],
+ ) -> Tuple[torch.Tensor, torch.Tensor]:
+ """Embeds different types of prompts, returning both sparse and dense
+ embeddings.
+
+ Arguments:
+ points (tuple(torch.Tensor, torch.Tensor) or none): point coordinates
+ and labels to embed.
+ boxes (torch.Tensor or none): boxes to embed
+ masks (torch.Tensor or none): masks to embed
+
+ Returns:
+ torch.Tensor: sparse embeddings for the points and boxes, with shape
+ BxNx(embed_dim), where N is determined by the number of input points
+ and boxes.
+ torch.Tensor: dense embeddings for the masks, in the shape
+ Bx(embed_dim)x(embed_H)x(embed_W)
+ """ # noqa
+ bs = self._get_batch_size(points, boxes, masks)
+ sparse_embeddings = torch.empty((bs, 0, self.embed_dim),
+ device=self._get_device())
+ if points is not None:
+ coords, labels = points
+ point_embeddings = self._embed_points(
+ coords, labels, pad=(boxes is None))
+ sparse_embeddings = torch.cat(
+ [sparse_embeddings, point_embeddings], dim=1)
+ if boxes is not None:
+ box_embeddings = self._embed_boxes(boxes)
+ sparse_embeddings = torch.cat([sparse_embeddings, box_embeddings],
+ dim=1)
+
+ if masks is not None:
+ dense_embeddings = self._embed_masks(masks)
+ else:
+ dense_embeddings = self.no_mask_embed.weight.reshape(
+ 1, -1, 1, 1).expand(bs, -1, self.image_embedding_size[0],
+ self.image_embedding_size[1])
+
+ return sparse_embeddings, dense_embeddings
+
+
+class PositionEmbeddingRandom(nn.Module):
+ """Positional encoding using random spatial frequencies."""
+
+ def __init__(self,
+ num_pos_feats: int = 64,
+ scale: Optional[float] = None) -> None:
+ super().__init__()
+ if scale is None or scale <= 0.0:
+ scale = 1.0
+ self.register_buffer(
+ 'positional_encoding_gaussian_matrix',
+ scale * torch.randn((2, num_pos_feats)),
+ )
+
+ def _pe_encoding(self, coords: torch.Tensor) -> torch.Tensor:
+ """Positionally encode points that are normalized to [0,1]."""
+ # assuming coords are in [0, 1]^2 square and have d_1 x ... x d_n x 2 shape # noqa
+ coords = 2 * coords - 1
+ coords = coords @ self.positional_encoding_gaussian_matrix
+ coords = 2 * np.pi * coords
+ # outputs d_1 x ... x d_n x C shape
+ return torch.cat([torch.sin(coords), torch.cos(coords)], dim=-1)
+
+ def forward(self, size: Tuple[int, int]) -> torch.Tensor:
+ """Generate positional encoding for a grid of the specified size."""
+ h, w = size
+ device: Any = self.positional_encoding_gaussian_matrix.device
+ grid = torch.ones((h, w), device=device, dtype=torch.float32)
+ y_embed = grid.cumsum(dim=0) - 0.5
+ x_embed = grid.cumsum(dim=1) - 0.5
+ y_embed = y_embed / h
+ x_embed = x_embed / w
+
+ pe = self._pe_encoding(torch.stack([x_embed, y_embed], dim=-1))
+ return pe.permute(2, 0, 1) # C x H x W
+
+ def forward_with_coords(self, coords_input: torch.Tensor,
+ image_size: Tuple[int, int]) -> torch.Tensor:
+ """Positionally encode points that are not normalized to [0,1]."""
+ coords = coords_input.clone()
+ coords[:, :, 0] = coords[:, :, 0] / image_size[1]
+ coords[:, :, 1] = coords[:, :, 1] / image_size[0]
+ return self._pe_encoding(coords.to(torch.float)) # B x N x C
diff --git a/projects/sam_inference_demo/sam/modeling/sam.py b/projects/sam_inference_demo/sam/modeling/sam.py
new file mode 100644
index 00000000000..c61c1eca4e7
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/sam.py
@@ -0,0 +1,188 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# Borrowed from https://github.com/facebookresearch/segment-anything
+
+from typing import Any, Dict, List, Tuple
+
+import torch
+from torch import nn
+from torch.nn import functional as F
+
+from mmseg.registry import MODELS
+from .mask_decoder import MaskDecoder
+from .prompt_encoder import PromptEncoder
+
+
+@MODELS.register_module()
+class SAM(nn.Module):
+ mask_threshold: float = 0.0
+ image_format: str = 'RGB'
+
+ def __init__(
+ self,
+ image_encoder_cfg: dict,
+ prompt_encoder_cfg: dict,
+ mask_decoder_cfg: dict,
+ pixel_mean: List[float] = [123.675, 116.28, 103.53],
+ pixel_std: List[float] = [58.395, 57.12, 57.375],
+ ) -> None:
+ """SAM predicts object masks from an image and input prompts. Borrowed
+ from https://github.com/facebookresearch/segment-anything.
+
+ Arguments:
+ image_encoder (ViTSAM): The backbone used to encode the
+ image into image embeddings that allow for efficient mask
+ prediction.
+ prompt_encoder (PromptEncoder): Encodes various types of input
+ prompts.
+ mask_decoder (MaskDecoder): Predicts masks from the image embeddings
+ and encoded prompts.
+ pixel_mean (list(float)): Mean values for normalizing pixels in the
+ input image.
+ pixel_std (list(float)): Std values for normalizing pixels in the
+ input image.
+ """
+ super().__init__()
+ self.image_encoder = MODELS.build(image_encoder_cfg)
+ self.prompt_encoder: PromptEncoder = MODELS.build(prompt_encoder_cfg)
+ self.mask_decoder: MaskDecoder = MODELS.build(mask_decoder_cfg)
+ self.register_buffer('pixel_mean',
+ torch.Tensor(pixel_mean).view(-1, 1, 1), False)
+ self.register_buffer('pixel_std',
+ torch.Tensor(pixel_std).view(-1, 1, 1), False)
+
+ @property
+ def device(self) -> Any:
+ return self.pixel_mean.device
+
+ @torch.no_grad()
+ def forward(
+ self,
+ batched_input: List[Dict[str, Any]],
+ multimask_output: bool,
+ ) -> List[Dict[str, torch.Tensor]]:
+ """Predicts masks end-to-end from provided images and prompts. If
+ prompts are not known in advance, using SamPredictor is recommended
+ over calling the model directly.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ batched_input (list(dict)): A list over input images, each a
+ dictionary with the following keys. A prompt key can be
+ excluded if it is not present.
+ 'image': The image as a torch tensor in 3xHxW format,
+ already transformed for input to the model.
+ 'original_size': (tuple(int, int)) The original size of
+ the image before transformation, as (H, W).
+ 'point_coords': (torch.Tensor) Batched point prompts for
+ this image, with shape BxNx2. Already transformed to the
+ input frame of the model.
+ 'point_labels': (torch.Tensor) Batched labels for point prompts,
+ with shape BxN.
+ 'boxes': (torch.Tensor) Batched box inputs, with shape Bx4.
+ Already transformed to the input frame of the model.
+ 'mask_inputs': (torch.Tensor) Batched mask inputs to the model,
+ in the form Bx1xHxW.
+ multimask_output (bool): Whether the model should predict multiple
+ disambiguating masks, or return a single mask.
+
+ Returns:
+ (list(dict)): A list over input images, where each element is
+ as dictionary with the following keys.
+ 'masks': (torch.Tensor) Batched binary mask predictions,
+ with shape BxCxHxW, where B is the number of input prompts,
+ C is determiend by multimask_output, and (H, W) is the
+ original size of the image.
+ 'iou_predictions': (torch.Tensor) The model's predictions
+ of mask quality, in shape BxC.
+ 'low_res_logits': (torch.Tensor) Low resolution logits with
+ shape BxCxHxW, where H=W=256. Can be passed as mask input
+ to subsequent iterations of prediction.
+ """
+ input_images = torch.stack(
+ [self.preprocess(x['image']) for x in batched_input], dim=0)
+ image_embeddings = self.image_encoder(input_images)
+
+ outputs = []
+ for image_record, curr_embedding in zip(batched_input,
+ image_embeddings):
+ if 'point_coords' in image_record:
+ points = (image_record['point_coords'],
+ image_record['point_labels'])
+ else:
+ points = None
+ sparse_embeddings, dense_embeddings = self.prompt_encoder(
+ points=points,
+ boxes=image_record.get('boxes', None),
+ masks=image_record.get('mask_inputs', None),
+ )
+ low_res_masks, iou_predictions = self.mask_decoder(
+ image_embeddings=curr_embedding.unsqueeze(0),
+ image_pe=self.prompt_encoder.get_dense_pe(),
+ sparse_prompt_embeddings=sparse_embeddings,
+ dense_prompt_embeddings=dense_embeddings,
+ multimask_output=multimask_output,
+ )
+ masks = self.postprocess_masks(
+ low_res_masks,
+ input_size=image_record['image'].shape[-2:],
+ original_size=image_record['original_size'],
+ )
+ masks = masks > self.mask_threshold
+ outputs.append({
+ 'masks': masks,
+ 'iou_predictions': iou_predictions,
+ 'low_res_logits': low_res_masks,
+ })
+ return outputs
+
+ def postprocess_masks(
+ self,
+ masks: torch.Tensor,
+ input_size: Tuple[int, ...],
+ original_size: Tuple[int, ...],
+ ) -> torch.Tensor:
+ """Remove padding and upscale masks to the original image size.
+
+ Borrowed from https://github.com/facebookresearch/segment-anything
+
+ Arguments:
+ masks (torch.Tensor): Batched masks from the mask_decoder,
+ in BxCxHxW format.
+ input_size (tuple(int, int)): The size of the image input to the
+ model, in (H, W) format. Used to remove padding.
+ original_size (tuple(int, int)): The original size of the image
+ before resizing for input to the model, in (H, W) format.
+
+ Returns:
+ (torch.Tensor): Batched masks in BxCxHxW format, where (H, W)
+ is given by original_size.
+ """
+ masks = F.interpolate(
+ masks,
+ self.image_encoder.img_size,
+ mode='bilinear',
+ align_corners=False,
+ )
+ masks = masks[..., :input_size[0], :input_size[1]]
+ masks = F.interpolate(
+ masks, original_size, mode='bilinear', align_corners=False)
+ return masks
+
+ def preprocess(self, x: torch.Tensor) -> torch.Tensor:
+ """Normalize pixel values and pad to a square input."""
+ # Normalize colors
+ x = (x - self.pixel_mean) / self.pixel_std
+
+ # Pad
+ h, w = x.shape[-2:]
+ img_size = max(self.image_encoder.img_size)
+ padh = img_size - h
+ padw = img_size - w
+ x = F.pad(x, (0, padw, 0, padh))
+ return x
diff --git a/projects/sam_inference_demo/sam/modeling/transformer.py b/projects/sam_inference_demo/sam/modeling/transformer.py
new file mode 100644
index 00000000000..c56f602487d
--- /dev/null
+++ b/projects/sam_inference_demo/sam/modeling/transformer.py
@@ -0,0 +1,241 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+import math
+from typing import Tuple, Type
+
+import torch
+from torch import Tensor, nn
+
+from mmseg.registry import MODELS
+from .common import MLPBlock
+
+
+@MODELS.register_module()
+class TwoWayTransformer(nn.Module):
+
+ def __init__(
+ self,
+ depth: int,
+ embedding_dim: int,
+ num_heads: int,
+ mlp_dim: int,
+ activation: Type[nn.Module] = nn.ReLU,
+ attention_downsample_rate: int = 2,
+ ) -> None:
+ """A transformer decoder that attends to an input image using queries
+ whose positional embedding is supplied.
+
+ Args:
+ depth (int): number of layers in the transformer
+ embedding_dim (int): the channel dimension for the input embeddings
+ num_heads (int): the number of heads for multihead attention. Must
+ divide embedding_dim
+ mlp_dim (int): the channel dimension internal to the MLP block
+ activation (nn.Module): the activation to use in the MLP block
+ """
+ super().__init__()
+ self.depth = depth
+ self.embedding_dim = embedding_dim
+ self.num_heads = num_heads
+ self.mlp_dim = mlp_dim
+ self.layers = nn.ModuleList()
+
+ for i in range(depth):
+ self.layers.append(
+ TwoWayAttentionBlock(
+ embedding_dim=embedding_dim,
+ num_heads=num_heads,
+ mlp_dim=mlp_dim,
+ activation=activation,
+ attention_downsample_rate=attention_downsample_rate,
+ skip_first_layer_pe=(i == 0),
+ ))
+
+ self.final_attn_token_to_image = Attention(
+ embedding_dim,
+ num_heads,
+ downsample_rate=attention_downsample_rate)
+ self.norm_final_attn = nn.LayerNorm(embedding_dim)
+
+ def forward(
+ self,
+ image_embedding: Tensor,
+ image_pe: Tensor,
+ point_embedding: Tensor,
+ ) -> Tuple[Tensor, Tensor]:
+ """
+ Args:
+ image_embedding (torch.Tensor): image to attend to. Should be shape
+ B x embedding_dim x h x w for any h and w.
+ image_pe (torch.Tensor): the positional encoding to add to the image. Must
+ have the same shape as image_embedding.
+ point_embedding (torch.Tensor): the embedding to add to the query points.
+ Must have shape B x N_points x embedding_dim for any N_points.
+
+ Returns:
+ torch.Tensor: the processed point_embedding
+ torch.Tensor: the processed image_embedding
+ """ # noqa E501
+ # BxCxHxW -> BxHWxC == B x N_image_tokens x C
+ bs, c, h, w = image_embedding.shape
+ image_embedding = image_embedding.flatten(2).permute(0, 2, 1)
+ image_pe = image_pe.flatten(2).permute(0, 2, 1)
+
+ # Prepare queries
+ queries = point_embedding
+ keys = image_embedding
+
+ # Apply transformer blocks and final layernorm
+ for layer in self.layers:
+ queries, keys = layer(
+ queries=queries,
+ keys=keys,
+ query_pe=point_embedding,
+ key_pe=image_pe,
+ )
+
+ # Apply the final attenion layer from the points to the image
+ q = queries + point_embedding
+ k = keys + image_pe
+ attn_out = self.final_attn_token_to_image(q=q, k=k, v=keys)
+ queries = queries + attn_out
+ queries = self.norm_final_attn(queries)
+
+ return queries, keys
+
+
+class TwoWayAttentionBlock(nn.Module):
+
+ def __init__(
+ self,
+ embedding_dim: int,
+ num_heads: int,
+ mlp_dim: int = 2048,
+ activation: Type[nn.Module] = nn.ReLU,
+ attention_downsample_rate: int = 2,
+ skip_first_layer_pe: bool = False,
+ ) -> None:
+ """A transformer block with four layers: (1) self-attention of sparse
+ inputs, (2) cross attention of sparse inputs to dense inputs, (3) mlp
+ block on sparse inputs, and (4) cross attention of dense inputs to
+ sparse inputs.
+
+ Arguments:
+ embedding_dim (int): the channel dimension of the embeddings
+ num_heads (int): the number of heads in the attention layers
+ mlp_dim (int): the hidden dimension of the mlp block
+ activation (nn.Module): the activation of the mlp block
+ skip_first_layer_pe (bool): skip the PE on the first layer
+ """
+ super().__init__()
+ self.self_attn = Attention(embedding_dim, num_heads)
+ self.norm1 = nn.LayerNorm(embedding_dim)
+
+ self.cross_attn_token_to_image = Attention(
+ embedding_dim,
+ num_heads,
+ downsample_rate=attention_downsample_rate)
+ self.norm2 = nn.LayerNorm(embedding_dim)
+
+ self.mlp = MLPBlock(embedding_dim, mlp_dim, activation)
+ self.norm3 = nn.LayerNorm(embedding_dim)
+
+ self.norm4 = nn.LayerNorm(embedding_dim)
+ self.cross_attn_image_to_token = Attention(
+ embedding_dim,
+ num_heads,
+ downsample_rate=attention_downsample_rate)
+
+ self.skip_first_layer_pe = skip_first_layer_pe
+
+ def forward(self, queries: Tensor, keys: Tensor, query_pe: Tensor,
+ key_pe: Tensor) -> Tuple[Tensor, Tensor]:
+ # Self attention block
+ if self.skip_first_layer_pe:
+ queries = self.self_attn(q=queries, k=queries, v=queries)
+ else:
+ q = queries + query_pe
+ attn_out = self.self_attn(q=q, k=q, v=queries)
+ queries = queries + attn_out
+ queries = self.norm1(queries)
+
+ # Cross attention block, tokens attending to image embedding
+ q = queries + query_pe
+ k = keys + key_pe
+ attn_out = self.cross_attn_token_to_image(q=q, k=k, v=keys)
+ queries = queries + attn_out
+ queries = self.norm2(queries)
+
+ # MLP block
+ mlp_out = self.mlp(queries)
+ queries = queries + mlp_out
+ queries = self.norm3(queries)
+
+ # Cross attention block, image embedding attending to tokens
+ q = queries + query_pe
+ k = keys + key_pe
+ attn_out = self.cross_attn_image_to_token(q=k, k=q, v=queries)
+ keys = keys + attn_out
+ keys = self.norm4(keys)
+
+ return queries, keys
+
+
+class Attention(nn.Module):
+ """An attention layer that allows for downscaling the size of the embedding
+ after projection to queries, keys, and values."""
+
+ def __init__(
+ self,
+ embedding_dim: int,
+ num_heads: int,
+ downsample_rate: int = 1,
+ ) -> None:
+ super().__init__()
+ self.embedding_dim = embedding_dim
+ self.internal_dim = embedding_dim // downsample_rate
+ self.num_heads = num_heads
+ assert self.internal_dim % num_heads == 0, 'num_heads must divide embedding_dim.' # noqa E501
+
+ self.q_proj = nn.Linear(embedding_dim, self.internal_dim)
+ self.k_proj = nn.Linear(embedding_dim, self.internal_dim)
+ self.v_proj = nn.Linear(embedding_dim, self.internal_dim)
+ self.out_proj = nn.Linear(self.internal_dim, embedding_dim)
+
+ def _separate_heads(self, x: Tensor, num_heads: int) -> Tensor:
+ b, n, c = x.shape
+ x = x.reshape(b, n, num_heads, c // num_heads)
+ return x.transpose(1, 2) # B x N_heads x N_tokens x C_per_head
+
+ def _recombine_heads(self, x: Tensor) -> Tensor:
+ b, n_heads, n_tokens, c_per_head = x.shape
+ x = x.transpose(1, 2)
+ return x.reshape(b, n_tokens, n_heads * c_per_head) # B x N_tokens x C
+
+ def forward(self, q: Tensor, k: Tensor, v: Tensor) -> Tensor:
+ # Input projections
+ q = self.q_proj(q)
+ k = self.k_proj(k)
+ v = self.v_proj(v)
+
+ # Separate into heads
+ q = self._separate_heads(q, self.num_heads)
+ k = self._separate_heads(k, self.num_heads)
+ v = self._separate_heads(v, self.num_heads)
+
+ # Attention
+ _, _, _, c_per_head = q.shape
+ attn = q @ k.permute(0, 1, 3, 2) # B x N_heads x N_tokens x N_tokens
+ attn = attn / math.sqrt(c_per_head)
+ attn = torch.softmax(attn, dim=-1)
+
+ # Get output
+ out = attn @ v
+ out = self._recombine_heads(out)
+ out = self.out_proj(out)
+
+ return out
diff --git a/projects/sam_inference_demo/sam/sam_inferencer.py b/projects/sam_inference_demo/sam/sam_inferencer.py
new file mode 100644
index 00000000000..2da2e959c09
--- /dev/null
+++ b/projects/sam_inference_demo/sam/sam_inferencer.py
@@ -0,0 +1,688 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from typing import Any, Dict, List, Optional, Tuple
+
+import numpy as np
+import torch
+from mmengine.runner.checkpoint import load_checkpoint
+# yapf: disable
+from sam.utils import (MaskData, area_from_rle, batch_iterator,
+ batched_mask_to_box, box_xyxy_to_xywh,
+ build_all_layer_point_grids, calculate_stability_score,
+ coco_encode_rle, generate_crop_boxes,
+ is_box_near_crop_edge, mask_to_rle_pytorch,
+ remove_small_regions, rle_to_mask, uncrop_boxes_xyxy,
+ uncrop_masks, uncrop_points)
+from torchvision.ops.boxes import batched_nms, box_area
+
+from mmseg.registry import MODELS, TRANSFORMS
+
+# yapf: enable
+
+model_zoo = {
+ 'base':
+ 'https://download.openmmlab.com/mmsegmentation/v0.5/sam/sam_vit-base-p16_3rdparty_sa1b-1024x1024_20230413-78a25eed.pth', # noqa
+ 'large':
+ 'https://download.openmmlab.com/mmsegmentation/v0.5/sam/sam_vit-large-p16_3rdparty_sa1b-1024x1024_20230413-940520da.pth', # noqa
+ 'huge':
+ 'https://download.openmmlab.com/mmsegmentation/v0.5/sam/sam_vit-huge-p16_3rdparty_sa1b-1024x1024_20230413-faaf96f6.pth', # noqa
+}
+
+
+class SAMInferencer:
+
+ def __init__(self, arch: str = 'base') -> None:
+ assert arch in ['base', 'large', 'huge']
+ self.model = self.init_model(arch)
+ self.transform = TRANSFORMS.build(
+ dict(
+ type='ResizeLongestSide',
+ target_length=max(self.model.image_encoder.img_size)))
+
+ def set_image(
+ self,
+ image: np.ndarray,
+ image_format: str = 'RGB',
+ ) -> None:
+ """Calculates the image embeddings for the provided image, allowing
+ masks to be predicted with the 'predict' method.
+
+ Arguments:
+ image (np.ndarray): The image for calculating masks. Expects an
+ image in HWC uint8 format, with pixel values in [0, 255].
+ image_format (str): The color format of the image, in ['RGB', 'BGR'].
+ """
+ assert image_format in [
+ 'RGB',
+ 'BGR',
+ ], f"image_format must be in ['RGB', 'BGR'], is {image_format}."
+ if image_format != self.model.image_format:
+ image = image[..., ::-1]
+
+ # Transform the image to the form expected by the model
+ input_image = self.transform.apply_image(image)
+ input_image_torch = torch.as_tensor(input_image, device=self.device)
+ input_image_torch = input_image_torch.permute(
+ 2, 0, 1).contiguous()[None, :, :, :]
+
+ self.set_torch_image(input_image_torch, image.shape[:2])
+
+ @torch.no_grad()
+ def set_torch_image(
+ self,
+ transformed_image: torch.Tensor,
+ original_image_size: Tuple[int, ...],
+ ) -> None:
+ """Calculates the image embeddings for the provided image, allowing
+ masks to be predicted with the 'predict' method. Expects the input
+ image to be already transformed to the format expected by the model.
+
+ Arguments:
+ transformed_image (torch.Tensor): The input image, with shape
+ 1x3xHxW, which has been transformed with ResizeLongestSide.
+ original_image_size (tuple(int, int)): The size of the image
+ before transformation, in (H, W) format.
+ """
+ assert (len(transformed_image.shape) == 4
+ and transformed_image.shape[1] == 3
+ and max(*transformed_image.shape[2:]) == max(
+ self.model.image_encoder.img_size)
+ ), 'set_torch_image input must be BCHW with long side'
+ f' {self.model.image_encoder.img_size}.'
+ self.reset_image()
+
+ self.original_size = original_image_size
+ self.input_size = tuple(transformed_image.shape[-2:])
+ input_image = self.model.preprocess(transformed_image)
+ self.features = self.model.image_encoder(input_image)[0]
+ self.is_image_set = True
+
+ def predict(
+ self,
+ point_coords: Optional[np.ndarray] = None,
+ point_labels: Optional[np.ndarray] = None,
+ box: Optional[np.ndarray] = None,
+ mask_input: Optional[np.ndarray] = None,
+ multimask_output: bool = True,
+ return_logits: bool = False,
+ ) -> Tuple[torch.Tensor, torch.Tensor, torch.Tensor]:
+ """Predict masks for the given input prompts, using the currently set
+ image.
+
+ Arguments:
+ point_coords (np.ndarray or None): A Nx2 array of point prompts to the
+ model. Each point is in (X,Y) in pixels.
+ point_labels (np.ndarray or None): A length N array of labels for the
+ point prompts. 1 indicates a foreground point and 0 indicates a
+ background point.
+ box (np.ndarray or None): A length 4 array given a box prompt to the
+ model, in XYXY format.
+ mask_input (np.ndarray): A low resolution mask input to the model, typically
+ coming from a previous prediction iteration. Has form 1xHxW, where
+ for SAM, H=W=256.
+ multimask_output (bool): If true, the model will return three masks.
+ For ambiguous input prompts (such as a single click), this will often
+ produce better masks than a single prediction. If only a single
+ mask is needed, the model's predicted quality score can be used
+ to select the best mask. For non-ambiguous prompts, such as multiple
+ input prompts, multimask_output=False can give better results.
+ return_logits (bool): If true, returns un-thresholded masks logits
+ instead of a binary mask.
+
+ Returns:
+ (np.ndarray): The output masks in CxHxW format, where C is the
+ number of masks, and (H, W) is the original image size.
+ (np.ndarray): An array of length C containing the model's
+ predictions for the quality of each mask.
+ (np.ndarray): An array of shape CxHxW, where C is the number
+ of masks and H=W=256. These low resolution logits can be passed to
+ a subsequent iteration as mask input.
+ """ # noqa
+ if not self.is_image_set:
+ raise RuntimeError(
+ 'An image must be set with .set_image(...) before mask'
+ 'prediction.')
+
+ # Transform input prompts
+ coords_torch = None
+ labels_torch = None
+ box_torch = None
+ mask_input_torch = None
+
+ if point_coords is not None:
+ assert (
+ point_labels is not None
+ ), 'point_labels must be supplied if point_coords is supplied.'
+ point_coords = self.transform.apply_coords(point_coords,
+ self.original_size)
+ coords_torch = torch.as_tensor(
+ point_coords, dtype=torch.float, device=self.device)
+ labels_torch = torch.as_tensor(
+ point_labels, dtype=torch.int, device=self.device)
+ coords_torch, labels_torch = coords_torch[
+ None, :, :], labels_torch[None, :]
+ if box is not None:
+ box = self.transform.apply_boxes(box, self.original_size)
+ box_torch = torch.as_tensor(
+ box, dtype=torch.float, device=self.device)
+ box_torch = box_torch[None, :]
+ if mask_input is not None:
+ mask_input_torch = torch.as_tensor(
+ mask_input, dtype=torch.float, device=self.device)
+ mask_input_torch = mask_input_torch[None, :, :, :]
+
+ masks, iou_predictions, low_res_masks = self.predict_torch(
+ coords_torch,
+ labels_torch,
+ box_torch,
+ mask_input_torch,
+ multimask_output,
+ return_logits=return_logits,
+ )
+
+ masks = masks[0].detach().cpu().numpy()
+ iou_predictions = iou_predictions[0].detach().cpu().numpy()
+ low_res_masks = low_res_masks[0].detach().cpu().numpy()
+ return masks, iou_predictions, low_res_masks
+
+ @torch.no_grad()
+ def predict_torch(
+ self,
+ point_coords: Optional[torch.Tensor],
+ point_labels: Optional[torch.Tensor],
+ boxes: Optional[torch.Tensor] = None,
+ mask_input: Optional[torch.Tensor] = None,
+ multimask_output: bool = True,
+ return_logits: bool = False,
+ ) -> Tuple[torch.Tensor, torch.Tensor, torch.Tensor]:
+ """Predict masks for the given input prompts, using the currently set
+ image. Input prompts are batched torch tensors and are expected to
+ already be transformed to the input frame using ResizeLongestSide.
+
+ Arguments:
+ point_coords (torch.Tensor or None): A BxNx2 array of point prompts to the
+ model. Each point is in (X,Y) in pixels.
+ point_labels (torch.Tensor or None): A BxN array of labels for the
+ point prompts. 1 indicates a foreground point and 0 indicates a
+ background point.
+ box (np.ndarray or None): A Bx4 array given a box prompt to the
+ model, in XYXY format.
+ mask_input (np.ndarray): A low resolution mask input to the model, typically
+ coming from a previous prediction iteration. Has form Bx1xHxW, where
+ for SAM, H=W=256. Masks returned by a previous iteration of the
+ predict method do not need further transformation.
+ multimask_output (bool): If true, the model will return three masks.
+ For ambiguous input prompts (such as a single click), this will often
+ produce better masks than a single prediction. If only a single
+ mask is needed, the model's predicted quality score can be used
+ to select the best mask. For non-ambiguous prompts, such as multiple
+ input prompts, multimask_output=False can give better results.
+ return_logits (bool): If true, returns un-thresholded masks logits
+ instead of a binary mask.
+
+ Returns:
+ (torch.Tensor): The output masks in BxCxHxW format, where C is the
+ number of masks, and (H, W) is the original image size.
+ (torch.Tensor): An array of shape BxC containing the model's
+ predictions for the quality of each mask.
+ (torch.Tensor): An array of shape BxCxHxW, where C is the number
+ of masks and H=W=256. These low res logits can be passed to
+ a subsequent iteration as mask input.
+ """ # noqa
+ if not self.is_image_set:
+ raise RuntimeError(
+ 'An image must be set with .set_image(...) before mask '
+ 'prediction.')
+
+ if point_coords is not None:
+ points = (point_coords, point_labels)
+ else:
+ points = None
+
+ # Embed prompts
+ sparse_embeddings, dense_embeddings = self.model.prompt_encoder(
+ points=points,
+ boxes=boxes,
+ masks=mask_input,
+ )
+
+ # Predict masks
+ low_res_masks, iou_predictions = self.model.mask_decoder(
+ image_embeddings=self.features,
+ image_pe=self.model.prompt_encoder.get_dense_pe(),
+ sparse_prompt_embeddings=sparse_embeddings,
+ dense_prompt_embeddings=dense_embeddings,
+ multimask_output=multimask_output,
+ )
+
+ # Upscale the masks to the original image resolution
+ masks = self.model.postprocess_masks(low_res_masks, self.input_size,
+ self.original_size)
+
+ if not return_logits:
+ masks = masks > self.model.mask_threshold
+
+ return masks, iou_predictions, low_res_masks
+
+ def get_image_embedding(self) -> torch.Tensor:
+ """Returns the image embeddings for the currently set image, with shape
+ 1xCxHxW, where C is the embedding dimension and (H,W) are the embedding
+ spatial dimension of SAM (typically C=256, H=W=64)."""
+ if not self.is_image_set:
+ raise RuntimeError(
+ 'An image must be set with .set_image(...) to generate an '
+ 'embedding.')
+ assert self.features is not None, 'Features must exist if an image has'
+ ' been set.'
+ return self.features
+
+ @property
+ def device(self) -> torch.device:
+ return self.model.device
+
+ def reset_image(self) -> None:
+ """Resets the currently set image."""
+ self.is_image_set = False
+ self.features = None
+ self.orig_h = None
+ self.orig_w = None
+ self.input_h = None
+ self.input_w = None
+
+ def init_model(self, arch: str):
+ model = MODELS.build(
+ dict(
+ type='SAM',
+ image_encoder_cfg=dict(
+ type='mmpretrain.ViTSAM',
+ arch=arch,
+ img_size=1024,
+ patch_size=16,
+ out_channels=256,
+ use_abs_pos=True,
+ use_rel_pos=True,
+ window_size=14,
+ ),
+ prompt_encoder_cfg=dict(
+ type='PromptEncoder',
+ embed_dim=256,
+ image_embedding_size=(64, 64),
+ input_image_size=(1024, 1024),
+ mask_in_chans=16,
+ ),
+ mask_decoder_cfg=dict(
+ type='MaskDecoder',
+ num_multimask_outputs=3,
+ transformer=dict(
+ type='TwoWayTransformer',
+ depth=2,
+ embedding_dim=256,
+ mlp_dim=2048,
+ num_heads=8,
+ ),
+ transformer_dim=256,
+ iou_head_depth=3,
+ iou_head_hidden_dim=256,
+ )))
+ load_checkpoint(model, model_zoo.get(arch), strict=True)
+ if torch.cuda.is_available():
+ model = model.cuda()
+ return model
+
+
+class SamAutomaticMaskGenerator:
+
+ def __init__(
+ self,
+ arch: str = 'base',
+ points_per_side: Optional[int] = 32,
+ points_per_batch: int = 64,
+ pred_iou_thresh: float = 0.88,
+ stability_score_thresh: float = 0.95,
+ stability_score_offset: float = 1.0,
+ box_nms_thresh: float = 0.7,
+ crop_n_layers: int = 0,
+ crop_nms_thresh: float = 0.7,
+ crop_overlap_ratio: float = 512 / 1500,
+ crop_n_points_downscale_factor: int = 1,
+ point_grids: Optional[List[np.ndarray]] = None,
+ min_mask_region_area: int = 0,
+ output_mode: str = 'binary_mask',
+ ) -> None:
+ """Using a SAM model, generates masks for the entire image. Generates a
+ grid of point prompts over the image, then filters low quality and
+ duplicate masks. The default settings are chosen for SAM with a ViT-H
+ backbone.
+
+ Arguments:
+ arch (str): The SAM model to use for mask prediction.
+ points_per_side (int or None): The number of points to be sampled
+ along one side of the image. The total number of points is
+ points_per_side**2. If None, 'point_grids' must provide explicit
+ point sampling.
+ points_per_batch (int): Sets the number of points run simultaneously
+ by the model. Higher numbers may be faster but use more GPU memory.
+ pred_iou_thresh (float): A filtering threshold in [0,1], using the
+ model's predicted mask quality.
+ stability_score_thresh (float): A filtering threshold in [0,1], using
+ the stability of the mask under changes to the cutoff used to binarize
+ the model's mask predictions.
+ stability_score_offset (float): The amount to shift the cutoff when
+ calculated the stability score.
+ box_nms_thresh (float): The box IoU cutoff used by non-maximal
+ suppression to filter duplicate masks.
+ crops_n_layers (int): If >0, mask prediction will be run again on
+ crops of the image. Sets the number of layers to run, where each
+ layer has 2**i_layer number of image crops.
+ crops_nms_thresh (float): The box IoU cutoff used by non-maximal
+ suppression to filter duplicate masks between different crops.
+ crop_overlap_ratio (float): Sets the degree to which crops overlap.
+ In the first crop layer, crops will overlap by this fraction of
+ the image length. Later layers with more crops scale down this overlap.
+ crop_n_points_downscale_factor (int): The number of points-per-side
+ sampled in layer n is scaled down by crop_n_points_downscale_factor**n.
+ point_grids (list(np.ndarray) or None): A list over explicit grids
+ of points used for sampling, normalized to [0,1]. The nth grid in the
+ list is used in the nth crop layer. Exclusive with points_per_side.
+ min_mask_region_area (int): If >0, postprocessing will be applied
+ to remove disconnected regions and holes in masks with area smaller
+ than min_mask_region_area. Requires opencv.
+ output_mode (str): The form masks are returned in. Can be 'binary_mask',
+ 'uncompressed_rle', or 'coco_rle'. 'coco_rle' requires pycocotools.
+ For large resolutions, 'binary_mask' may consume large amounts of
+ memory.
+ """ # noqa
+
+ assert (points_per_side is None) != (
+ point_grids is None
+ ), 'Exactly one of points_per_side or point_grid must be provided.'
+ if points_per_side is not None:
+ self.point_grids = build_all_layer_point_grids(
+ points_per_side,
+ crop_n_layers,
+ crop_n_points_downscale_factor,
+ )
+ elif point_grids is not None:
+ self.point_grids = point_grids
+ else:
+ raise ValueError(
+ "Can't have both points_per_side and point_grid be None.")
+
+ assert output_mode in [
+ 'binary_mask',
+ 'uncompressed_rle',
+ 'coco_rle',
+ ], f'Unknown output_mode {output_mode}.'
+ if output_mode == 'coco_rle':
+ from pycocotools import \
+ mask as mask_utils # type: ignore # noqa: F401
+
+ if min_mask_region_area > 0:
+ import cv2 # type: ignore # noqa: F401
+
+ self.predictor = SAMInferencer(arch)
+ self.points_per_batch = points_per_batch
+ self.pred_iou_thresh = pred_iou_thresh
+ self.stability_score_thresh = stability_score_thresh
+ self.stability_score_offset = stability_score_offset
+ self.box_nms_thresh = box_nms_thresh
+ self.crop_n_layers = crop_n_layers
+ self.crop_nms_thresh = crop_nms_thresh
+ self.crop_overlap_ratio = crop_overlap_ratio
+ self.crop_n_points_downscale_factor = crop_n_points_downscale_factor
+ self.min_mask_region_area = min_mask_region_area
+ self.output_mode = output_mode
+
+ @torch.no_grad()
+ def generate(self, image: np.ndarray) -> List[Dict[str, Any]]:
+ """Generates masks for the given image.
+
+ Arguments:
+ image (np.ndarray): The image to generate masks for, in HWC uint8 format.
+
+ Returns:
+ list(dict(str, any)): A list over records for masks. Each record is
+ a dict containing the following keys:
+ segmentation (dict(str, any) or np.ndarray): The mask. If
+ output_mode='binary_mask', is an array of shape HW. Otherwise,
+ is a dictionary containing the RLE.
+ bbox (list(float)): The box around the mask, in XYWH format.
+ area (int): The area in pixels of the mask.
+ predicted_iou (float): The model's own prediction of the mask's
+ quality. This is filtered by the pred_iou_thresh parameter.
+ point_coords (list(list(float))): The point coordinates input
+ to the model to generate this mask.
+ stability_score (float): A measure of the mask's quality. This
+ is filtered on using the stability_score_thresh parameter.
+ crop_box (list(float)): The crop of the image used to generate
+ the mask, given in XYWH format.
+ """ # noqa
+
+ # Generate masks
+ mask_data = self._generate_masks(image)
+
+ # Filter small disconnected regions and holes in masks
+ if self.min_mask_region_area > 0:
+ mask_data = self.postprocess_small_regions(
+ mask_data,
+ self.min_mask_region_area,
+ max(self.box_nms_thresh, self.crop_nms_thresh),
+ )
+
+ # Encode masks
+ if self.output_mode == 'coco_rle':
+ mask_data['segmentations'] = [
+ coco_encode_rle(rle) for rle in mask_data['rles']
+ ]
+ elif self.output_mode == 'binary_mask':
+ mask_data['segmentations'] = [
+ rle_to_mask(rle) for rle in mask_data['rles']
+ ]
+ else:
+ mask_data['segmentations'] = mask_data['rles']
+
+ # Write mask records
+ curr_anns = []
+ for idx in range(len(mask_data['segmentations'])):
+ ann = {
+ 'segmentation':
+ mask_data['segmentations'][idx],
+ 'area':
+ area_from_rle(mask_data['rles'][idx]),
+ 'bbox':
+ box_xyxy_to_xywh(mask_data['boxes'][idx]).tolist(),
+ 'predicted_iou':
+ mask_data['iou_preds'][idx].item(),
+ 'point_coords': [mask_data['points'][idx].tolist()],
+ 'stability_score':
+ mask_data['stability_score'][idx].item(),
+ 'crop_box':
+ box_xyxy_to_xywh(mask_data['crop_boxes'][idx]).tolist(),
+ }
+ curr_anns.append(ann)
+
+ return curr_anns
+
+ def _generate_masks(self, image: np.ndarray) -> MaskData:
+ orig_size = image.shape[:2]
+ crop_boxes, layer_idxs = generate_crop_boxes(orig_size,
+ self.crop_n_layers,
+ self.crop_overlap_ratio)
+
+ # Iterate over image crops
+ data = MaskData()
+ for crop_box, layer_idx in zip(crop_boxes, layer_idxs):
+ crop_data = self._process_crop(image, crop_box, layer_idx,
+ orig_size)
+ data.cat(crop_data)
+
+ # Remove duplicate masks between crops
+ if len(crop_boxes) > 1:
+ # Prefer masks from smaller crops
+ scores = 1 / box_area(data['crop_boxes'])
+ scores = scores.to(data['boxes'].device)
+ keep_by_nms = batched_nms(
+ data['boxes'].float(),
+ scores,
+ torch.zeros(len(data['boxes'])), # categories
+ iou_threshold=self.crop_nms_thresh,
+ )
+ data.filter(keep_by_nms)
+
+ data.to_numpy()
+ return data
+
+ def _process_crop(
+ self,
+ image: np.ndarray,
+ crop_box: List[int],
+ crop_layer_idx: int,
+ orig_size: Tuple[int, ...],
+ ) -> MaskData:
+ # Crop the image and calculate embeddings
+ x0, y0, x1, y1 = crop_box
+ cropped_im = image[y0:y1, x0:x1, :]
+ cropped_im_size = cropped_im.shape[:2]
+ self.predictor.set_image(cropped_im)
+
+ # Get points for this crop
+ points_scale = np.array(cropped_im_size)[None, ::-1]
+ points_for_image = self.point_grids[crop_layer_idx] * points_scale
+
+ # Generate masks for this crop in batches
+ data = MaskData()
+ for (points, ) in batch_iterator(self.points_per_batch,
+ points_for_image):
+ batch_data = self._process_batch(points, cropped_im_size, crop_box,
+ orig_size)
+ data.cat(batch_data)
+ del batch_data
+ self.predictor.reset_image()
+
+ # Remove duplicates within this crop.
+ keep_by_nms = batched_nms(
+ data['boxes'].float(),
+ data['iou_preds'],
+ torch.zeros(len(data['boxes'])), # categories
+ iou_threshold=self.box_nms_thresh,
+ )
+ data.filter(keep_by_nms)
+
+ # Return to the original image frame
+ data['boxes'] = uncrop_boxes_xyxy(data['boxes'], crop_box)
+ data['points'] = uncrop_points(data['points'], crop_box)
+ data['crop_boxes'] = torch.tensor(
+ [crop_box for _ in range(len(data['rles']))])
+
+ return data
+
+ def _process_batch(
+ self,
+ points: np.ndarray,
+ im_size: Tuple[int, ...],
+ crop_box: List[int],
+ orig_size: Tuple[int, ...],
+ ) -> MaskData:
+ orig_h, orig_w = orig_size
+
+ # Run model on this batch
+ transformed_points = self.predictor.transform.apply_coords(
+ points, im_size)
+ in_points = torch.as_tensor(
+ transformed_points, device=self.predictor.device)
+ in_labels = torch.ones(
+ in_points.shape[0], dtype=torch.int, device=in_points.device)
+ masks, iou_preds, _ = self.predictor.predict_torch(
+ in_points[:, None, :],
+ in_labels[:, None],
+ multimask_output=True,
+ return_logits=True,
+ )
+
+ # Serialize predictions and store in MaskData
+ data = MaskData(
+ masks=masks.flatten(0, 1),
+ iou_preds=iou_preds.flatten(0, 1),
+ points=torch.as_tensor(points.repeat(masks.shape[1], axis=0)),
+ )
+ del masks
+
+ # Filter by predicted IoU
+ if self.pred_iou_thresh > 0.0:
+ keep_mask = data['iou_preds'] > self.pred_iou_thresh
+ data.filter(keep_mask)
+
+ # Calculate stability score
+ data['stability_score'] = calculate_stability_score(
+ data['masks'], self.predictor.model.mask_threshold,
+ self.stability_score_offset)
+ if self.stability_score_thresh > 0.0:
+ keep_mask = data['stability_score'] >= self.stability_score_thresh
+ data.filter(keep_mask)
+
+ # Threshold masks and calculate boxes
+ data['masks'] = data['masks'] > self.predictor.model.mask_threshold
+ data['boxes'] = batched_mask_to_box(data['masks'])
+
+ # Filter boxes that touch crop boundaries
+ keep_mask = ~is_box_near_crop_edge(data['boxes'], crop_box,
+ [0, 0, orig_w, orig_h])
+ if not torch.all(keep_mask):
+ data.filter(keep_mask)
+
+ # Compress to RLE
+ data['masks'] = uncrop_masks(data['masks'], crop_box, orig_h, orig_w)
+ data['rles'] = mask_to_rle_pytorch(data['masks'])
+ del data['masks']
+
+ return data
+
+ @staticmethod
+ def postprocess_small_regions(mask_data: MaskData, min_area: int,
+ nms_thresh: float) -> MaskData:
+ """Removes small disconnected regions and holes in masks, then reruns
+ box NMS to remove any new duplicates.
+
+ Edits mask_data in place.
+
+ Requires open-cv as a dependency.
+ """
+ if len(mask_data['rles']) == 0:
+ return mask_data
+
+ # Filter small disconnected regions and holes
+ new_masks = []
+ scores = []
+ for rle in mask_data['rles']:
+ mask = rle_to_mask(rle)
+
+ mask, changed = remove_small_regions(mask, min_area, mode='holes')
+ unchanged = not changed
+ mask, changed = remove_small_regions(
+ mask, min_area, mode='islands')
+ unchanged = unchanged and not changed
+
+ new_masks.append(torch.as_tensor(mask).unsqueeze(0))
+ # Give score=0 to changed masks and score=1 to unchanged masks
+ # so NMS will prefer ones that didn't need postprocessing
+ scores.append(float(unchanged))
+
+ # Recalculate boxes and remove any new duplicates
+ masks = torch.cat(new_masks, dim=0)
+ boxes = batched_mask_to_box(masks)
+ keep_by_nms = batched_nms(
+ boxes.float(),
+ torch.as_tensor(scores),
+ torch.zeros(len(boxes)), # categories
+ iou_threshold=nms_thresh,
+ )
+
+ # Only recalculate RLEs for masks that have changed
+ for i_mask in keep_by_nms:
+ if scores[i_mask] == 0.0:
+ mask_torch = masks[i_mask].unsqueeze(0)
+ mask_data['rles'][i_mask] = mask_to_rle_pytorch(mask_torch)[0]
+ mask_data['boxes'][i_mask] = boxes[
+ i_mask] # update res directly
+ mask_data.filter(keep_by_nms)
+
+ return mask_data
diff --git a/projects/sam_inference_demo/sam/utils/__init__.py b/projects/sam_inference_demo/sam/utils/__init__.py
new file mode 100644
index 00000000000..5d33e33aeef
--- /dev/null
+++ b/projects/sam_inference_demo/sam/utils/__init__.py
@@ -0,0 +1,2 @@
+from .amg import * # noqa: F403 F401
+from .transforms import ResizeLongestSide # noqa: F403 F401
diff --git a/projects/sam_inference_demo/sam/utils/amg.py b/projects/sam_inference_demo/sam/utils/amg.py
new file mode 100644
index 00000000000..3ba359901f7
--- /dev/null
+++ b/projects/sam_inference_demo/sam/utils/amg.py
@@ -0,0 +1,355 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+# https://github.com/facebookresearch/segment-anything
+
+import math
+from copy import deepcopy
+from itertools import product
+from typing import Any, Dict, Generator, ItemsView, List, Tuple
+
+import numpy as np
+import torch
+
+
+class MaskData:
+ """A structure for storing masks and their related data in batched format.
+
+ Implements basic filtering and concatenation.
+ """
+
+ def __init__(self, **kwargs) -> None:
+ for v in kwargs.values():
+ assert isinstance(
+ v, (list, np.ndarray, torch.Tensor)
+ ), 'MaskData only supports list, numpy arrays, and torch tensors.'
+ self._stats = dict(**kwargs)
+
+ def __setitem__(self, key: str, item: Any) -> None:
+ assert isinstance(
+ item, (list, np.ndarray, torch.Tensor)
+ ), 'MaskData only supports list, numpy arrays, and torch tensors.'
+ self._stats[key] = item
+
+ def __delitem__(self, key: str) -> None:
+ del self._stats[key]
+
+ def __getitem__(self, key: str) -> Any:
+ return self._stats[key]
+
+ def items(self) -> ItemsView[str, Any]:
+ return self._stats.items()
+
+ def filter(self, keep: torch.Tensor) -> None:
+ for k, v in self._stats.items():
+ if v is None:
+ self._stats[k] = None
+ elif isinstance(v, torch.Tensor):
+ self._stats[k] = v[torch.as_tensor(keep, device=v.device)]
+ elif isinstance(v, np.ndarray):
+ self._stats[k] = v[keep.detach().cpu().numpy()]
+ elif isinstance(v, list) and keep.dtype == torch.bool:
+ self._stats[k] = [a for i, a in enumerate(v) if keep[i]]
+ elif isinstance(v, list):
+ self._stats[k] = [v[i] for i in keep]
+ else:
+ raise TypeError(
+ f'MaskData key {k} has an unsupported type {type(v)}.')
+
+ def cat(self, new_stats: 'MaskData') -> None:
+ for k, v in new_stats.items():
+ if k not in self._stats or self._stats[k] is None:
+ self._stats[k] = deepcopy(v)
+ elif isinstance(v, torch.Tensor):
+ self._stats[k] = torch.cat([self._stats[k], v], dim=0)
+ elif isinstance(v, np.ndarray):
+ self._stats[k] = np.concatenate([self._stats[k], v], axis=0)
+ elif isinstance(v, list):
+ self._stats[k] = self._stats[k] + deepcopy(v)
+ else:
+ raise TypeError(
+ f'MaskData key {k} has an unsupported type {type(v)}.')
+
+ def to_numpy(self) -> None:
+ for k, v in self._stats.items():
+ if isinstance(v, torch.Tensor):
+ self._stats[k] = v.detach().cpu().numpy()
+
+
+def is_box_near_crop_edge(boxes: torch.Tensor,
+ crop_box: List[int],
+ orig_box: List[int],
+ atol: float = 20.0) -> torch.Tensor:
+ """Filter masks at the edge of a crop, but not at the edge of the original
+ image."""
+ crop_box_torch = torch.as_tensor(
+ crop_box, dtype=torch.float, device=boxes.device)
+ orig_box_torch = torch.as_tensor(
+ orig_box, dtype=torch.float, device=boxes.device)
+ boxes = uncrop_boxes_xyxy(boxes, crop_box).float()
+ near_crop_edge = torch.isclose(
+ boxes, crop_box_torch[None, :], atol=atol, rtol=0)
+ near_image_edge = torch.isclose(
+ boxes, orig_box_torch[None, :], atol=atol, rtol=0)
+ near_crop_edge = torch.logical_and(near_crop_edge, ~near_image_edge)
+ return torch.any(near_crop_edge, dim=1)
+
+
+def box_xyxy_to_xywh(box_xyxy: torch.Tensor) -> torch.Tensor:
+ box_xywh = deepcopy(box_xyxy)
+ box_xywh[2] = box_xywh[2] - box_xywh[0]
+ box_xywh[3] = box_xywh[3] - box_xywh[1]
+ return box_xywh
+
+
+def batch_iterator(batch_size: int, *args) -> Generator[List[Any], None, None]:
+ assert len(args) > 0 and all(
+ len(a) == len(args[0]) for a in
+ args), 'Batched iteration must have inputs of all the same size.'
+ n_batches = len(args[0]) // batch_size + int(
+ len(args[0]) % batch_size != 0)
+ for b in range(n_batches):
+ yield [arg[b * batch_size:(b + 1) * batch_size] for arg in args]
+
+
+def mask_to_rle_pytorch(tensor: torch.Tensor) -> List[Dict[str, Any]]:
+ """Encodes masks to an uncompressed RLE, in the format expected by pycoco
+ tools."""
+ # Put in fortran order and flatten h,w
+ b, h, w = tensor.shape
+ tensor = tensor.permute(0, 2, 1).flatten(1)
+
+ # Compute change indices
+ diff = tensor[:, 1:] ^ tensor[:, :-1]
+ change_indices = diff.nonzero()
+
+ # Encode run length
+ out = []
+ for i in range(b):
+ cur_idxs = change_indices[change_indices[:, 0] == i, 1]
+ cur_idxs = torch.cat([
+ torch.tensor([0], dtype=cur_idxs.dtype, device=cur_idxs.device),
+ cur_idxs + 1,
+ torch.tensor([h * w], dtype=cur_idxs.dtype,
+ device=cur_idxs.device),
+ ])
+ btw_idxs = cur_idxs[1:] - cur_idxs[:-1]
+ counts = [] if tensor[i, 0] == 0 else [0]
+ counts.extend(btw_idxs.detach().cpu().tolist())
+ out.append({'size': [h, w], 'counts': counts})
+ return out
+
+
+def rle_to_mask(rle: Dict[str, Any]) -> np.ndarray:
+ """Compute a binary mask from an uncompressed RLE."""
+ h, w = rle['size']
+ mask = np.empty(h * w, dtype=bool)
+ idx = 0
+ parity = False
+ for count in rle['counts']:
+ mask[idx:idx + count] = parity
+ idx += count
+ parity ^= True
+ mask = mask.reshape(w, h)
+ return mask.transpose() # Put in C order
+
+
+def area_from_rle(rle: Dict[str, Any]) -> int:
+ return sum(rle['counts'][1::2])
+
+
+def calculate_stability_score(masks: torch.Tensor, mask_threshold: float,
+ threshold_offset: float) -> torch.Tensor:
+ """Computes the stability score for a batch of masks.
+
+ The stability score is the IoU between the binary masks obtained by
+ thresholding the predicted mask logits at high and low values.
+ """
+ # One mask is always contained inside the other.
+ # Save memory by preventing unnecessary cast to torch.int64
+ intersections = ((masks > (mask_threshold + threshold_offset)).sum(
+ -1, dtype=torch.int16).sum(-1, dtype=torch.int32))
+ unions = ((masks > (mask_threshold - threshold_offset)).sum(
+ -1, dtype=torch.int16).sum(-1, dtype=torch.int32))
+ return intersections / unions
+
+
+def build_point_grid(n_per_side: int) -> np.ndarray:
+ """Generates a 2D grid of points evenly spaced in [0,1]x[0,1]."""
+ offset = 1 / (2 * n_per_side)
+ points_one_side = np.linspace(offset, 1 - offset, n_per_side)
+ points_x = np.tile(points_one_side[None, :], (n_per_side, 1))
+ points_y = np.tile(points_one_side[:, None], (1, n_per_side))
+ points = np.stack([points_x, points_y], axis=-1).reshape(-1, 2)
+ return points
+
+
+def build_all_layer_point_grids(n_per_side: int, n_layers: int,
+ scale_per_layer: int) -> List[np.ndarray]:
+ """Generates point grids for all crop layers."""
+ points_by_layer = []
+ for i in range(n_layers + 1):
+ n_points = int(n_per_side / (scale_per_layer**i))
+ points_by_layer.append(build_point_grid(n_points))
+ return points_by_layer
+
+
+def generate_crop_boxes(
+ im_size: Tuple[int, ...], n_layers: int,
+ overlap_ratio: float) -> Tuple[List[List[int]], List[int]]:
+ """Generates a list of crop boxes of different sizes.
+
+ Each layer has (2**i)**2 boxes for the ith layer.
+ """
+ crop_boxes, layer_idxs = [], []
+ im_h, im_w = im_size
+ short_side = min(im_h, im_w)
+
+ # Original image
+ crop_boxes.append([0, 0, im_w, im_h])
+ layer_idxs.append(0)
+
+ def crop_len(orig_len, n_crops, overlap):
+ return int(math.ceil((overlap * (n_crops - 1) + orig_len) / n_crops))
+
+ for i_layer in range(n_layers):
+ n_crops_per_side = 2**(i_layer + 1)
+ overlap = int(overlap_ratio * short_side * (2 / n_crops_per_side))
+
+ crop_w = crop_len(im_w, n_crops_per_side, overlap)
+ crop_h = crop_len(im_h, n_crops_per_side, overlap)
+
+ crop_box_x0 = [
+ int((crop_w - overlap) * i) for i in range(n_crops_per_side)
+ ]
+ crop_box_y0 = [
+ int((crop_h - overlap) * i) for i in range(n_crops_per_side)
+ ]
+
+ # Crops in XYWH format
+ for x0, y0 in product(crop_box_x0, crop_box_y0):
+ box = [x0, y0, min(x0 + crop_w, im_w), min(y0 + crop_h, im_h)]
+ crop_boxes.append(box)
+ layer_idxs.append(i_layer + 1)
+
+ return crop_boxes, layer_idxs
+
+
+def uncrop_boxes_xyxy(boxes: torch.Tensor,
+ crop_box: List[int]) -> torch.Tensor:
+ x0, y0, _, _ = crop_box
+ offset = torch.tensor([[x0, y0, x0, y0]], device=boxes.device)
+ # Check if boxes has a channel dimension
+ if len(boxes.shape) == 3:
+ offset = offset.unsqueeze(1)
+ return boxes + offset
+
+
+def uncrop_points(points: torch.Tensor, crop_box: List[int]) -> torch.Tensor:
+ x0, y0, _, _ = crop_box
+ offset = torch.tensor([[x0, y0]], device=points.device)
+ # Check if points has a channel dimension
+ if len(points.shape) == 3:
+ offset = offset.unsqueeze(1)
+ return points + offset
+
+
+def uncrop_masks(masks: torch.Tensor, crop_box: List[int], orig_h: int,
+ orig_w: int) -> torch.Tensor:
+ x0, y0, x1, y1 = crop_box
+ if x0 == 0 and y0 == 0 and x1 == orig_w and y1 == orig_h:
+ return masks
+ # Coordinate transform masks
+ pad_x, pad_y = orig_w - (x1 - x0), orig_h - (y1 - y0)
+ pad = (x0, pad_x - x0, y0, pad_y - y0)
+ return torch.nn.functional.pad(masks, pad, value=0)
+
+
+def remove_small_regions(mask: np.ndarray, area_thresh: float,
+ mode: str) -> Tuple[np.ndarray, bool]:
+ """Removes small disconnected regions and holes in a mask.
+
+ Returns the mask and an indicator of if the mask has been modified.
+ """
+ import cv2 # type: ignore
+
+ assert mode in ['holes', 'islands']
+ correct_holes = mode == 'holes'
+ working_mask = (correct_holes ^ mask).astype(np.uint8)
+ n_labels, regions, stats, _ = cv2.connectedComponentsWithStats(
+ working_mask, 8)
+ sizes = stats[:, -1][1:] # Row 0 is background label
+ small_regions = [i + 1 for i, s in enumerate(sizes) if s < area_thresh]
+ if len(small_regions) == 0:
+ return mask, False
+ fill_labels = [0] + small_regions
+ if not correct_holes:
+ fill_labels = [i for i in range(n_labels) if i not in fill_labels]
+ # If every region is below threshold, keep largest
+ if len(fill_labels) == 0:
+ fill_labels = [int(np.argmax(sizes)) + 1]
+ mask = np.isin(regions, fill_labels)
+ return mask, True
+
+
+def coco_encode_rle(uncompressed_rle: Dict[str, Any]) -> Dict[str, Any]:
+ from pycocotools import mask as mask_utils # type: ignore
+
+ h, w = uncompressed_rle['size']
+ rle = mask_utils.frPyObjects(uncompressed_rle, h, w)
+ rle['counts'] = rle['counts'].decode(
+ 'utf-8') # Necessary to serialize with json
+ return rle
+
+
+def batched_mask_to_box(masks: torch.Tensor) -> torch.Tensor:
+ """Calculates boxes in XYXY format around masks.
+
+ Return [0,0,0,0] for an empty mask. For input shape C1xC2x...xHxW, the
+ output shape is C1xC2x...x4.
+ """
+ # torch.max below raises an error on empty inputs, just skip in this case
+ if torch.numel(masks) == 0:
+ return torch.zeros(*masks.shape[:-2], 4, device=masks.device)
+
+ # Normalize shape to CxHxW
+ shape = masks.shape
+ h, w = shape[-2:]
+ if len(shape) > 2:
+ masks = masks.flatten(0, -3)
+ else:
+ masks = masks.unsqueeze(0)
+
+ # Get top and bottom edges
+ in_height, _ = torch.max(masks, dim=-1)
+ in_height_coords = in_height * torch.arange(
+ h, device=in_height.device)[None, :]
+ bottom_edges, _ = torch.max(in_height_coords, dim=-1)
+ in_height_coords = in_height_coords + h * (~in_height)
+ top_edges, _ = torch.min(in_height_coords, dim=-1)
+
+ # Get left and right edges
+ in_width, _ = torch.max(masks, dim=-2)
+ in_width_coords = in_width * torch.arange(
+ w, device=in_width.device)[None, :]
+ right_edges, _ = torch.max(in_width_coords, dim=-1)
+ in_width_coords = in_width_coords + w * (~in_width)
+ left_edges, _ = torch.min(in_width_coords, dim=-1)
+
+ # If the mask is empty the right edge will be to the left of the left edge.
+ # Replace these boxes with [0, 0, 0, 0]
+ empty_filter = (right_edges < left_edges) | (bottom_edges < top_edges)
+ out = torch.stack([left_edges, top_edges, right_edges, bottom_edges],
+ dim=-1)
+ out = out * (~empty_filter).unsqueeze(-1)
+
+ # Return to original shape
+ if len(shape) > 2:
+ out = out.reshape(*shape[:-2], 4)
+ else:
+ out = out[0]
+
+ return out
diff --git a/projects/sam_inference_demo/sam/utils/transforms.py b/projects/sam_inference_demo/sam/utils/transforms.py
new file mode 100644
index 00000000000..484fd6691cf
--- /dev/null
+++ b/projects/sam_inference_demo/sam/utils/transforms.py
@@ -0,0 +1,110 @@
+# Copyright (c) Meta Platforms, Inc. and affiliates.
+# All rights reserved.
+
+# This source code is licensed under the license found in the
+# LICENSE file in the root directory of this source tree.
+
+from copy import deepcopy
+from typing import Tuple
+
+import numpy as np
+import torch
+from torch.nn import functional as F
+from torchvision.transforms.functional import resize # type: ignore
+from torchvision.transforms.functional import to_pil_image
+
+from mmseg.registry import TRANSFORMS
+
+
+@TRANSFORMS.register_module()
+class ResizeLongestSide:
+ """Resizes images to longest side 'target_length', as well as provides
+ methods for resizing coordinates and boxes.
+
+ Provides methods for transforming both numpy array and batched torch
+ tensors.
+ """
+
+ def __init__(self, target_length: int) -> None:
+ self.target_length = target_length
+
+ def apply_image(self, image: np.ndarray) -> np.ndarray:
+ """Expects a numpy array with shape HxWxC in uint8 format."""
+ target_size = self.get_preprocess_shape(image.shape[0], image.shape[1],
+ self.target_length)
+ return np.array(resize(to_pil_image(image), target_size))
+
+ def apply_coords(self, coords: np.ndarray,
+ original_size: Tuple[int, ...]) -> np.ndarray:
+ """Expects a numpy array of length 2 in the final dimension.
+
+ Requires the original image size in (H, W) format.
+ """
+ old_h, old_w = original_size
+ new_h, new_w = self.get_preprocess_shape(original_size[0],
+ original_size[1],
+ self.target_length)
+ coords = deepcopy(coords).astype(float)
+ coords[..., 0] = coords[..., 0] * (new_w / old_w)
+ coords[..., 1] = coords[..., 1] * (new_h / old_h)
+ return coords
+
+ def apply_boxes(self, boxes: np.ndarray,
+ original_size: Tuple[int, ...]) -> np.ndarray:
+ """Expects a numpy array shape Bx4.
+
+ Requires the original image size in (H, W) format.
+ """
+ boxes = self.apply_coords(boxes.reshape(-1, 2, 2), original_size)
+ return boxes.reshape(-1, 4)
+
+ def apply_image_torch(self, image: torch.Tensor) -> torch.Tensor:
+ """Expects batched images with shape BxCxHxW and float format.
+
+ This transformation may not exactly match apply_image. apply_image is
+ the transformation expected by the model.
+ """
+ # Expects an image in BCHW format. May not exactly match apply_image.
+ target_size = self.get_preprocess_shape(image.shape[0], image.shape[1],
+ self.target_length)
+ return F.interpolate(
+ image,
+ target_size,
+ mode='bilinear',
+ align_corners=False,
+ antialias=True)
+
+ def apply_coords_torch(self, coords: torch.Tensor,
+ original_size: Tuple[int, ...]) -> torch.Tensor:
+ """Expects a torch tensor with length 2 in the last dimension.
+
+ Requires the original image size in (H, W) format.
+ """
+ old_h, old_w = original_size
+ new_h, new_w = self.get_preprocess_shape(original_size[0],
+ original_size[1],
+ self.target_length)
+ coords = deepcopy(coords).to(torch.float)
+ coords[..., 0] = coords[..., 0] * (new_w / old_w)
+ coords[..., 1] = coords[..., 1] * (new_h / old_h)
+ return coords
+
+ def apply_boxes_torch(self, boxes: torch.Tensor,
+ original_size: Tuple[int, ...]) -> torch.Tensor:
+ """Expects a torch tensor with shape Bx4.
+
+ Requires the original image size in (H, W) format.
+ """
+ boxes = self.apply_coords_torch(boxes.reshape(-1, 2, 2), original_size)
+ return boxes.reshape(-1, 4)
+
+ @staticmethod
+ def get_preprocess_shape(oldh: int, oldw: int,
+ long_side_length: int) -> Tuple[int, int]:
+ """Compute the output size given input size and target long side
+ length."""
+ scale = long_side_length * 1.0 / max(oldh, oldw)
+ newh, neww = oldh * scale, oldw * scale
+ neww = int(neww + 0.5)
+ newh = int(newh + 0.5)
+ return (newh, neww)
diff --git a/projects/sam_inference_demo/sam_image_demo.ipynb b/projects/sam_inference_demo/sam_image_demo.ipynb
new file mode 100644
index 00000000000..1cb433fae9b
--- /dev/null
+++ b/projects/sam_inference_demo/sam_image_demo.ipynb
@@ -0,0 +1,122 @@
+{
+ "cells": [
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import numpy as np\n",
+ "import matplotlib.pyplot as plt\n",
+ "import cv2\n",
+ "\n",
+ "import sam # noqa: F401\n",
+ "from sam.sam_inferencer import SAMInferencer\n",
+ "\n",
+ "\n",
+ "def show_mask(mask, ax, random_color=False):\n",
+ " if random_color:\n",
+ " color = np.concatenate([np.random.random(3), np.array([0.6])], axis=0)\n",
+ " else:\n",
+ " color = np.array([30/255, 144/255, 255/255, 0.6])\n",
+ " h, w = mask.shape[-2:]\n",
+ " mask_image = mask.reshape(h, w, 1) * color.reshape(1, 1, -1)\n",
+ " ax.imshow(mask_image)\n",
+ " \n",
+ "def show_points(coords, labels, ax, marker_size=375):\n",
+ " pos_points = coords[labels==1]\n",
+ " neg_points = coords[labels==0]\n",
+ " ax.scatter(pos_points[:, 0], pos_points[:, 1], color='green', marker='*', s=marker_size, edgecolor='white', linewidth=1.25)\n",
+ " ax.scatter(neg_points[:, 0], neg_points[:, 1], color='red', marker='*', s=marker_size, edgecolor='white', linewidth=1.25) \n",
+ " \n",
+ "def show_box(box, ax):\n",
+ " x0, y0 = box[0], box[1]\n",
+ " w, h = box[2] - box[0], box[3] - box[1]\n",
+ " ax.add_patch(plt.Rectangle((x0, y0), w, h, edgecolor='green', facecolor=(0,0,0,0), lw=2))\n",
+ "\n",
+ "image = cv2.imread('../../demo/demo.png')\n",
+ "image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)\n",
+ "plt.figure(figsize=(10,10))\n",
+ "plt.imshow(image)\n",
+ "plt.axis('on')\n",
+ "plt.show()\n",
+ "print(image.shape)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "inferencer = SAMInferencer(arch='huge')\n",
+ "inferencer.set_image(image)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "input_point = np.array([[280, 230], [500, 300]])\n",
+ "input_label = np.array([1, 1])\n",
+ "plt.figure(figsize=(10,10))\n",
+ "plt.imshow(image)\n",
+ "show_points(input_point, input_label, plt.gca())\n",
+ "plt.axis('on')\n",
+ "plt.show() "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "masks, scores, logits = inferencer.predict(\n",
+ " point_coords=input_point,\n",
+ " point_labels=input_label,\n",
+ " multimask_output=True,\n",
+ ")\n",
+ "for i, (mask, score) in enumerate(zip(masks, scores)):\n",
+ " plt.figure(figsize=(10,10))\n",
+ " plt.imshow(image)\n",
+ " show_mask(mask, plt.gca(), random_color=True)\n",
+ " show_points(input_point, input_label, plt.gca())\n",
+ " plt.title(f\"Mask {i+1}, Score: {score:.3f}\", fontsize=18)\n",
+ " plt.axis('off')\n",
+ " plt.show()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": []
+ }
+ ],
+ "metadata": {
+ "kernelspec": {
+ "display_name": "pt1.13",
+ "language": "python",
+ "name": "python3"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.10.9"
+ },
+ "orig_nbformat": 4
+ },
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
diff --git a/projects/van/README.md b/projects/van/README.md
new file mode 100644
index 00000000000..be0ba362faa
--- /dev/null
+++ b/projects/van/README.md
@@ -0,0 +1,101 @@
+# Visual Attention Network (VAN) for Segmentation
+
+This repo is a PyTorch implementation of applying **VAN** (**Visual Attention Network**) to semantic segmentation.
+
+The code is an integration from [VAN-Segmentation](https://github.com/Visual-Attention-Network/VAN-Segmentation/blob/main/README.md?plain=1)
+
+More details can be found in [**Visual Attention Network**](https://arxiv.org/abs/2202.09741).
+
+## Citation
+
+```bib
+@article{guo2022visual,
+ title={Visual Attention Network},
+ author={Guo, Meng-Hao and Lu, Cheng-Ze and Liu, Zheng-Ning and Cheng, Ming-Ming and Hu, Shi-Min},
+ journal={arXiv preprint arXiv:2202.09741},
+ year={2022}
+}
+```
+
+## Results
+
+**Notes**: Pre-trained models can be found in [TsingHua Cloud](https://cloud.tsinghua.edu.cn/d/0100f0cea37d41ba8d08/).
+
+Results can be found in [VAN-Segmentation](https://github.com/Visual-Attention-Network/VAN-Segmentation/blob/main/README.md?plain=1)
+
+We provide evaluation results of the converted weights.
+
+| Method | Backbone | mIoU | Download |
+| :-----: | :----------: | :---: | :--------------------------------------------------------------------------------------------------------------------------------------------: |
+| UPerNet | VAN-B2 | 49.35 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2-in1kpre_upernet_3rdparty_512x512-ade20k_20230522-19c58aee.pth) |
+| UPerNet | VAN-B3 | 49.71 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3-in1kpre_upernet_3rdparty_512x512-ade20k_20230522-653bd6b7.pth) |
+| UPerNet | VAN-B4 | 51.56 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4-in1kpre_upernet_3rdparty_512x512-ade20k_20230522-653bd6b7.pth) |
+| UPerNet | VAN-B4-in22k | 52.61 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4-in22kpre_upernet_3rdparty_512x512-ade20k_20230522-4a4d744a.pth) |
+| UPerNet | VAN-B5-in22k | 53.11 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b5-in22kpre_upernet_3rdparty_512x512-ade20k_20230522-5bb6f2b4.pth) |
+| UPerNet | VAN-B6-in22k | 54.25 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b6-in22kpre_upernet_3rdparty_512x512-ade20k_20230522-e226b363.pth) |
+| FPN | VAN-B0 | 38.65 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b0-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-75a76298.pth) |
+| FPN | VAN-B1 | 43.22 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b1-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-104499ff.pth) |
+| FPN | VAN-B2 | 46.84 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-7074e6f8.pth) |
+| FPN | VAN-B3 | 48.32 | [model](https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3-in1kpre_fpn_3rdparty_512x512-ade20k_20230522-2c3b7f5e.pth) |
+
+## Preparation
+
+Install MMSegmentation and download ADE20K according to the guidelines in MMSegmentation.
+
+## Requirement
+
+**Step 0.** Install [MMCV](https://github.com/open-mmlab/mmcv) using [MIM](https://github.com/open-mmlab/mim).
+
+```shell
+pip install -U openmim
+mim install mmengine
+mim install "mmcv>=2.0.0"
+```
+
+**Step 1.** Install MMSegmentation.
+
+Case a: If you develop and run mmseg directly, install it from source:
+
+```shell
+git clone -b main https://github.com/open-mmlab/mmsegmentation.git
+cd mmsegmentation
+pip install -v -e .
+```
+
+Case b: If you use mmsegmentation as a dependency or third-party package, install it with pip:
+
+```shell
+pip install "mmsegmentation>=1.0.0"
+```
+
+## Training
+
+If you use 4 GPUs for training by default. Run:
+
+```bash
+bash tools/dist_train.sh projects/van/configs/van/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512.py 4
+```
+
+## Evaluation
+
+To evaluate the model, an example is:
+
+```bash
+bash tools/dist_train.sh projects/van/configs/van/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512.py work_dirs/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512/iter_160000.pth 4 --eval mIoU
+```
+
+## FLOPs
+
+To calculate FLOPs for a model, run:
+
+```bash
+bash tools/analysis_tools/get_flops.py projects/van/configs/van/van-b2_pre1k_upernet_4xb2-160k_ade20k-512x512.py --shape 512 512
+```
+
+## Acknowledgment
+
+Our implementation is mainly based on [mmsegmentation](https://github.com/open-mmlab/mmsegmentation/tree/v0.12.0), [Swin-Transformer](https://github.com/SwinTransformer/Swin-Transformer-Semantic-Segmentation), [PoolFormer](https://github.com/sail-sg/poolformer), [Enjoy-Hamburger](https://github.com/Gsunshine/Enjoy-Hamburger) and [VAN-Segmentation](https://github.com/Visual-Attention-Network/VAN-Segmentation/blob/main/README.md?plain=1). Thanks for their authors.
+
+## LICENSE
+
+This repo is under the Apache-2.0 license. For commercial use, please contact the authors.
diff --git a/projects/van/backbones/__init__.py b/projects/van/backbones/__init__.py
new file mode 100644
index 00000000000..071995de294
--- /dev/null
+++ b/projects/van/backbones/__init__.py
@@ -0,0 +1,4 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from .van import VAN
+
+__all__ = ['VAN']
diff --git a/projects/van/backbones/van.py b/projects/van/backbones/van.py
new file mode 100644
index 00000000000..301834a7587
--- /dev/null
+++ b/projects/van/backbones/van.py
@@ -0,0 +1,124 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import warnings
+
+import torch
+import torch.nn as nn
+from mmengine.model import BaseModule
+
+from mmseg.models.backbones.mscan import (MSCAN, MSCABlock,
+ MSCASpatialAttention,
+ OverlapPatchEmbed)
+from mmseg.registry import MODELS
+
+
+class VANAttentionModule(BaseModule):
+
+ def __init__(self, in_channels):
+ super().__init__()
+ self.conv0 = nn.Conv2d(
+ in_channels, in_channels, 5, padding=2, groups=in_channels)
+ self.conv_spatial = nn.Conv2d(
+ in_channels,
+ in_channels,
+ 7,
+ stride=1,
+ padding=9,
+ groups=in_channels,
+ dilation=3)
+ self.conv1 = nn.Conv2d(in_channels, in_channels, 1)
+
+ def forward(self, x):
+ u = x.clone()
+ attn = self.conv0(x)
+ attn = self.conv_spatial(attn)
+ attn = self.conv1(attn)
+ return u * attn
+
+
+class VANSpatialAttention(MSCASpatialAttention):
+
+ def __init__(self, in_channels, act_cfg=dict(type='GELU')):
+ super().__init__(in_channels, act_cfg=act_cfg)
+ self.spatial_gating_unit = VANAttentionModule(in_channels)
+
+
+class VANBlock(MSCABlock):
+
+ def __init__(self,
+ channels,
+ mlp_ratio=4.,
+ drop=0.,
+ drop_path=0.,
+ act_cfg=dict(type='GELU'),
+ norm_cfg=dict(type='SyncBN', requires_grad=True)):
+ super().__init__(
+ channels,
+ mlp_ratio=mlp_ratio,
+ drop=drop,
+ drop_path=drop_path,
+ act_cfg=act_cfg,
+ norm_cfg=norm_cfg)
+ self.attn = VANSpatialAttention(channels)
+
+
+@MODELS.register_module()
+class VAN(MSCAN):
+
+ def __init__(self,
+ in_channels=3,
+ embed_dims=[64, 128, 256, 512],
+ mlp_ratios=[8, 8, 4, 4],
+ drop_rate=0.,
+ drop_path_rate=0.,
+ depths=[3, 4, 6, 3],
+ num_stages=4,
+ act_cfg=dict(type='GELU'),
+ norm_cfg=dict(type='SyncBN', requires_grad=True),
+ pretrained=None,
+ init_cfg=None):
+ super(MSCAN, self).__init__(init_cfg=init_cfg)
+
+ assert not (init_cfg and pretrained), \
+ 'init_cfg and pretrained cannot be set at the same time'
+ if isinstance(pretrained, str):
+ warnings.warn('DeprecationWarning: pretrained is deprecated, '
+ 'please use "init_cfg" instead')
+ self.init_cfg = dict(type='Pretrained', checkpoint=pretrained)
+ elif pretrained is not None:
+ raise TypeError('pretrained must be a str or None')
+
+ self.depths = depths
+ self.num_stages = num_stages
+
+ # stochastic depth decay rule
+ dpr = [
+ x.item() for x in torch.linspace(0, drop_path_rate, sum(depths))
+ ]
+ cur = 0
+
+ for i in range(num_stages):
+ patch_embed = OverlapPatchEmbed(
+ patch_size=7 if i == 0 else 3,
+ stride=4 if i == 0 else 2,
+ in_channels=in_channels if i == 0 else embed_dims[i - 1],
+ embed_dim=embed_dims[i],
+ norm_cfg=norm_cfg)
+
+ block = nn.ModuleList([
+ VANBlock(
+ channels=embed_dims[i],
+ mlp_ratio=mlp_ratios[i],
+ drop=drop_rate,
+ drop_path=dpr[cur + j],
+ act_cfg=act_cfg,
+ norm_cfg=norm_cfg) for j in range(depths[i])
+ ])
+ norm = nn.LayerNorm(embed_dims[i])
+ cur += depths[i]
+
+ setattr(self, f'patch_embed{i + 1}', patch_embed)
+ setattr(self, f'block{i + 1}', block)
+ setattr(self, f'norm{i + 1}', norm)
+
+ def init_weights(self):
+ return super().init_weights()
diff --git a/projects/van/configs/_base_/datasets/ade20k.py b/projects/van/configs/_base_/datasets/ade20k.py
new file mode 100644
index 00000000000..69b3c2a73b8
--- /dev/null
+++ b/projects/van/configs/_base_/datasets/ade20k.py
@@ -0,0 +1,14 @@
+# dataset settings
+_base_ = '../../../../../configs/_base_/datasets/ade20k.py'
+
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 512), keep_ratio=True),
+ dict(type='ResizeToMultiple', size_divisor=32),
+ # add loading annotation after ``Resize`` because ground truth
+ # does not need to do resize data transform
+ dict(type='LoadAnnotations', reduce_zero_label=True),
+ dict(type='PackSegInputs')
+]
+val_dataloader = dict(dataset=dict(pipeline=test_pipeline))
+test_dataloader = val_dataloader
diff --git a/projects/van/configs/_base_/models/van_fpn.py b/projects/van/configs/_base_/models/van_fpn.py
new file mode 100644
index 00000000000..c7fd7391f77
--- /dev/null
+++ b/projects/van/configs/_base_/models/van_fpn.py
@@ -0,0 +1,43 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255,
+ size=(512, 512))
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ type='VAN',
+ embed_dims=[32, 64, 160, 256],
+ drop_rate=0.0,
+ drop_path_rate=0.1,
+ depths=[3, 3, 5, 2],
+ act_cfg=dict(type='GELU'),
+ norm_cfg=norm_cfg,
+ init_cfg=dict()),
+ neck=dict(
+ type='FPN',
+ in_channels=[32, 64, 160, 256],
+ out_channels=256,
+ num_outs=4),
+ decode_head=dict(
+ type='FPNHead',
+ in_channels=[256, 256, 256, 256],
+ in_index=[0, 1, 2, 3],
+ feature_strides=[4, 8, 16, 32],
+ channels=128,
+ dropout_ratio=0.1,
+ num_classes=150,
+ norm_cfg=norm_cfg,
+ align_corners=False,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0)),
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/van/configs/_base_/models/van_upernet.py b/projects/van/configs/_base_/models/van_upernet.py
new file mode 100644
index 00000000000..8f94c0d9d84
--- /dev/null
+++ b/projects/van/configs/_base_/models/van_upernet.py
@@ -0,0 +1,51 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[123.675, 116.28, 103.53],
+ std=[58.395, 57.12, 57.375],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255,
+ size=(512, 512))
+model = dict(
+ type='EncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ type='VAN',
+ embed_dims=[32, 64, 160, 256],
+ drop_rate=0.0,
+ drop_path_rate=0.1,
+ depths=[3, 3, 5, 2],
+ act_cfg=dict(type='GELU'),
+ norm_cfg=norm_cfg,
+ init_cfg=dict()),
+ decode_head=dict(
+ type='UPerHead',
+ in_channels=[32, 64, 160, 256],
+ in_index=[0, 1, 2, 3],
+ pool_scales=(1, 2, 3, 6),
+ channels=512,
+ dropout_ratio=0.1,
+ num_classes=150,
+ norm_cfg=norm_cfg,
+ align_corners=False,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=1.0)),
+ auxiliary_head=dict(
+ type='FCNHead',
+ in_channels=160,
+ in_index=2,
+ channels=256,
+ num_convs=1,
+ concat_input=False,
+ dropout_ratio=0.1,
+ num_classes=150,
+ norm_cfg=norm_cfg,
+ align_corners=False,
+ loss_decode=dict(
+ type='CrossEntropyLoss', use_sigmoid=False, loss_weight=0.4)),
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole'))
diff --git a/projects/van/configs/van/van-b0_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b0_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..2faf3788a71
--- /dev/null
+++ b/projects/van/configs/van/van-b0_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,8 @@
+_base_ = './van-b2_fpn_8xb4-40k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b0_3rdparty_20230522-956f5e0d.pth' # noqa
+model = dict(
+ backbone=dict(
+ embed_dims=[32, 64, 160, 256],
+ depths=[3, 3, 5, 2],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path)),
+ neck=dict(in_channels=[32, 64, 160, 256]))
diff --git a/projects/van/configs/van/van-b1_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b1_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..cf64a7138b2
--- /dev/null
+++ b/projects/van/configs/van/van-b1_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,6 @@
+_base_ = './van-b2_fpn_8xb4-40k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b1_3rdparty_20230522-3adb117f.pth' # noqa
+model = dict(
+ backbone=dict(
+ depths=[2, 2, 4, 2],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path)))
diff --git a/projects/van/configs/van/van-b2_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b2_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..965fa1cd363
--- /dev/null
+++ b/projects/van/configs/van/van-b2_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,53 @@
+_base_ = [
+ '../_base_/models/van_fpn.py',
+ '../_base_/datasets/ade20k.py',
+ '../../../../configs/_base_/default_runtime.py',
+]
+custom_imports = dict(imports=['projects.van.backbones'])
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2_3rdparty_20230522-636fac93.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ embed_dims=[64, 128, 320, 512],
+ depths=[3, 3, 12, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.2),
+ neck=dict(in_channels=[64, 128, 320, 512]),
+ decode_head=dict(num_classes=150))
+
+train_dataloader = dict(batch_size=4)
+
+# we use 8 gpu instead of 4 in mmsegmentation, so lr*2 and max_iters/2
+gpu_multiples = 2
+max_iters = 80000 // gpu_multiples
+interval = 8000 // gpu_multiples
+optim_wrapper = dict(
+ type='OptimWrapper',
+ optimizer=dict(
+ type='AdamW',
+ lr=0.0001 * gpu_multiples,
+ # betas=(0.9, 0.999),
+ weight_decay=0.0001),
+ clip_grad=None)
+# learning policy
+param_scheduler = [
+ dict(
+ type='PolyLR',
+ power=0.9,
+ eta_min=0.0,
+ begin=0,
+ end=max_iters,
+ by_epoch=False,
+ )
+]
+train_cfg = dict(
+ type='IterBasedTrainLoop', max_iters=max_iters, val_interval=interval)
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
+default_hooks = dict(
+ timer=dict(type='IterTimerHook'),
+ logger=dict(type='LoggerHook', interval=50, log_metric_by_epoch=False),
+ param_scheduler=dict(type='ParamSchedulerHook'),
+ checkpoint=dict(type='CheckpointHook', by_epoch=False, interval=interval),
+ sampler_seed=dict(type='DistSamplerSeedHook'),
+ visualization=dict(type='SegVisualizationHook'))
diff --git a/projects/van/configs/van/van-b2_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b2_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..c529606a202
--- /dev/null
+++ b/projects/van/configs/van/van-b2_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,46 @@
+_base_ = [
+ '../_base_/models/van_upernet.py', '../_base_/datasets/ade20k.py',
+ '../../../../configs/_base_/default_runtime.py',
+ '../../../../configs/_base_/schedules/schedule_160k.py'
+]
+custom_imports = dict(imports=['projects.van.backbones'])
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b2_3rdparty_20230522-636fac93.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ embed_dims=[64, 128, 320, 512],
+ depths=[3, 3, 12, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path)),
+ decode_head=dict(in_channels=[64, 128, 320, 512], num_classes=150),
+ auxiliary_head=dict(in_channels=320, num_classes=150))
+
+# AdamW optimizer
+# no weight decay for position embedding & layer norm in backbone
+optim_wrapper = dict(
+ _delete_=True,
+ type='OptimWrapper',
+ optimizer=dict(
+ type='AdamW', lr=0.00006, betas=(0.9, 0.999), weight_decay=0.01),
+ clip_grad=None,
+ paramwise_cfg=dict(
+ custom_keys={
+ 'absolute_pos_embed': dict(decay_mult=0.),
+ 'relative_position_bias_table': dict(decay_mult=0.),
+ 'norm': dict(decay_mult=0.)
+ }))
+# learning policy
+param_scheduler = [
+ dict(
+ type='LinearLR', start_factor=1e-6, by_epoch=False, begin=0, end=1500),
+ dict(
+ type='PolyLR',
+ power=1.0,
+ begin=1500,
+ end=_base_.train_cfg.max_iters,
+ eta_min=0.0,
+ by_epoch=False,
+ )
+]
+
+# By default, models are trained on 8 GPUs with 2 images per GPU
+train_dataloader = dict(batch_size=2)
diff --git a/projects/van/configs/van/van-b3_fpn_8xb4-40k_ade20k-512x512.py b/projects/van/configs/van/van-b3_fpn_8xb4-40k_ade20k-512x512.py
new file mode 100644
index 00000000000..b0493fe4f9f
--- /dev/null
+++ b/projects/van/configs/van/van-b3_fpn_8xb4-40k_ade20k-512x512.py
@@ -0,0 +1,11 @@
+_base_ = './van-b2_fpn_8xb4-40k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3_3rdparty_20230522-a184e051.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ embed_dims=[64, 128, 320, 512],
+ depths=[3, 5, 27, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.3),
+ neck=dict(in_channels=[64, 128, 320, 512]))
+train_dataloader = dict(batch_size=4)
diff --git a/projects/van/configs/van/van-b3_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b3_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..8201801d992
--- /dev/null
+++ b/projects/van/configs/van/van-b3_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,8 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b3_3rdparty_20230522-a184e051.pth' # noqa
+model = dict(
+ type='EncoderDecoder',
+ backbone=dict(
+ depths=[3, 5, 27, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.3))
diff --git a/projects/van/configs/van/van-b4-in22kpre_upernet_4xb4-160k_ade20k-512x512.py b/projects/van/configs/van/van-b4-in22kpre_upernet_4xb4-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..15c8f7ca6ee
--- /dev/null
+++ b/projects/van/configs/van/van-b4-in22kpre_upernet_4xb4-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4-in22k_3rdparty_20230522-5e31cafb.pth' # noqa
+model = dict(
+ backbone=dict(
+ depths=[3, 6, 40, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.4))
+
+# By default, models are trained on 8 GPUs with 2 images per GPU
+train_dataloader = dict(batch_size=4)
diff --git a/projects/van/configs/van/van-b4_upernet_4xb4-160k_ade20k-512x512.py b/projects/van/configs/van/van-b4_upernet_4xb4-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..33ae049d0c0
--- /dev/null
+++ b/projects/van/configs/van/van-b4_upernet_4xb4-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b4_3rdparty_20230522-1d71c077.pth' # noqa
+model = dict(
+ backbone=dict(
+ depths=[3, 6, 40, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.4))
+
+# By default, models are trained on 4 GPUs with 4 images per GPU
+train_dataloader = dict(batch_size=4)
diff --git a/projects/van/configs/van/van-b5-in22kpre_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b5-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..f36c6242bdf
--- /dev/null
+++ b/projects/van/configs/van/van-b5-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b5-in22k_3rdparty_20230522-b26134d7.pth' # noqa
+model = dict(
+ backbone=dict(
+ embed_dims=[96, 192, 480, 768],
+ depths=[3, 3, 24, 3],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.4),
+ decode_head=dict(in_channels=[96, 192, 480, 768], num_classes=150),
+ auxiliary_head=dict(in_channels=480, num_classes=150))
diff --git a/projects/van/configs/van/van-b6-in22kpre_upernet_4xb2-160k_ade20k-512x512.py b/projects/van/configs/van/van-b6-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
new file mode 100644
index 00000000000..aa529efed8c
--- /dev/null
+++ b/projects/van/configs/van/van-b6-in22kpre_upernet_4xb2-160k_ade20k-512x512.py
@@ -0,0 +1,10 @@
+_base_ = './van-b2_upernet_4xb2-160k_ade20k-512x512.py'
+ckpt_path = 'https://download.openmmlab.com/mmsegmentation/v0.5/van_3rdparty/van-b6-in22k_3rdparty_20230522-5e5172a3.pth' # noqa
+model = dict(
+ backbone=dict(
+ embed_dims=[96, 192, 384, 768],
+ depths=[6, 6, 90, 6],
+ init_cfg=dict(type='Pretrained', checkpoint=ckpt_path),
+ drop_path_rate=0.5),
+ decode_head=dict(in_channels=[96, 192, 384, 768], num_classes=150),
+ auxiliary_head=dict(in_channels=384, num_classes=150))
diff --git a/requirements.txt b/requirements.txt
index 6da5adea757..501bddc8843 100644
--- a/requirements.txt
+++ b/requirements.txt
@@ -1,3 +1,4 @@
-r requirements/optional.txt
-r requirements/runtime.txt
-r requirements/tests.txt
+-r requirements/multimodal.txt
diff --git a/requirements/albu.txt b/requirements/albu.txt
new file mode 100644
index 00000000000..f421fbbdc47
--- /dev/null
+++ b/requirements/albu.txt
@@ -0,0 +1 @@
+albumentations>=0.3.2 --no-binary qudida,albumentations
diff --git a/requirements/docs.txt b/requirements/docs.txt
index 8e98c16fc72..19632d36aba 100644
--- a/requirements/docs.txt
+++ b/requirements/docs.txt
@@ -4,3 +4,4 @@ myst-parser
sphinx==4.0.2
sphinx_copybutton
sphinx_markdown_tables
+urllib3<2.0.0
diff --git a/requirements/multimodal.txt b/requirements/multimodal.txt
new file mode 100644
index 00000000000..2195d0d9ef8
--- /dev/null
+++ b/requirements/multimodal.txt
@@ -0,0 +1,2 @@
+ftfy
+regex
diff --git a/requirements/optional.txt b/requirements/optional.txt
index 5eca649247e..b0310f52960 100644
--- a/requirements/optional.txt
+++ b/requirements/optional.txt
@@ -1,2 +1,22 @@
cityscapesscripts
+-e git+https://github.com/openai/CLIP.git@main#egg=clip
+
+# for vpd model
+diffusers
+einops==0.3.0
+imageio==2.9.0
+imageio-ffmpeg==0.4.2
+invisible-watermark
+kornia==0.6
+-e git+https://github.com/CompVis/stable-diffusion@21f890f#egg=latent-diffusion
nibabel
+omegaconf==2.1.1
+pudb==2019.2
+pytorch-lightning==1.4.2
+streamlit>=0.73.1
+-e git+https://github.com/CompVis/taming-transformers.git@master#egg=taming-transformers
+test-tube>=0.7.5
+timm
+torch-fidelity==0.3.0
+torchmetrics==0.6.0
+transformers==4.19.2
diff --git a/requirements/tests.txt b/requirements/tests.txt
index 74fc76146d8..3fff2520d7c 100644
--- a/requirements/tests.txt
+++ b/requirements/tests.txt
@@ -1,6 +1,8 @@
codecov
flake8
+ftfy
interrogate
pytest
+regex
xdoctest>=0.10.0
yapf
diff --git a/resources/miaomiao_qrcode.jpg b/resources/miaomiao_qrcode.jpg
new file mode 100644
index 00000000000..d34cbae6fd1
Binary files /dev/null and b/resources/miaomiao_qrcode.jpg differ
diff --git a/setup.cfg b/setup.cfg
index dc5ea071110..2ea07600c0b 100644
--- a/setup.cfg
+++ b/setup.cfg
@@ -16,4 +16,4 @@ default_section = THIRDPARTY
skip = *.po,*.ts,*.ipynb
count =
quiet-level = 3
-ignore-words-list = formating,sur,hist,dota,warmup
+ignore-words-list = formating,sur,hist,dota,warmup,damon
diff --git a/setup.py b/setup.py
index 854dd186054..45d923db60e 100755
--- a/setup.py
+++ b/setup.py
@@ -123,7 +123,7 @@ def add_mim_extension():
else:
return
- filenames = ['tools', 'configs', 'model-index.yml']
+ filenames = ['tools', 'configs', 'model-index.yml', 'dataset-index.yml']
repo_path = osp.dirname(__file__)
mim_path = osp.join(repo_path, 'mmseg', '.mim')
os.makedirs(mim_path, exist_ok=True)
@@ -175,7 +175,7 @@ def add_mim_extension():
author='MMSegmentation Contributors',
author_email='openmmlab@gmail.com',
keywords='computer vision, semantic segmentation',
- url='http://github.com/open-mmlab/mmsegmentation',
+ url='https://github.com/open-mmlab/mmsegmentation',
packages=find_packages(exclude=('configs', 'tools', 'demo')),
include_package_data=True,
classifiers=[
@@ -194,6 +194,7 @@ def add_mim_extension():
'tests': parse_requirements('requirements/tests.txt'),
'optional': parse_requirements('requirements/optional.txt'),
'mim': parse_requirements('requirements/mminstall.txt'),
+ 'multimodal': parse_requirements('requirements/multimodal.txt'),
},
ext_modules=[],
zip_safe=False)
diff --git a/tests/data/dsdl_seg/config.py b/tests/data/dsdl_seg/config.py
new file mode 100755
index 00000000000..8eed751c2f7
--- /dev/null
+++ b/tests/data/dsdl_seg/config.py
@@ -0,0 +1,13 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+local = dict(
+ type='LocalFileReader',
+ working_dir='/nvme/share_data/VOC2012',
+)
+
+ali_oss = dict(
+ type='AliOSSFileReader',
+ access_key_secret='your secret key of aliyun oss',
+ endpoint='your endpoint of aliyun oss',
+ access_key_id='your access key of aliyun oss',
+ bucket_name='your bucket name of aliyun oss',
+ working_dir='the relative path of your media dir in the bucket')
diff --git a/tests/data/dsdl_seg/defs/class-dom.yaml b/tests/data/dsdl_seg/defs/class-dom.yaml
new file mode 100755
index 00000000000..e5dd598c4ae
--- /dev/null
+++ b/tests/data/dsdl_seg/defs/class-dom.yaml
@@ -0,0 +1,24 @@
+$dsdl-version: "0.5.0"
+VOCClassDom:
+ $def: class_domain
+ classes:
+ - aeroplane
+ - bicycle
+ - bird
+ - boat
+ - bottle
+ - bus
+ - car
+ - cat
+ - chair
+ - cow
+ - diningtable
+ - dog
+ - horse
+ - motorbike
+ - person
+ - pottedplant
+ - sheep
+ - sofa
+ - train
+ - tvmonitor
diff --git a/tests/data/dsdl_seg/defs/segmentation-def.yaml b/tests/data/dsdl_seg/defs/segmentation-def.yaml
new file mode 100755
index 00000000000..057139ed57e
--- /dev/null
+++ b/tests/data/dsdl_seg/defs/segmentation-def.yaml
@@ -0,0 +1,15 @@
+$dsdl-version: "0.5.0"
+
+ImageMedia:
+ $def: struct
+ $fields:
+ image: Image
+ image_shape: ImageShape
+
+SegmentationSample:
+ $def: struct
+ $params: ['cdom']
+ $fields:
+ media: ImageMedia
+ label_map: LabelMap[dom=$cdom]
+ instance_map: InstanceMap
diff --git a/tests/data/dsdl_seg/set-train/train.yaml b/tests/data/dsdl_seg/set-train/train.yaml
new file mode 100755
index 00000000000..69872445a53
--- /dev/null
+++ b/tests/data/dsdl_seg/set-train/train.yaml
@@ -0,0 +1,15 @@
+$dsdl-version: "0.5.0"
+$import:
+ - ../defs/segmentation-def
+ - ../defs/class-dom
+meta:
+ dataset_name: "VOC2012"
+ sub_dataset_name: "train"
+ task_type: "Segmentation"
+ dataset_homepage: "http://host.robots.ox.ac.uk/pascal/VOC/voc2012/index.html"
+ dataset_publisher: "University of Leeds | ETHZ, Zurich | University of Edinburgh\
+ \ |Microsoft Research Cambridge | University of Oxford"
+ OpenDataLab_address: "https://opendatalab.com/PASCAL_VOC2012/download"
+data:
+ sample-type: SegmentationSample[cdom=VOCClassDom]
+ sample-path: train_samples.json
diff --git a/tests/data/dsdl_seg/set-train/train_samples.json b/tests/data/dsdl_seg/set-train/train_samples.json
new file mode 100755
index 00000000000..559f5845721
--- /dev/null
+++ b/tests/data/dsdl_seg/set-train/train_samples.json
@@ -0,0 +1 @@
+{"samples": [{"media": {"image": "JPEGImages/2007_000032.jpg", "image_shape": [281, 500]}, "label_map": "SegmentationClass/2007_000032.png", "instance_map": "SegmentationObject/2007_000032.png"}, {"media": {"image": "JPEGImages/2007_000039.jpg", "image_shape": [375, 500]}, "label_map": "SegmentationClass/2007_000039.png", "instance_map": "SegmentationObject/2007_000039.png"}, {"media": {"image": "JPEGImages/2007_000063.jpg", "image_shape": [375, 500]}, "label_map": "SegmentationClass/2007_000063.png", "instance_map": "SegmentationObject/2007_000063.png"}]}
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/train/0004a4c0-d4dff0ad.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/train/0004a4c0-d4dff0ad.jpg
new file mode 100644
index 00000000000..4724a3d9302
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/train/0004a4c0-d4dff0ad.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/train/00054602-3bf57337.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00054602-3bf57337.jpg
new file mode 100644
index 00000000000..5efe06b99a4
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00054602-3bf57337.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/train/00067cfb-e535423e.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00067cfb-e535423e.jpg
new file mode 100644
index 00000000000..2233c03c763
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/train/00067cfb-e535423e.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d06fefd-f7be05a6.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d06fefd-f7be05a6.jpg
new file mode 100644
index 00000000000..535087b5a95
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d06fefd-f7be05a6.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d128593-0ccfea4c.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d128593-0ccfea4c.jpg
new file mode 100644
index 00000000000..7f2971afdee
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d128593-0ccfea4c.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d15b18b-1e0d6e3f.jpg b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d15b18b-1e0d6e3f.jpg
new file mode 100644
index 00000000000..31a951d4830
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/images/10k/val/7d15b18b-1e0d6e3f.jpg differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/0004a4c0-d4dff0ad.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/0004a4c0-d4dff0ad.png
new file mode 100644
index 00000000000..086a8d50647
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/0004a4c0-d4dff0ad.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00054602-3bf57337.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00054602-3bf57337.png
new file mode 100644
index 00000000000..43338c283cc
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00054602-3bf57337.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00067cfb-e535423e.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00067cfb-e535423e.png
new file mode 100644
index 00000000000..7c0ad1d5d90
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/train/00067cfb-e535423e.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d128593-0ccfea4c.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d128593-0ccfea4c.png
new file mode 100644
index 00000000000..43338c283cc
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d128593-0ccfea4c.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d15b18b-1e0d6e3f.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d15b18b-1e0d6e3f.png
new file mode 100644
index 00000000000..43338c283cc
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d15b18b-1e0d6e3f.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d2f7975-e0c1c5a7.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d2f7975-e0c1c5a7.png
new file mode 100644
index 00000000000..7c0ad1d5d90
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/colormaps/val/7d2f7975-e0c1c5a7.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/0004a4c0-d4dff0ad.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/0004a4c0-d4dff0ad.png
new file mode 100644
index 00000000000..5c6bf5e1581
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/0004a4c0-d4dff0ad.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00054602-3bf57337.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00054602-3bf57337.png
new file mode 100644
index 00000000000..c525a768885
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00054602-3bf57337.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00067cfb-e535423e.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00067cfb-e535423e.png
new file mode 100644
index 00000000000..7dfd3af4e3a
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/train/00067cfb-e535423e.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d06fefd-f7be05a6.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d06fefd-f7be05a6.png
new file mode 100644
index 00000000000..7dfd3af4e3a
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d06fefd-f7be05a6.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d128593-0ccfea4c.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d128593-0ccfea4c.png
new file mode 100644
index 00000000000..c525a768885
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d128593-0ccfea4c.png differ
diff --git a/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d15b18b-1e0d6e3f.png b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d15b18b-1e0d6e3f.png
new file mode 100644
index 00000000000..c525a768885
Binary files /dev/null and b/tests/data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val/7d15b18b-1e0d6e3f.png differ
diff --git a/tests/data/pseudo_nyu_dataset/annotations/bookstore_0001d_00001.png b/tests/data/pseudo_nyu_dataset/annotations/bookstore_0001d_00001.png
new file mode 100644
index 00000000000..77e343603ad
Binary files /dev/null and b/tests/data/pseudo_nyu_dataset/annotations/bookstore_0001d_00001.png differ
diff --git a/tests/data/pseudo_nyu_dataset/images/bookstore_0001d_00001.jpg b/tests/data/pseudo_nyu_dataset/images/bookstore_0001d_00001.jpg
new file mode 100644
index 00000000000..7892ed47e7d
Binary files /dev/null and b/tests/data/pseudo_nyu_dataset/images/bookstore_0001d_00001.jpg differ
diff --git a/tests/test_apis/test_inferencer.py b/tests/test_apis/test_inferencer.py
index 497eae4a011..d8dbce8f385 100644
--- a/tests/test_apis/test_inferencer.py
+++ b/tests/test_apis/test_inferencer.py
@@ -3,76 +3,16 @@
import numpy as np
import torch
-import torch.nn as nn
from mmengine import ConfigDict
-from torch.utils.data import DataLoader, Dataset
+from utils import * # noqa: F401, F403
from mmseg.apis import MMSegInferencer
-from mmseg.models import EncoderDecoder
-from mmseg.models.decode_heads.decode_head import BaseDecodeHead
from mmseg.registry import MODELS
from mmseg.utils import register_all_modules
-@MODELS.register_module(name='InferExampleHead')
-class ExampleDecodeHead(BaseDecodeHead):
-
- def __init__(self, num_classes=19, out_channels=None):
- super().__init__(
- 3, 3, num_classes=num_classes, out_channels=out_channels)
-
- def forward(self, inputs):
- return self.cls_seg(inputs[0])
-
-
-@MODELS.register_module(name='InferExampleBackbone')
-class ExampleBackbone(nn.Module):
-
- def __init__(self):
- super().__init__()
- self.conv = nn.Conv2d(3, 3, 3)
-
- def init_weights(self, pretrained=None):
- pass
-
- def forward(self, x):
- return [self.conv(x)]
-
-
-@MODELS.register_module(name='InferExampleModel')
-class ExampleModel(EncoderDecoder):
-
- def __init__(self, **kwargs):
- super().__init__(**kwargs)
-
-
-class ExampleDataset(Dataset):
-
- def __init__(self) -> None:
- super().__init__()
- self.pipeline = [
- dict(type='LoadImageFromFile'),
- dict(type='LoadAnnotations'),
- dict(type='PackSegInputs')
- ]
-
- def __getitem__(self, idx):
- return dict(img=torch.tensor([1]), img_metas=dict())
-
- def __len__(self):
- return 1
-
-
def test_inferencer():
register_all_modules()
- test_dataset = ExampleDataset()
- data_loader = DataLoader(
- test_dataset,
- batch_size=1,
- sampler=None,
- num_workers=0,
- shuffle=False,
- )
visualizer = dict(
type='SegLocalVisualizer',
@@ -87,7 +27,14 @@ def test_inferencer():
decode_head=dict(type='InferExampleHead'),
test_cfg=dict(mode='whole')),
visualizer=visualizer,
- test_dataloader=data_loader)
+ test_dataloader=dict(
+ dataset=dict(
+ type='ExampleDataset',
+ pipeline=[
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+ ]), ))
cfg = ConfigDict(cfg_dict)
model = MODELS.build(cfg.model)
diff --git a/tests/test_apis/test_rs_inferencer.py b/tests/test_apis/test_rs_inferencer.py
new file mode 100644
index 00000000000..03423d9680e
--- /dev/null
+++ b/tests/test_apis/test_rs_inferencer.py
@@ -0,0 +1,73 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import os.path as osp
+from unittest import TestCase
+
+import numpy as np
+from mmengine import ConfigDict, init_default_scope
+from utils import * # noqa: F401, F403
+
+from mmseg.apis import RSImage, RSInferencer
+from mmseg.registry import MODELS
+
+
+class TestRSImage(TestCase):
+
+ def test_read_whole_image(self):
+ init_default_scope('mmseg')
+ img_path = osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_loveda_dataset/img_dir/0.png')
+ rs_image = RSImage(img_path)
+ window_size = (16, 16)
+ rs_image.create_grids(window_size)
+ image_data = rs_image.read(rs_image.grids[0])
+ self.assertIsNotNone(image_data)
+
+ def test_write_image_data(self):
+ init_default_scope('mmseg')
+ img_path = osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_loveda_dataset/img_dir/0.png')
+ rs_image = RSImage(img_path)
+ window_size = (16, 16)
+ rs_image.create_grids(window_size)
+ data = np.random.random((16, 16)).astype(np.int8)
+ rs_image.write(data, rs_image.grids[0])
+
+
+class TestRSInferencer(TestCase):
+
+ def test_read_and_inference(self):
+ init_default_scope('mmseg')
+ cfg_dict = dict(
+ model=dict(
+ type='InferExampleModel',
+ data_preprocessor=dict(type='SegDataPreProcessor'),
+ backbone=dict(type='InferExampleBackbone'),
+ decode_head=dict(type='InferExampleHead'),
+ test_cfg=dict(mode='whole')),
+ test_dataloader=dict(
+ dataset=dict(
+ type='ExampleDataset',
+ pipeline=[
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+ ])),
+ test_pipeline=[
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+ ])
+ cfg = ConfigDict(cfg_dict)
+ model = MODELS.build(cfg.model)
+ model.cfg = cfg
+ inferencer = RSInferencer.from_model(model)
+
+ img_path = osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_loveda_dataset/img_dir/0.png')
+ rs_image = RSImage(img_path)
+ window_size = (16, 16)
+ stride = (16, 16)
+ inferencer.run(rs_image, window_size, stride)
diff --git a/tests/test_apis/utils.py b/tests/test_apis/utils.py
new file mode 100644
index 00000000000..0a9928fccf1
--- /dev/null
+++ b/tests/test_apis/utils.py
@@ -0,0 +1,38 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch.nn as nn
+
+from mmseg.models import EncoderDecoder
+from mmseg.models.decode_heads.decode_head import BaseDecodeHead
+from mmseg.registry import MODELS
+
+
+@MODELS.register_module(name='InferExampleHead')
+class ExampleDecodeHead(BaseDecodeHead):
+
+ def __init__(self, num_classes=19, out_channels=None):
+ super().__init__(
+ 3, 3, num_classes=num_classes, out_channels=out_channels)
+
+ def forward(self, inputs):
+ return self.cls_seg(inputs[0])
+
+
+@MODELS.register_module(name='InferExampleBackbone')
+class ExampleBackbone(nn.Module):
+
+ def __init__(self):
+ super().__init__()
+ self.conv = nn.Conv2d(3, 3, 3)
+
+ def init_weights(self, pretrained=None):
+ pass
+
+ def forward(self, x):
+ return [self.conv(x)]
+
+
+@MODELS.register_module(name='InferExampleModel')
+class ExampleModel(EncoderDecoder):
+
+ def __init__(self, **kwargs):
+ super().__init__(**kwargs)
diff --git a/tests/test_config.py b/tests/test_config.py
index 13de460181c..cdd85ff57cd 100644
--- a/tests/test_config.py
+++ b/tests/test_config.py
@@ -30,6 +30,7 @@ def _get_config_directory():
def test_config_build_segmentor():
"""Test that all segmentation models defined in the configs can be
initialized."""
+ init_default_scope('mmseg')
config_dpath = _get_config_directory()
print(f'Found config_dpath = {config_dpath!r}')
@@ -92,7 +93,12 @@ def test_config_data_pipeline():
del config_mod.train_pipeline[0]
del config_mod.test_pipeline[0]
# remove loading annotation in test pipeline
- del config_mod.test_pipeline[-2]
+ load_anno_idx = -1
+ for i in range(len(config_mod.test_pipeline)):
+ if config_mod.test_pipeline[i].type in ('LoadAnnotations',
+ 'LoadDepthAnnotation'):
+ load_anno_idx = i
+ del config_mod.test_pipeline[load_anno_idx]
train_pipeline = Compose(config_mod.train_pipeline)
test_pipeline = Compose(config_mod.test_pipeline)
@@ -101,6 +107,7 @@ def test_config_data_pipeline():
if to_float32:
img = img.astype(np.float32)
seg = np.random.randint(0, 255, size=(1024, 2048, 1), dtype=np.uint8)
+ depth = np.random.rand(1024, 2048).astype(np.float32)
results = dict(
filename='test_img.png',
@@ -108,24 +115,30 @@ def test_config_data_pipeline():
img=img,
img_shape=img.shape,
ori_shape=img.shape,
- gt_seg_map=seg)
+ gt_seg_map=seg,
+ gt_depth_map=depth)
results['seg_fields'] = ['gt_seg_map']
-
+ _check_concat_cd_input(config_mod, results)
print(f'Test training data pipeline: \n{train_pipeline!r}')
output_results = train_pipeline(results)
assert output_results is not None
- results = dict(
- filename='test_img.png',
- ori_filename='test_img.png',
- img=img,
- img_shape=img.shape,
- ori_shape=img.shape)
+ _check_concat_cd_input(config_mod, results)
print(f'Test testing data pipeline: \n{test_pipeline!r}')
output_results = test_pipeline(results)
assert output_results is not None
+def _check_concat_cd_input(config_mod: Config, results: dict):
+ keys = []
+ pipeline = config_mod.train_pipeline.copy()
+ pipeline.extend(config_mod.test_pipeline)
+ for t in pipeline:
+ keys.append(t.type)
+ if 'ConcatCDInput' in keys:
+ results.update({'img2': results['img']})
+
+
def _check_decode_head(decode_head_cfg, decode_head):
if isinstance(decode_head_cfg, list):
assert isinstance(decode_head, nn.ModuleList)
@@ -149,14 +162,14 @@ def _check_decode_head(decode_head_cfg, decode_head):
elif input_transform == 'resize_concat':
assert sum(in_channels) == decode_head.in_channels
else:
- assert isinstance(in_channels, int)
assert in_channels == decode_head.in_channels
- assert isinstance(decode_head.in_index, int)
if decode_head_cfg['type'] == 'PointHead':
assert decode_head_cfg.channels+decode_head_cfg.num_classes == \
decode_head.fc_seg.in_channels
assert decode_head.fc_seg.out_channels == decode_head_cfg.num_classes
+ elif decode_head_cfg['type'] == 'VPDDepthHead':
+ assert decode_head.out_channels == 1
else:
assert decode_head_cfg.channels == decode_head.conv_seg.in_channels
assert decode_head.conv_seg.out_channels == decode_head_cfg.num_classes
diff --git a/tests/test_datasets/test_dataset.py b/tests/test_datasets/test_dataset.py
index db4a7799069..2904e09cedd 100644
--- a/tests/test_datasets/test_dataset.py
+++ b/tests/test_datasets/test_dataset.py
@@ -5,15 +5,21 @@
import pytest
-from mmseg.datasets import (ADE20KDataset, BaseSegDataset, CityscapesDataset,
- COCOStuffDataset, DecathlonDataset, ISPRSDataset,
+from mmseg.datasets import (ADE20KDataset, BaseSegDataset, BDD100KDataset,
+ CityscapesDataset, COCOStuffDataset,
+ DecathlonDataset, DSDLSegDataset, ISPRSDataset,
LIPDataset, LoveDADataset, MapillaryDataset_v1,
- MapillaryDataset_v2, PascalVOCDataset,
+ MapillaryDataset_v2, NYUDataset, PascalVOCDataset,
PotsdamDataset, REFUGEDataset, SynapseDataset,
iSAIDDataset)
from mmseg.registry import DATASETS
from mmseg.utils import get_classes, get_palette
+try:
+ from dsdl.dataset import DSDLDataset
+except ImportError:
+ DSDLDataset = None
+
def test_classes():
assert list(
@@ -32,7 +38,7 @@ def test_classes():
MapillaryDataset_v1.METAINFO['classes']) == get_classes('mapillary_v1')
assert list(
MapillaryDataset_v2.METAINFO['classes']) == get_classes('mapillary_v2')
-
+ assert list(BDD100KDataset.METAINFO['classes']) == get_classes('bdd100k')
with pytest.raises(ValueError):
get_classes('unsupported')
@@ -89,7 +95,7 @@ def test_palette():
MapillaryDataset_v1.METAINFO['palette']) == get_palette('mapillary_v1')
assert list(
MapillaryDataset_v2.METAINFO['palette']) == get_palette('mapillary_v2')
-
+ assert list(BDD100KDataset.METAINFO['palette']) == get_palette('bdd100k')
with pytest.raises(ValueError):
get_palette('unsupported')
@@ -326,6 +332,19 @@ def test_mapillary():
assert len(test_dataset) == 1
+def test_bdd100k():
+ test_dataset = BDD100KDataset(
+ pipeline=[],
+ data_prefix=dict(
+ img_path=osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_bdd100k_dataset/images/10k/val'),
+ seg_map_path=osp.join(
+ osp.dirname(__file__),
+ '../data/pseudo_bdd100k_dataset/labels/sem_seg/masks/val')))
+ assert len(test_dataset) == 3
+
+
@pytest.mark.parametrize('dataset, classes', [
('ADE20KDataset', ('wall', 'building')),
('CityscapesDataset', ('road', 'sidewalk')),
@@ -433,3 +452,24 @@ def test_custom_dataset_custom_palette():
ann_file=tempfile.mkdtemp(),
metainfo=dict(classes=('bus', 'car'), palette=[[200, 200, 200]]),
lazy_init=True)
+
+
+def test_dsdlseg_dataset():
+ if DSDLDataset is not None:
+ dataset = DSDLSegDataset(
+ data_root='tests/data/dsdl_seg', ann_file='set-train/train.yaml')
+ assert len(dataset) == 3
+ assert len(dataset.metainfo['classes']) == 21
+ else:
+ ImportWarning('Package `dsdl` is not installed.')
+
+
+def test_nyu_dataset():
+ dataset = NYUDataset(
+ data_root='tests/data/pseudo_nyu_dataset',
+ data_prefix=dict(img_path='images', depth_map_path='annotations'),
+ )
+ assert len(dataset) == 1
+ data = dataset[0]
+ assert data.get('depth_map_path', None) is not None
+ assert data.get('category_id', -1) == 26
diff --git a/tests/test_datasets/test_loading.py b/tests/test_datasets/test_loading.py
index 5ce624bff6a..3eea6e3f9dd 100644
--- a/tests/test_datasets/test_loading.py
+++ b/tests/test_datasets/test_loading.py
@@ -7,10 +7,11 @@
import numpy as np
from mmcv.transforms import LoadImageFromFile
-from mmseg.datasets.transforms import (LoadAnnotations,
- LoadBiomedicalAnnotation,
+from mmseg.datasets.transforms import LoadAnnotations # noqa
+from mmseg.datasets.transforms import (LoadBiomedicalAnnotation,
LoadBiomedicalData,
LoadBiomedicalImageFromFile,
+ LoadDepthAnnotation,
LoadImageFromNDArray)
@@ -276,3 +277,19 @@ def test_load_biomedical_data(self):
"decode_backend='numpy', "
'to_xyz=False, '
'backend_args=None)')
+
+ def test_load_depth_annotation(self):
+ input_results = dict(
+ img_path='tests/data/pseudo_nyu_dataset/images/'
+ 'bookstore_0001d_00001.jpg',
+ depth_map_path='tests/data/pseudo_nyu_dataset/'
+ 'annotations/bookstore_0001d_00001.png',
+ category_id=-1,
+ seg_fields=[])
+ transform = LoadDepthAnnotation(depth_rescale_factor=0.001)
+ results = transform(input_results)
+ assert 'gt_depth_map' in results
+ assert results['gt_depth_map'].shape[:2] == mmcv.imread(
+ input_results['depth_map_path']).shape[:2]
+ assert results['gt_depth_map'].dtype == np.float32
+ assert 'gt_depth_map' in results['seg_fields']
diff --git a/tests/test_datasets/test_transform.py b/tests/test_datasets/test_transform.py
index 92d6c6106d2..e73e558ee8e 100644
--- a/tests/test_datasets/test_transform.py
+++ b/tests/test_datasets/test_transform.py
@@ -1,6 +1,7 @@
# Copyright (c) OpenMMLab. All rights reserved.
import copy
import os.path as osp
+from unittest import TestCase
import mmcv
import numpy as np
@@ -11,7 +12,8 @@
from mmseg.datasets.transforms import * # noqa
from mmseg.datasets.transforms import (LoadBiomedicalData,
LoadBiomedicalImageFromFile,
- PhotoMetricDistortion, RandomCrop)
+ PhotoMetricDistortion, RandomCrop,
+ RandomDepthMix)
from mmseg.registry import TRANSFORMS
init_default_scope('mmseg')
@@ -184,6 +186,14 @@ def test_flip():
assert np.equal(original_img, results['img']).all()
assert np.equal(original_seg, results['gt_semantic_seg']).all()
+ results['gt_depth_map'] = seg
+ results['seg_fields'] = ['gt_depth_map']
+ results = flip_module(results)
+ flip_module = TRANSFORMS.build(transform)
+ results = flip_module(results)
+ assert np.equal(original_img, results['img']).all()
+ assert np.equal(original_seg, results['gt_depth_map']).all()
+
def test_random_rotate_flip():
with pytest.raises(AssertionError):
@@ -1160,3 +1170,104 @@ def test_biomedical_3d_flip():
results = transform(results)
assert np.equal(original_img, results['img']).all()
assert np.equal(original_seg, results['gt_seg_map']).all()
+
+
+def test_albu_transform():
+ results = dict(
+ img_path=osp.join(osp.dirname(__file__), '../data/color.jpg'))
+
+ # Define simple pipeline
+ load = dict(type='LoadImageFromFile')
+ load = TRANSFORMS.build(load)
+
+ albu_transform = dict(
+ type='Albu', transforms=[dict(type='ChannelShuffle', p=1)])
+ albu_transform = TRANSFORMS.build(albu_transform)
+
+ normalize = dict(type='Normalize', mean=[0] * 3, std=[0] * 3, to_rgb=True)
+ normalize = TRANSFORMS.build(normalize)
+
+ # Execute transforms
+ results = load(results)
+ results = albu_transform(results)
+ results = normalize(results)
+
+ assert results['img'].dtype == np.float32
+
+
+def test_albu_channel_order():
+ results = dict(
+ img_path=osp.join(osp.dirname(__file__), '../data/color.jpg'))
+
+ # Define simple pipeline
+ load = dict(type='LoadImageFromFile')
+ load = TRANSFORMS.build(load)
+
+ # Transform is modifying B channel
+ albu_transform = dict(
+ type='Albu',
+ transforms=[
+ dict(
+ type='RGBShift',
+ r_shift_limit=0,
+ g_shift_limit=0,
+ b_shift_limit=200,
+ p=1)
+ ])
+ albu_transform = TRANSFORMS.build(albu_transform)
+
+ # Execute transforms
+ results_load = load(results)
+ results_albu = albu_transform(results_load)
+
+ # assert only Green and Red channel are not modified
+ np.testing.assert_array_equal(results_albu['img'][..., 1:],
+ results_load['img'][..., 1:])
+
+ # assert Blue channel is modified
+ with pytest.raises(AssertionError):
+ np.testing.assert_array_equal(results_albu['img'][..., 0],
+ results_load['img'][..., 0])
+
+
+class TestRandomDepthMix(TestCase):
+
+ def setUp(self):
+ self.transform = RandomDepthMix(prob=1.0)
+
+ def test_transform_shape(self):
+ # Create a dummy result dict
+ results = {
+ 'img_shape': (10, 10),
+ 'img': np.random.rand(10, 10, 3),
+ 'gt_depth_map': np.random.rand(10, 10)
+ }
+ transformed = self.transform.transform(results)
+
+ # Check if the shape remains the same
+ self.assertEqual(results['img'].shape, transformed['img'].shape)
+
+ def test_transform_values(self):
+ # Create a dummy result dict
+ results = {
+ 'img_shape': (10, 10),
+ 'img': np.zeros((10, 10, 3)),
+ 'gt_depth_map': np.ones((10, 10))
+ }
+ transformed = self.transform.transform(results)
+
+ # Assuming the transformation modifies a portion of the image,
+ # it shouldn't remain all zeros
+ self.assertFalse(np.all(transformed['img'] == 0))
+
+ def test_invalid_image_dimension(self):
+ # Create a dummy result dict with invalid image dimension
+ results = {
+ 'img_shape': (10, 10),
+ 'img': np.random.rand(10, 10, 3, 3),
+ 'gt_depth_map': np.random.rand(10, 10)
+ }
+
+ # Check if a ValueError is raised for invalid dimension
+ with self.assertRaises(ValueError):
+ self.transform.transform(results)
diff --git a/tests/test_evaluation/test_metrics/test_depth_metric.py b/tests/test_evaluation/test_metrics/test_depth_metric.py
new file mode 100644
index 00000000000..a172db8fa20
--- /dev/null
+++ b/tests/test_evaluation/test_metrics/test_depth_metric.py
@@ -0,0 +1,85 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import os.path as osp
+import shutil
+from unittest import TestCase
+
+import torch
+from mmengine.structures import PixelData
+
+from mmseg.evaluation import DepthMetric
+from mmseg.structures import SegDataSample
+
+
+class TestDepthMetric(TestCase):
+
+ def _demo_mm_inputs(self,
+ batch_size=2,
+ image_shapes=(3, 64, 64),
+ num_classes=5):
+ """Create a superset of inputs needed to run test or train batches.
+
+ Args:
+ batch_size (int): batch size. Default to 2.
+ image_shapes (List[tuple], Optional): image shape.
+ Default to (3, 64, 64)
+ num_classes (int): number of different classes.
+ Default to 5.
+ """
+ if isinstance(image_shapes, list):
+ assert len(image_shapes) == batch_size
+ else:
+ image_shapes = [image_shapes] * batch_size
+
+ data_samples = []
+ for idx in range(batch_size):
+ image_shape = image_shapes[idx]
+ _, h, w = image_shape
+
+ data_sample = SegDataSample()
+ gt_depth_map = torch.rand((1, h, w)) * 10
+ data_sample.gt_depth_map = PixelData(data=gt_depth_map)
+
+ data_samples.append(data_sample.to_dict())
+
+ return data_samples
+
+ def _demo_mm_model_output(self,
+ data_samples,
+ batch_size=2,
+ image_shapes=(3, 64, 64),
+ num_classes=5):
+
+ _, h, w = image_shapes
+
+ for data_sample in data_samples:
+ data_sample['pred_depth_map'] = dict(data=torch.randn(1, h, w))
+
+ data_sample[
+ 'img_path'] = 'tests/data/pseudo_dataset/imgs/00000_img.jpg'
+ return data_samples
+
+ def test_evaluate(self):
+ """Test using the metric in the same way as Evalutor."""
+
+ data_samples = self._demo_mm_inputs()
+ data_samples = self._demo_mm_model_output(data_samples)
+
+ depth_metric = DepthMetric()
+ depth_metric.process([0] * len(data_samples), data_samples)
+ res = depth_metric.compute_metrics(depth_metric.results)
+ self.assertIsInstance(res, dict)
+
+ # test save depth map file in output_dir
+ depth_metric = DepthMetric(output_dir='tmp')
+ depth_metric.process([0] * len(data_samples), data_samples)
+ assert osp.exists('tmp')
+ assert osp.isfile('tmp/00000_img.png')
+ shutil.rmtree('tmp')
+
+ # test format_only
+ depth_metric = DepthMetric(output_dir='tmp', format_only=True)
+ depth_metric.process([0] * len(data_samples), data_samples)
+ assert depth_metric.results == []
+ assert osp.exists('tmp')
+ assert osp.isfile('tmp/00000_img.png')
+ shutil.rmtree('tmp')
diff --git a/tests/test_models/test_assigners/test_hungarian_assigner.py b/tests/test_models/test_assigners/test_hungarian_assigner.py
new file mode 100644
index 00000000000..2cdb1de839d
--- /dev/null
+++ b/tests/test_models/test_assigners/test_hungarian_assigner.py
@@ -0,0 +1,77 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from unittest import TestCase
+
+import torch
+from mmengine.structures import InstanceData
+
+from mmseg.models.assigners import HungarianAssigner
+
+
+class TestHungarianAssigner(TestCase):
+
+ def test_init(self):
+ with self.assertRaises(AssertionError):
+ HungarianAssigner([])
+
+ def test_hungarian_match_assigner(self):
+ assigner = HungarianAssigner([
+ dict(type='ClassificationCost', weight=2.0),
+ dict(type='CrossEntropyLossCost', weight=5.0, use_sigmoid=True),
+ dict(type='DiceCost', weight=5.0, pred_act=True, eps=1.0)
+ ])
+ num_classes = 3
+ num_masks = 10
+ num_points = 20
+ gt_instances = InstanceData()
+ gt_instances.labels = torch.randint(0, num_classes, (num_classes, ))
+ gt_instances.masks = torch.randint(0, 2, (num_classes, num_points))
+ pred_instances = InstanceData()
+ pred_instances.scores = torch.rand((num_masks, num_classes))
+ pred_instances.masks = torch.rand((num_masks, num_points))
+
+ matched_quiery_inds, matched_label_inds = \
+ assigner.assign(pred_instances, gt_instances)
+ unique_quiery_inds = torch.unique(matched_quiery_inds)
+ unique_label_inds = torch.unique(matched_label_inds)
+ self.assertTrue(len(unique_quiery_inds) == len(matched_quiery_inds))
+ self.assertTrue(
+ torch.equal(unique_label_inds, torch.arange(0, num_classes)))
+
+ def test_cls_match_cost(self):
+ num_classes = 3
+ num_masks = 10
+ gt_instances = InstanceData()
+ gt_instances.labels = torch.randint(0, num_classes, (num_classes, ))
+ pred_instances = InstanceData()
+ pred_instances.scores = torch.rand((num_masks, num_classes))
+
+ # test ClassificationCost
+ assigner = HungarianAssigner(dict(type='ClassificationCost'))
+ matched_quiery_inds, matched_label_inds = \
+ assigner.assign(pred_instances, gt_instances)
+ unique_quiery_inds = torch.unique(matched_quiery_inds)
+ unique_label_inds = torch.unique(matched_label_inds)
+ self.assertTrue(len(unique_quiery_inds) == len(matched_quiery_inds))
+ self.assertTrue(
+ torch.equal(unique_label_inds, torch.arange(0, num_classes)))
+
+ def test_mask_match_cost(self):
+ num_classes = 3
+ num_masks = 10
+ num_points = 20
+ gt_instances = InstanceData()
+ gt_instances.masks = torch.randint(0, 2, (num_classes, num_points))
+ pred_instances = InstanceData()
+ pred_instances.masks = torch.rand((num_masks, num_points))
+
+ # test DiceCost
+ assigner = HungarianAssigner(
+ dict(type='DiceCost', pred_act=True, eps=1.0))
+ assign_result = assigner.assign(pred_instances, gt_instances)
+ self.assertTrue(len(assign_result[0]) == len(assign_result[1]))
+
+ # test CrossEntropyLossCost
+ assigner = HungarianAssigner(
+ dict(type='CrossEntropyLossCost', use_sigmoid=True))
+ assign_result = assigner.assign(pred_instances, gt_instances)
+ self.assertTrue(len(assign_result[0]) == len(assign_result[1]))
diff --git a/tests/test_models/test_backbones/test_clip_text_encoder.py b/tests/test_models/test_backbones/test_clip_text_encoder.py
new file mode 100644
index 00000000000..ea06c5b5b3f
--- /dev/null
+++ b/tests/test_models/test_backbones/test_clip_text_encoder.py
@@ -0,0 +1,43 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch
+from mmengine import Config
+from mmengine.registry import init_default_scope
+
+from mmseg.models.text_encoder import CLIPTextEncoder
+from mmseg.utils import get_classes
+
+
+def test_clip_text_encoder():
+ init_default_scope('mmseg')
+ # test vocabulary
+ output_dims = 8
+ embed_dims = 32
+ vocabulary = ['cat', 'dog', 'bird', 'car', 'bike']
+ cfg = dict(
+ vocabulary=vocabulary,
+ templates=['a photo of a {}.'],
+ embed_dims=embed_dims,
+ output_dims=output_dims)
+ cfg = Config(cfg)
+
+ text_encoder = CLIPTextEncoder(**cfg)
+ if torch.cuda.is_available():
+ text_encoder = text_encoder.cuda()
+
+ with torch.no_grad():
+ class_embeds = text_encoder()
+ assert class_embeds.shape == (len(vocabulary) + 1, output_dims)
+
+ # test dataset name
+ cfg = dict(
+ dataset_name='vaihingen',
+ templates=['a photo of a {}.'],
+ embed_dims=embed_dims,
+ output_dims=output_dims)
+ cfg = Config(cfg)
+
+ text_encoder = CLIPTextEncoder(**cfg)
+ with torch.no_grad():
+ class_embeds = text_encoder()
+ class_nums = len(get_classes('vaihingen'))
+ assert class_embeds.shape == (class_nums + 1, output_dims)
diff --git a/tests/test_models/test_backbones/test_vpd.py b/tests/test_models/test_backbones/test_vpd.py
new file mode 100644
index 00000000000..a268159155e
--- /dev/null
+++ b/tests/test_models/test_backbones/test_vpd.py
@@ -0,0 +1,51 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from os.path import dirname, join
+from unittest import TestCase
+
+import torch
+from mmengine import Config
+
+import mmseg
+from mmseg.models.backbones import VPD
+
+
+class TestVPD(TestCase):
+
+ def setUp(self) -> None:
+
+ repo_dpath = dirname(dirname(mmseg.__file__))
+ config_dpath = join(repo_dpath, 'configs/_base_/models/vpd_sd.py')
+ vpd_cfg = Config.fromfile(config_dpath).stable_diffusion_cfg
+ vpd_cfg.pop('checkpoint')
+
+ self.vpd_model = VPD(
+ diffusion_cfg=vpd_cfg,
+ class_embed_path='https://download.openmmlab.com/mmsegmentation/'
+ 'v0.5/vpd/nyu_class_embeddings.pth',
+ class_embed_select=True,
+ pad_shape=64,
+ unet_cfg=dict(use_attn=False),
+ )
+
+ def test_forward(self):
+ # test forward without class_id
+ x = torch.randn(1, 3, 60, 60)
+ with torch.no_grad():
+ out = self.vpd_model(x)
+
+ self.assertEqual(len(out), 4)
+ self.assertListEqual(list(out[0].shape), [1, 320, 8, 8])
+ self.assertListEqual(list(out[1].shape), [1, 640, 4, 4])
+ self.assertListEqual(list(out[2].shape), [1, 1280, 2, 2])
+ self.assertListEqual(list(out[3].shape), [1, 1280, 1, 1])
+
+ # test forward with class_id
+ x = torch.randn(1, 3, 60, 60)
+ with torch.no_grad():
+ out = self.vpd_model((x, torch.tensor([2])))
+
+ self.assertEqual(len(out), 4)
+ self.assertListEqual(list(out[0].shape), [1, 320, 8, 8])
+ self.assertListEqual(list(out[1].shape), [1, 640, 4, 4])
+ self.assertListEqual(list(out[2].shape), [1, 1280, 2, 2])
+ self.assertListEqual(list(out[3].shape), [1, 1280, 1, 1])
diff --git a/tests/test_models/test_heads/test_san_head.py b/tests/test_models/test_heads/test_san_head.py
new file mode 100644
index 00000000000..af85a6e2ca0
--- /dev/null
+++ b/tests/test_models/test_heads/test_san_head.py
@@ -0,0 +1,126 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch
+from mmengine import Config
+from mmengine.structures import PixelData
+
+from mmseg.models.decode_heads import SideAdapterCLIPHead
+from mmseg.structures import SegDataSample
+from .utils import list_to_cuda
+
+
+def test_san_head():
+ H, W = (64, 64)
+ clip_channels = 64
+ img_channels = 4
+ num_queries = 40
+ out_dims = 64
+ num_classes = 19
+ cfg = dict(
+ num_classes=num_classes,
+ deep_supervision_idxs=[4],
+ san_cfg=dict(
+ in_channels=img_channels,
+ embed_dims=128,
+ clip_channels=clip_channels,
+ num_queries=num_queries,
+ cfg_encoder=dict(num_encode_layer=4, mlp_ratio=2, num_heads=2),
+ cfg_decoder=dict(
+ num_heads=4,
+ num_layers=1,
+ embed_channels=32,
+ mlp_channels=32,
+ num_mlp=2,
+ rescale=True)),
+ maskgen_cfg=dict(
+ sos_token_num=num_queries,
+ embed_dims=clip_channels,
+ out_dims=out_dims,
+ num_heads=4,
+ mlp_ratio=2),
+ train_cfg=dict(
+ num_points=100,
+ oversample_ratio=3.0,
+ importance_sample_ratio=0.75,
+ assigner=dict(
+ type='HungarianAssigner',
+ match_costs=[
+ dict(type='ClassificationCost', weight=2.0),
+ dict(
+ type='CrossEntropyLossCost',
+ weight=5.0,
+ use_sigmoid=True),
+ dict(type='DiceCost', weight=5.0, pred_act=True, eps=1.0)
+ ])),
+ loss_decode=[
+ dict(
+ type='CrossEntropyLoss',
+ loss_name='loss_cls_ce',
+ loss_weight=2.0,
+ class_weight=[1.0] * num_classes + [0.1]),
+ dict(
+ type='CrossEntropyLoss',
+ use_sigmoid=True,
+ loss_name='loss_mask_ce',
+ loss_weight=5.0),
+ dict(
+ type='DiceLoss',
+ ignore_index=None,
+ naive_dice=True,
+ eps=1,
+ loss_name='loss_mask_dice',
+ loss_weight=5.0)
+ ])
+
+ cfg = Config(cfg)
+ head = SideAdapterCLIPHead(**cfg)
+
+ inputs = torch.rand((2, img_channels, H, W))
+ clip_feature = [[
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ],
+ [
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ],
+ [
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ],
+ [
+ torch.rand((2, clip_channels, H // 2, W // 2)),
+ torch.rand((2, clip_channels))
+ ]]
+ class_embed = torch.rand((num_classes + 1, out_dims))
+
+ data_samples = []
+ for i in range(2):
+ data_sample = SegDataSample()
+ img_meta = {}
+ img_meta['img_shape'] = (H, W)
+ img_meta['ori_shape'] = (H, W)
+ data_sample.gt_sem_seg = PixelData(
+ data=torch.randint(0, num_classes, (1, H, W)))
+ data_sample.set_metainfo(img_meta)
+ data_samples.append(data_sample)
+
+ batch_img_metas = []
+ for data_sample in data_samples:
+ batch_img_metas.append(data_sample.metainfo)
+
+ if torch.cuda.is_available():
+ head = head.cuda()
+ data = list_to_cuda([inputs, clip_feature, class_embed])
+ for data_sample in data_samples:
+ data_sample.gt_sem_seg.data = data_sample.gt_sem_seg.data.cuda()
+ else:
+ data = [inputs, clip_feature, class_embed]
+
+ # loss test
+ loss_dict = head.loss(data, data_samples, None)
+ assert isinstance(loss_dict, dict)
+
+ # prediction test
+ with torch.no_grad():
+ seg_logits = head.predict(data, batch_img_metas, None)
+ assert seg_logits.shape == torch.Size((2, num_classes, H, W))
diff --git a/tests/test_models/test_heads/test_vpd_depth_head.py b/tests/test_models/test_heads/test_vpd_depth_head.py
new file mode 100644
index 00000000000..e3a4f7558ec
--- /dev/null
+++ b/tests/test_models/test_heads/test_vpd_depth_head.py
@@ -0,0 +1,50 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from unittest import TestCase
+
+import torch
+from mmengine.structures import PixelData
+
+from mmseg.models.decode_heads import VPDDepthHead
+from mmseg.structures import SegDataSample
+
+
+class TestVPDDepthHead(TestCase):
+
+ def setUp(self):
+ """Set up common resources."""
+ self.in_channels = [320, 640, 1280, 1280]
+ self.max_depth = 10.0
+ self.loss_decode = dict(
+ type='SiLogLoss'
+ ) # Replace with your actual loss type and parameters
+ self.vpd_depth_head = VPDDepthHead(
+ max_depth=self.max_depth,
+ in_channels=self.in_channels,
+ loss_decode=self.loss_decode)
+
+ def test_forward(self):
+ """Test the forward method."""
+ # Create a mock input tensor. Replace shape as per your needs.
+ x = [
+ torch.randn(1, 320, 32, 32),
+ torch.randn(1, 640, 16, 16),
+ torch.randn(1, 1280, 8, 8),
+ torch.randn(1, 1280, 4, 4)
+ ]
+
+ output = self.vpd_depth_head.forward(x)
+ print(output.shape)
+
+ self.assertEqual(output.shape, (1, 1, 256, 256))
+
+ def test_loss_by_feat(self):
+ """Test the loss_by_feat method."""
+ # Create mock data for `pred_depth_map` and `batch_data_samples`.
+ pred_depth_map = torch.randn(1, 1, 32, 32)
+ gt_depth_map = PixelData(data=torch.rand(1, 32, 32))
+ batch_data_samples = [SegDataSample(gt_depth_map=gt_depth_map)]
+
+ loss = self.vpd_depth_head.loss_by_feat(pred_depth_map,
+ batch_data_samples)
+
+ self.assertIsNotNone(loss)
diff --git a/tests/test_models/test_heads/utils.py b/tests/test_models/test_heads/utils.py
index 335e261a5e5..72823401552 100644
--- a/tests/test_models/test_heads/utils.py
+++ b/tests/test_models/test_heads/utils.py
@@ -20,3 +20,12 @@ def to_cuda(module, data):
for i in range(len(data)):
data[i] = data[i].cuda()
return module, data
+
+
+def list_to_cuda(data):
+ if isinstance(data, list):
+ for i in range(len(data)):
+ data[i] = list_to_cuda(data[i])
+ return data
+ else:
+ return data.cuda()
diff --git a/tests/test_models/test_losses/test_dice_loss.py b/tests/test_models/test_losses/test_dice_loss.py
new file mode 100644
index 00000000000..34253dae12e
--- /dev/null
+++ b/tests/test_models/test_losses/test_dice_loss.py
@@ -0,0 +1,96 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import pytest
+import torch
+
+from mmseg.models.losses import DiceLoss
+
+
+@pytest.mark.parametrize('naive_dice', [True, False])
+def test_dice_loss(naive_dice):
+ loss_class = DiceLoss
+ pred = torch.rand((1, 10, 4, 4))
+ target = torch.randint(0, 10, (1, 4, 4))
+ weight = torch.rand(1)
+ # Test loss forward
+ loss = loss_class(naive_dice=naive_dice)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with weight
+ loss = loss_class(naive_dice=naive_dice)(pred, target, weight)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with reduction_override
+ loss = loss_class(naive_dice=naive_dice)(
+ pred, target, reduction_override='mean')
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with avg_factor
+ loss = loss_class(naive_dice=naive_dice)(pred, target, avg_factor=10)
+ assert isinstance(loss, torch.Tensor)
+
+ with pytest.raises(ValueError):
+ # loss can evaluate with avg_factor only if
+ # reduction is None, 'none' or 'mean'.
+ reduction_override = 'sum'
+ loss_class(naive_dice=naive_dice)(
+ pred, target, avg_factor=10, reduction_override=reduction_override)
+
+ # Test loss forward with avg_factor and reduction
+ for reduction_override in [None, 'none', 'mean']:
+ loss_class(naive_dice=naive_dice)(
+ pred, target, avg_factor=10, reduction_override=reduction_override)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with has_acted=False and use_sigmoid=False
+ for use_sigmoid in [True, False]:
+ loss_class(
+ use_sigmoid=use_sigmoid, activate=True,
+ naive_dice=naive_dice)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with weight.ndim != loss.ndim
+ with pytest.raises(AssertionError):
+ weight = torch.rand((2, 8))
+ loss_class(naive_dice=naive_dice)(pred, target, weight)
+
+ # Test loss forward with len(weight) != len(pred)
+ with pytest.raises(AssertionError):
+ weight = torch.rand(8)
+ loss_class(naive_dice=naive_dice)(pred, target, weight)
+
+ # Test _expand_onehot_labels_dice
+ pred = torch.tensor([[[[1, 1], [1, 0]], [[0, 1], [1, 1]]]]).float()
+ target = torch.tensor([[[0, 0], [0, 1]]])
+ target_onehot = torch.tensor([[[[1, 1], [1, 0]], [[0, 0], [0, 1]]]])
+ weight = torch.rand(1)
+ loss = loss_class(naive_dice=naive_dice)(pred, target, weight)
+ loss_onehot = loss_class(naive_dice=naive_dice)(pred, target_onehot,
+ weight)
+ assert torch.equal(loss, loss_onehot)
+
+ # Test Whether Loss is 0 when pred == target, eps == 0 and naive_dice=False
+ target = torch.randint(0, 2, (1, 10, 4, 4))
+ pred = target.float()
+ target = target.sigmoid()
+ weight = torch.rand(1)
+ loss = loss_class(
+ naive_dice=False, use_sigmoid=True, eps=0)(pred, target, weight)
+ assert loss.item() == 0
+
+ # Test ignore_index when ignore_index is the only class
+ with pytest.raises(AssertionError):
+ pred = torch.ones((1, 1, 4, 4))
+ target = torch.randint(0, 1, (1, 4, 4))
+ weight = torch.rand(1)
+ loss = loss_class(
+ naive_dice=naive_dice, use_sigmoid=False, ignore_index=0,
+ eps=0)(pred, target, weight)
+
+ # Test ignore_index with naive_dice = False
+ pred = torch.tensor([[[[1, 1], [1, 0]], [[0, 1], [1, 1]]]]).float()
+ target = torch.tensor([[[[1, 1], [1, 0]], [[1, 0], [0, 1]]]]).sigmoid()
+ weight = torch.rand(1)
+ loss = loss_class(
+ naive_dice=False, use_sigmoid=True, ignore_index=1,
+ eps=0)(pred, target, weight)
+ assert loss.item() == 0
diff --git a/tests/test_models/test_losses/test_huasdorff_distance_loss.py b/tests/test_models/test_losses/test_huasdorff_distance_loss.py
new file mode 100644
index 00000000000..29c2732d3f1
--- /dev/null
+++ b/tests/test_models/test_losses/test_huasdorff_distance_loss.py
@@ -0,0 +1,29 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import pytest
+import torch
+
+from mmseg.models.losses import HuasdorffDisstanceLoss
+
+
+def test_huasdorff_distance_loss():
+ loss_class = HuasdorffDisstanceLoss
+ pred = torch.rand((10, 8, 6, 6))
+ target = torch.rand((10, 6, 6))
+ class_weight = torch.rand(8)
+
+ # Test loss forward
+ loss = loss_class()(pred, target)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with avg_factor
+ loss = loss_class()(pred, target, avg_factor=10)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with avg_factor and reduction is None, 'sum' and 'mean'
+ for reduction in [None, 'sum', 'mean']:
+ loss = loss_class()(pred, target, avg_factor=10, reduction=reduction)
+ assert isinstance(loss, torch.Tensor)
+
+ # Test loss forward with class_weight
+ with pytest.raises(AssertionError):
+ loss_class(class_weight=class_weight)(pred, target)
diff --git a/tests/test_models/test_losses/test_kldiv_loss.py b/tests/test_models/test_losses/test_kldiv_loss.py
new file mode 100644
index 00000000000..48bcc4bfd9f
--- /dev/null
+++ b/tests/test_models/test_losses/test_kldiv_loss.py
@@ -0,0 +1,40 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import torch
+
+from mmseg.models.losses.kldiv_loss import KLDivLoss
+
+
+def test_kldiv_loss_with_none_reduction():
+ loss_class = KLDivLoss
+ pred = torch.rand((8, 5, 5))
+ target = torch.rand((8, 5, 5))
+ reduction = 'none'
+
+ # Test loss forward
+ loss = loss_class(reduction=reduction)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+ assert loss.shape == (8, 5, 5), f'{loss.shape}'
+
+
+def test_kldiv_loss_with_mean_reduction():
+ loss_class = KLDivLoss
+ pred = torch.rand((8, 5, 5))
+ target = torch.rand((8, 5, 5))
+ reduction = 'mean'
+
+ # Test loss forward
+ loss = loss_class(reduction=reduction)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+ assert loss.shape == (8, ), f'{loss.shape}'
+
+
+def test_kldiv_loss_with_sum_reduction():
+ loss_class = KLDivLoss
+ pred = torch.rand((8, 5, 5))
+ target = torch.rand((8, 5, 5))
+ reduction = 'sum'
+
+ # Test loss forward
+ loss = loss_class(reduction=reduction)(pred, target)
+ assert isinstance(loss, torch.Tensor)
+ assert loss.shape == (8, ), f'{loss.shape}'
diff --git a/tests/test_models/test_losses/test_silog_loss.py b/tests/test_models/test_losses/test_silog_loss.py
new file mode 100644
index 00000000000..022434bcc14
--- /dev/null
+++ b/tests/test_models/test_losses/test_silog_loss.py
@@ -0,0 +1,20 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from unittest import TestCase
+
+import torch
+
+from mmseg.models.losses import SiLogLoss
+
+
+class TestSiLogLoss(TestCase):
+
+ def test_SiLogLoss_forward(self):
+ pred = torch.tensor([[1.0, 2.0], [3.5, 4.0]], dtype=torch.float32)
+ target = torch.tensor([[0.0, 2.0], [3.0, 4.0]], dtype=torch.float32)
+ weight = torch.tensor([1.0, 0.5], dtype=torch.float32)
+
+ loss_module = SiLogLoss()
+ loss = loss_module.forward(pred, target, weight)
+
+ expected_loss = 0.02
+ self.assertAlmostEqual(loss.item(), expected_loss, places=2)
diff --git a/tests/test_models/test_segmentors/test_depth_estimator.py b/tests/test_models/test_segmentors/test_depth_estimator.py
new file mode 100644
index 00000000000..e819c9e7633
--- /dev/null
+++ b/tests/test_models/test_segmentors/test_depth_estimator.py
@@ -0,0 +1,64 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from copy import deepcopy
+from os.path import dirname, join
+from unittest import TestCase
+
+import torch
+from mmengine import Config, ConfigDict
+from mmengine.structures import PixelData
+
+import mmseg
+from mmseg.models.segmentors import DepthEstimator
+from mmseg.structures import SegDataSample
+
+
+class TestDepthEstimator(TestCase):
+
+ def setUp(self) -> None:
+ repo_dpath = dirname(dirname(mmseg.__file__))
+ config_dpath = join(repo_dpath, 'configs/_base_/models/vpd_sd.py')
+ vpd_cfg = Config.fromfile(config_dpath).stable_diffusion_cfg
+ vpd_cfg.pop('checkpoint')
+
+ backbone_cfg = dict(
+ type='VPD',
+ diffusion_cfg=vpd_cfg,
+ class_embed_path='https://download.openmmlab.com/mmsegmentation/'
+ 'v0.5/vpd/nyu_class_embeddings.pth',
+ class_embed_select=True,
+ pad_shape=64,
+ unet_cfg=dict(use_attn=False),
+ )
+
+ head_cfg = dict(
+ type='VPDDepthHead',
+ max_depth=10,
+ )
+
+ self.model = DepthEstimator(
+ backbone=backbone_cfg, decode_head=head_cfg)
+
+ inputs = torch.randn(1, 3, 64, 80)
+ data_sample = SegDataSample()
+ data_sample.gt_depth_map = PixelData(data=torch.rand(1, 64, 80))
+ data_sample.set_metainfo(dict(img_shape=(64, 80), ori_shape=(64, 80)))
+ self.data = dict(inputs=inputs, data_samples=[data_sample])
+
+ def test_slide_flip_inference(self):
+
+ self.model.test_cfg = ConfigDict(
+ dict(mode='slide_flip', crop_size=(64, 64), stride=(16, 16)))
+
+ with torch.no_grad():
+ out = self.model.predict(**deepcopy(self.data))
+
+ self.assertEqual(len(out), 1)
+ self.assertIn('pred_depth_map', out[0].keys())
+ self.assertListEqual(list(out[0].pred_depth_map.shape), [64, 80])
+
+ def test__forward(self):
+ data = deepcopy(self.data)
+ data['inputs'] = data['inputs'][:, :, :64, :64]
+ with torch.no_grad():
+ out = self.model._forward(**data)
+ self.assertListEqual(list(out.shape), [1, 1, 64, 64])
diff --git a/tests/test_models/test_segmentors/test_multimodal_encoder_decoder.py b/tests/test_models/test_segmentors/test_multimodal_encoder_decoder.py
new file mode 100644
index 00000000000..75258d89a7d
--- /dev/null
+++ b/tests/test_models/test_segmentors/test_multimodal_encoder_decoder.py
@@ -0,0 +1,24 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+from mmengine import ConfigDict
+
+from mmseg.models import build_segmentor
+from tests.test_models.test_segmentors.utils import \
+ _segmentor_forward_train_test
+
+
+def test_multimodal_encoder_decoder():
+
+ cfg = ConfigDict(
+ type='MultimodalEncoderDecoder',
+ asymetric_input=False,
+ image_encoder=dict(type='ExampleBackbone', out_indices=[1, 2, 3, 4]),
+ text_encoder=dict(
+ type='ExampleTextEncoder',
+ vocabulary=['A', 'B', 'C'],
+ output_dims=3),
+ decode_head=dict(
+ type='ExampleDecodeHead', out_channels=1, num_classes=2),
+ train_cfg=None,
+ test_cfg=dict(mode='whole'))
+ segmentor = build_segmentor(cfg)
+ _segmentor_forward_train_test(segmentor)
diff --git a/tests/test_models/test_segmentors/test_seg_tta_model.py b/tests/test_models/test_segmentors/test_seg_tta_model.py
index 3c9699e8df4..1e152ed0565 100644
--- a/tests/test_models/test_segmentors/test_seg_tta_model.py
+++ b/tests/test_models/test_segmentors/test_seg_tta_model.py
@@ -1,4 +1,6 @@
# Copyright (c) OpenMMLab. All rights reserved.
+import tempfile
+
import torch
from mmengine import ConfigDict
from mmengine.model import BaseTTAModel
@@ -37,7 +39,8 @@ def test_encoder_decoder_tta():
ori_shape=(10, 10),
img_shape=(10 + i, 10 + i),
flip=(i % 2 == 0),
- flip_direction=flip_direction),
+ flip_direction=flip_direction,
+ img_path=tempfile.mktemp()),
gt_sem_seg=PixelData(data=torch.randint(0, 19, (1, 10, 10))))
])
diff --git a/tests/test_models/test_segmentors/utils.py b/tests/test_models/test_segmentors/utils.py
index 6b440df906d..ac31e2b2774 100644
--- a/tests/test_models/test_segmentors/utils.py
+++ b/tests/test_models/test_segmentors/utils.py
@@ -52,15 +52,22 @@ def _demo_mm_inputs(input_shape=(1, 3, 8, 16), num_classes=10):
@MODELS.register_module()
class ExampleBackbone(nn.Module):
- def __init__(self):
+ def __init__(self, out_indices=None):
super().__init__()
self.conv = nn.Conv2d(3, 3, 3)
+ self.out_indices = out_indices
def init_weights(self, pretrained=None):
pass
def forward(self, x):
- return [self.conv(x)]
+ if self.out_indices is None:
+ return [self.conv(x)]
+ else:
+ outs = []
+ for i in self.out_indices:
+ outs.append(self.conv(x))
+ return outs
@MODELS.register_module()
@@ -74,6 +81,18 @@ def forward(self, inputs):
return self.cls_seg(inputs[0])
+@MODELS.register_module()
+class ExampleTextEncoder(nn.Module):
+
+ def __init__(self, vocabulary=None, output_dims=None):
+ super().__init__()
+ self.vocabulary = vocabulary
+ self.output_dims = output_dims
+
+ def forward(self):
+ return torch.randn((len(self.vocabulary), self.output_dims))
+
+
@MODELS.register_module()
class ExampleCascadeDecodeHead(BaseCascadeDecodeHead):
@@ -132,3 +151,32 @@ def _segmentor_forward_train_test(segmentor):
data_batch = dict(inputs=imgs, data_samples=data_samples)
results = segmentor.forward(imgs, data_samples, mode='tensor')
assert isinstance(results, torch.Tensor)
+
+
+def _segmentor_predict(segmentor):
+ if isinstance(segmentor.decode_head, nn.ModuleList):
+ num_classes = segmentor.decode_head[-1].num_classes
+ else:
+ num_classes = segmentor.decode_head.num_classes
+ # batch_size=2 for BatchNorm
+ mm_inputs = _demo_mm_inputs(num_classes=num_classes)
+
+ # convert to cuda Tensor if applicable
+ if torch.cuda.is_available():
+ segmentor = segmentor.cuda()
+
+ # check data preprocessor
+ if not hasattr(segmentor,
+ 'data_preprocessor') or segmentor.data_preprocessor is None:
+ segmentor.data_preprocessor = SegDataPreProcessor()
+
+ mm_inputs = segmentor.data_preprocessor(mm_inputs, True)
+ imgs = mm_inputs.pop('imgs')
+ data_samples = mm_inputs.pop('data_samples')
+
+ # Test predict
+ with torch.no_grad():
+ segmentor.eval()
+ data_batch = dict(inputs=imgs, data_samples=data_samples)
+ outputs = segmentor.predict(**data_batch)
+ assert isinstance(outputs, list)
diff --git a/tests/test_visualization/test_local_visualizer.py b/tests/test_visualization/test_local_visualizer.py
index b60a9b87507..e3b2a88cfb7 100644
--- a/tests/test_visualization/test_local_visualizer.py
+++ b/tests/test_visualization/test_local_visualizer.py
@@ -155,3 +155,59 @@ def _assert_image_and_shape(self, out_file, out_shape):
assert os.path.exists(out_file)
drawn_img = cv2.imread(out_file)
assert drawn_img.shape == out_shape
+
+ def test_add_datasample_depth(self):
+ h = 10
+ w = 12
+ out_file = 'out_file'
+
+ image = np.random.randint(0, 256, size=(h, w, 3)).astype('uint8')
+
+ # test gt_depth_map
+ gt_depth_map = PixelData(data=torch.rand(1, h, w))
+
+ def test_add_datasample_forward_depth(gt_depth_map):
+ data_sample = SegDataSample()
+ data_sample.gt_depth_map = gt_depth_map
+
+ with tempfile.TemporaryDirectory() as tmp_dir:
+ seg_local_visualizer = SegLocalVisualizer(
+ vis_backends=[dict(type='LocalVisBackend')],
+ save_dir=tmp_dir)
+ seg_local_visualizer.dataset_meta = dict(
+ classes=('background', 'foreground'),
+ palette=[[120, 120, 120], [6, 230, 230]])
+
+ # test out_file
+ seg_local_visualizer.add_datasample(out_file, image,
+ data_sample)
+
+ assert os.path.exists(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'))
+ drawn_img = cv2.imread(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'))
+ assert drawn_img.shape == (h * 2, w, 3)
+
+ # test gt_instances and pred_instances
+
+ pred_depth_map = PixelData(data=torch.rand(1, h, w))
+
+ data_sample.pred_depth_map = pred_depth_map
+
+ seg_local_visualizer.add_datasample(out_file, image,
+ data_sample)
+ self._assert_image_and_shape(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'), (h * 2, w * 2, 3))
+
+ seg_local_visualizer.add_datasample(
+ out_file, image, data_sample, draw_gt=False)
+ self._assert_image_and_shape(
+ osp.join(tmp_dir, 'vis_data', 'vis_image',
+ out_file + '_0.png'), (h * 2, w, 3))
+
+ if torch.cuda.is_available():
+ test_add_datasample_forward_depth(gt_depth_map.cuda())
+ test_add_datasample_forward_depth(gt_depth_map)
diff --git a/tools/analysis_tools/confusion_matrix.py b/tools/analysis_tools/confusion_matrix.py
index 9a87bc14c9b..39756cdfdd2 100644
--- a/tools/analysis_tools/confusion_matrix.py
+++ b/tools/analysis_tools/confusion_matrix.py
@@ -5,10 +5,14 @@
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.ticker import MultipleLocator
-from mmengine import Config, DictAction
-from mmengine.utils import ProgressBar, load
+from mmengine.config import Config, DictAction
+from mmengine.registry import init_default_scope
+from mmengine.utils import mkdir_or_exist, progressbar
+from PIL import Image
-from mmseg.datasets import build_dataset
+from mmseg.registry import DATASETS
+
+init_default_scope('mmseg')
def parse_args():
@@ -16,7 +20,7 @@ def parse_args():
description='Generate confusion matrix from segmentation results')
parser.add_argument('config', help='test config file path')
parser.add_argument(
- 'prediction_path', help='prediction path where test .pkl result')
+ 'prediction_path', help='prediction path where test folder result')
parser.add_argument(
'save_dir', help='directory where confusion matrix will be saved')
parser.add_argument(
@@ -50,15 +54,23 @@ def calculate_confusion_matrix(dataset, results):
dataset (Dataset): Test or val dataset.
results (list[ndarray]): A list of segmentation results in each image.
"""
- n = len(dataset.CLASSES)
+ n = len(dataset.METAINFO['classes'])
confusion_matrix = np.zeros(shape=[n, n])
assert len(dataset) == len(results)
- prog_bar = ProgressBar(len(results))
+ ignore_index = dataset.ignore_index
+ reduce_zero_label = dataset.reduce_zero_label
+ prog_bar = progressbar.ProgressBar(len(results))
for idx, per_img_res in enumerate(results):
res_segm = per_img_res
- gt_segm = dataset.get_gt_seg_map_by_idx(idx)
+ gt_segm = dataset[idx]['data_samples'] \
+ .gt_sem_seg.data.squeeze().numpy().astype(np.uint8)
+ gt_segm, res_segm = gt_segm.flatten(), res_segm.flatten()
+ if reduce_zero_label:
+ gt_segm = gt_segm - 1
+ to_ignore = gt_segm == ignore_index
+
+ gt_segm, res_segm = gt_segm[~to_ignore], res_segm[~to_ignore]
inds = n * gt_segm + res_segm
- inds = inds.flatten()
mat = np.bincount(inds, minlength=n**2).reshape(n, n)
confusion_matrix += mat
prog_bar.update()
@@ -70,7 +82,7 @@ def plot_confusion_matrix(confusion_matrix,
save_dir=None,
show=True,
title='Normalized Confusion Matrix',
- color_theme='winter'):
+ color_theme='OrRd'):
"""Draw confusion matrix with matplotlib.
Args:
@@ -89,14 +101,15 @@ def plot_confusion_matrix(confusion_matrix,
num_classes = len(labels)
fig, ax = plt.subplots(
- figsize=(2 * num_classes, 2 * num_classes * 0.8), dpi=180)
+ figsize=(2 * num_classes, 2 * num_classes * 0.8), dpi=300)
cmap = plt.get_cmap(color_theme)
im = ax.imshow(confusion_matrix, cmap=cmap)
- plt.colorbar(mappable=im, ax=ax)
+ colorbar = plt.colorbar(mappable=im, ax=ax)
+ colorbar.ax.tick_params(labelsize=20) # 设置 colorbar 标签的字体大小
- title_font = {'weight': 'bold', 'size': 12}
+ title_font = {'weight': 'bold', 'size': 20}
ax.set_title(title, fontdict=title_font)
- label_font = {'size': 10}
+ label_font = {'size': 40}
plt.ylabel('Ground Truth Label', fontdict=label_font)
plt.xlabel('Prediction Label', fontdict=label_font)
@@ -116,8 +129,8 @@ def plot_confusion_matrix(confusion_matrix,
# draw label
ax.set_xticks(np.arange(num_classes))
ax.set_yticks(np.arange(num_classes))
- ax.set_xticklabels(labels)
- ax.set_yticklabels(labels)
+ ax.set_xticklabels(labels, fontsize=20)
+ ax.set_yticklabels(labels, fontsize=20)
ax.tick_params(
axis='x', bottom=False, top=True, labelbottom=False, labeltop=True)
@@ -135,13 +148,14 @@ def plot_confusion_matrix(confusion_matrix,
) if not np.isnan(confusion_matrix[i, j]) else -1),
ha='center',
va='center',
- color='w',
- size=7)
+ color='k',
+ size=20)
ax.set_ylim(len(confusion_matrix) - 0.5, -0.5) # matplotlib>3.1.1
fig.tight_layout()
if save_dir is not None:
+ mkdir_or_exist(save_dir)
plt.savefig(
os.path.join(save_dir, 'confusion_matrix.png'), format='png')
if show:
@@ -155,7 +169,12 @@ def main():
if args.cfg_options is not None:
cfg.merge_from_dict(args.cfg_options)
- results = load(args.prediction_path)
+ results = []
+ for img in sorted(os.listdir(args.prediction_path)):
+ img = os.path.join(args.prediction_path, img)
+ image = Image.open(img)
+ image = np.copy(image)
+ results.append(image)
assert isinstance(results, list)
if isinstance(results[0], np.ndarray):
@@ -163,17 +182,11 @@ def main():
else:
raise TypeError('invalid type of prediction results')
- if isinstance(cfg.data.test, dict):
- cfg.data.test.test_mode = True
- elif isinstance(cfg.data.test, list):
- for ds_cfg in cfg.data.test:
- ds_cfg.test_mode = True
-
- dataset = build_dataset(cfg.data.test)
+ dataset = DATASETS.build(cfg.test_dataloader.dataset)
confusion_matrix = calculate_confusion_matrix(dataset, results)
plot_confusion_matrix(
confusion_matrix,
- dataset.CLASSES,
+ dataset.METAINFO['classes'],
save_dir=args.save_dir,
show=args.show,
title=args.title,
diff --git a/tools/analysis_tools/visualization_cam.py b/tools/analysis_tools/visualization_cam.py
new file mode 100644
index 00000000000..00cdb3e04ab
--- /dev/null
+++ b/tools/analysis_tools/visualization_cam.py
@@ -0,0 +1,127 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+"""Use the pytorch-grad-cam tool to visualize Class Activation Maps (CAM).
+
+requirement: pip install grad-cam
+"""
+
+from argparse import ArgumentParser
+
+import numpy as np
+import torch
+import torch.nn.functional as F
+from mmengine import Config
+from mmengine.model import revert_sync_batchnorm
+from PIL import Image
+from pytorch_grad_cam import GradCAM
+from pytorch_grad_cam.utils.image import preprocess_image, show_cam_on_image
+
+from mmseg.apis import inference_model, init_model, show_result_pyplot
+from mmseg.utils import register_all_modules
+
+
+class SemanticSegmentationTarget:
+ """wrap the model.
+
+ requirement: pip install grad-cam
+
+ Args:
+ category (int): Visualization class.
+ mask (ndarray): Mask of class.
+ size (tuple): Image size.
+ """
+
+ def __init__(self, category, mask, size):
+ self.category = category
+ self.mask = torch.from_numpy(mask)
+ self.size = size
+ if torch.cuda.is_available():
+ self.mask = self.mask.cuda()
+
+ def __call__(self, model_output):
+ model_output = torch.unsqueeze(model_output, dim=0)
+ model_output = F.interpolate(
+ model_output, size=self.size, mode='bilinear')
+ model_output = torch.squeeze(model_output, dim=0)
+
+ return (model_output[self.category, :, :] * self.mask).sum()
+
+
+def main():
+ parser = ArgumentParser()
+ parser.add_argument('img', help='Image file')
+ parser.add_argument('config', help='Config file')
+ parser.add_argument('checkpoint', help='Checkpoint file')
+ parser.add_argument(
+ '--out-file',
+ default='prediction.png',
+ help='Path to output prediction file')
+ parser.add_argument(
+ '--cam-file', default='vis_cam.png', help='Path to output cam file')
+ parser.add_argument(
+ '--target-layers',
+ default='backbone.layer4[2]',
+ help='Target layers to visualize CAM')
+ parser.add_argument(
+ '--category-index', default='7', help='Category to visualize CAM')
+ parser.add_argument(
+ '--device', default='cuda:0', help='Device used for inference')
+ args = parser.parse_args()
+
+ # build the model from a config file and a checkpoint file
+ register_all_modules()
+ model = init_model(args.config, args.checkpoint, device=args.device)
+ if args.device == 'cpu':
+ model = revert_sync_batchnorm(model)
+
+ # test a single image
+ result = inference_model(model, args.img)
+
+ # show the results
+ show_result_pyplot(
+ model,
+ args.img,
+ result,
+ draw_gt=False,
+ show=False if args.out_file is not None else True,
+ out_file=args.out_file)
+
+ # result data conversion
+ prediction_data = result.pred_sem_seg.data
+ pre_np_data = prediction_data.cpu().numpy().squeeze(0)
+
+ target_layers = args.target_layers
+ target_layers = [eval(f'model.{target_layers}')]
+
+ category = int(args.category_index)
+ mask_float = np.float32(pre_np_data == category)
+
+ # data processing
+ image = np.array(Image.open(args.img).convert('RGB'))
+ height, width = image.shape[0], image.shape[1]
+ rgb_img = np.float32(image) / 255
+ config = Config.fromfile(args.config)
+ image_mean = config.data_preprocessor['mean']
+ image_std = config.data_preprocessor['std']
+ input_tensor = preprocess_image(
+ rgb_img,
+ mean=[x / 255 for x in image_mean],
+ std=[x / 255 for x in image_std])
+
+ # Grad CAM(Class Activation Maps)
+ # Can also be LayerCAM, XGradCAM, GradCAMPlusPlus, EigenCAM, EigenGradCAM
+ targets = [
+ SemanticSegmentationTarget(category, mask_float, (height, width))
+ ]
+ with GradCAM(
+ model=model,
+ target_layers=target_layers,
+ use_cuda=torch.cuda.is_available()) as cam:
+ grayscale_cam = cam(input_tensor=input_tensor, targets=targets)[0, :]
+ cam_image = show_cam_on_image(rgb_img, grayscale_cam, use_rgb=True)
+
+ # save cam file
+ Image.fromarray(cam_image).save(args.cam_file)
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/dataset_converters/isaid.py b/tools/dataset_converters/isaid.py
index 1da264d975e..1d5ccd9c776 100644
--- a/tools/dataset_converters/isaid.py
+++ b/tools/dataset_converters/isaid.py
@@ -91,7 +91,7 @@ def slide_crop_image(src_path, out_dir, mode, patch_H, patch_W, overlap):
x_end) + '.png'
# print(image)
save_path_image = osp.join(out_dir, 'img_dir', mode, str(image))
- img_patch.save(save_path_image)
+ img_patch.save(save_path_image, format='BMP')
def slide_crop_label(src_path, out_dir, mode, patch_H, patch_W, overlap):
diff --git a/tools/dataset_converters/levircd.py b/tools/dataset_converters/levircd.py
new file mode 100644
index 00000000000..8717f3e856b
--- /dev/null
+++ b/tools/dataset_converters/levircd.py
@@ -0,0 +1,99 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import glob
+import math
+import os
+import os.path as osp
+
+import mmcv
+import numpy as np
+from mmengine.utils import ProgressBar
+
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='Convert levir-cd dataset to mmsegmentation format')
+ parser.add_argument('--dataset_path', help='potsdam folder path')
+ parser.add_argument('-o', '--out_dir', help='output path')
+ parser.add_argument(
+ '--clip_size',
+ type=int,
+ help='clipped size of image after preparation',
+ default=256)
+ parser.add_argument(
+ '--stride_size',
+ type=int,
+ help='stride of clipping original images',
+ default=256)
+ args = parser.parse_args()
+ return args
+
+
+def main():
+ args = parse_args()
+ input_folder = args.dataset_path
+ png_files = glob.glob(
+ os.path.join(input_folder, '**/*.png'), recursive=True)
+ output_folder = args.out_dir
+ prog_bar = ProgressBar(len(png_files))
+ for png_file in png_files:
+ new_path = os.path.join(
+ output_folder,
+ os.path.relpath(os.path.dirname(png_file), input_folder))
+ os.makedirs(os.path.dirname(new_path), exist_ok=True)
+ label = False
+ if 'label' in png_file:
+ label = True
+ clip_big_image(png_file, new_path, args, label)
+ prog_bar.update()
+
+
+def clip_big_image(image_path, clip_save_dir, args, to_label=False):
+ image = mmcv.imread(image_path)
+
+ h, w, c = image.shape
+ clip_size = args.clip_size
+ stride_size = args.stride_size
+
+ num_rows = math.ceil((h - clip_size) / stride_size) if math.ceil(
+ (h - clip_size) /
+ stride_size) * stride_size + clip_size >= h else math.ceil(
+ (h - clip_size) / stride_size) + 1
+ num_cols = math.ceil((w - clip_size) / stride_size) if math.ceil(
+ (w - clip_size) /
+ stride_size) * stride_size + clip_size >= w else math.ceil(
+ (w - clip_size) / stride_size) + 1
+
+ x, y = np.meshgrid(np.arange(num_cols + 1), np.arange(num_rows + 1))
+ xmin = x * clip_size
+ ymin = y * clip_size
+
+ xmin = xmin.ravel()
+ ymin = ymin.ravel()
+ xmin_offset = np.where(xmin + clip_size > w, w - xmin - clip_size,
+ np.zeros_like(xmin))
+ ymin_offset = np.where(ymin + clip_size > h, h - ymin - clip_size,
+ np.zeros_like(ymin))
+ boxes = np.stack([
+ xmin + xmin_offset, ymin + ymin_offset,
+ np.minimum(xmin + clip_size, w),
+ np.minimum(ymin + clip_size, h)
+ ],
+ axis=1)
+
+ if to_label:
+ image[image == 255] = 1
+ image = image[:, :, 0]
+ for box in boxes:
+ start_x, start_y, end_x, end_y = box
+ clipped_image = image[start_y:end_y, start_x:end_x] \
+ if to_label else image[start_y:end_y, start_x:end_x, :]
+ idx = osp.basename(image_path).split('.')[0]
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(clip_save_dir,
+ f'{idx}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/dataset_converters/nyu.py b/tools/dataset_converters/nyu.py
new file mode 100644
index 00000000000..49e09e7af68
--- /dev/null
+++ b/tools/dataset_converters/nyu.py
@@ -0,0 +1,89 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import os.path as osp
+import shutil
+import tempfile
+import zipfile
+
+from mmengine.utils import mkdir_or_exist
+
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='Convert NYU Depth dataset to mmsegmentation format')
+ parser.add_argument('raw_data', help='the path of raw data')
+ parser.add_argument(
+ '-o', '--out_dir', help='output path', default='./data/nyu')
+ args = parser.parse_args()
+ return args
+
+
+def reorganize(raw_data_dir: str, out_dir: str):
+ """Reorganize NYU Depth dataset files into the required directory
+ structure.
+
+ Args:
+ raw_data_dir (str): Path to the raw data directory.
+ out_dir (str): Output directory for the organized dataset.
+ """
+
+ def move_data(data_list, dst_prefix, fname_func):
+ """Move data files from source to destination directory.
+
+ Args:
+ data_list (list): List of data file paths.
+ dst_prefix (str): Prefix to be added to destination paths.
+ fname_func (callable): Function to process file names
+ """
+ for data_item in data_list:
+ data_item = data_item.strip().strip('/')
+ new_item = fname_func(data_item)
+ shutil.move(
+ osp.join(raw_data_dir, data_item),
+ osp.join(out_dir, dst_prefix, new_item))
+
+ def process_phase(phase):
+ """Process a dataset phase (e.g., 'train' or 'test')."""
+ with open(osp.join(raw_data_dir, f'nyu_{phase}.txt')) as f:
+ data = filter(lambda x: len(x.strip()) > 0, f.readlines())
+ data = map(lambda x: x.split()[:2], data)
+ images, annos = zip(*data)
+
+ move_data(images, f'images/{phase}',
+ lambda x: x.replace('/rgb', ''))
+ move_data(annos, f'annotations/{phase}',
+ lambda x: x.replace('/sync_depth', ''))
+
+ process_phase('train')
+ process_phase('test')
+
+
+def main():
+ args = parse_args()
+
+ print('Making directories...')
+ mkdir_or_exist(args.out_dir)
+ for subdir in [
+ 'images/train', 'images/test', 'annotations/train',
+ 'annotations/test'
+ ]:
+ mkdir_or_exist(osp.join(args.out_dir, subdir))
+
+ print('Generating images and annotations...')
+
+ if args.raw_data.endswith('.zip'):
+ with tempfile.TemporaryDirectory() as tmp_dir:
+ zip_file = zipfile.ZipFile(args.raw_data)
+ zip_file.extractall(tmp_dir)
+ reorganize(osp.join(tmp_dir, 'nyu'), args.out_dir)
+ else:
+ assert osp.isdir(
+ args.raw_data
+ ), 'the argument --raw-data should be either a zip file or directory.'
+ reorganize(args.raw_data, args.out_dir)
+
+ print('Done!')
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/misc/publish_model.py b/tools/misc/publish_model.py
index c1bbc9ac1ad..e035ad90e85 100644
--- a/tools/misc/publish_model.py
+++ b/tools/misc/publish_model.py
@@ -1,9 +1,12 @@
# Copyright (c) OpenMMLab. All rights reserved.
import argparse
import subprocess
+from hashlib import sha256
import torch
+BLOCK_SIZE = 128 * 1024
+
def parse_args():
parser = argparse.ArgumentParser(
@@ -14,6 +17,17 @@ def parse_args():
return args
+def sha256sum(filename: str) -> str:
+ """Compute SHA256 message digest from a file."""
+ hash_func = sha256()
+ byte_array = bytearray(BLOCK_SIZE)
+ memory_view = memoryview(byte_array)
+ with open(filename, 'rb', buffering=0) as file:
+ for block in iter(lambda: file.readinto(memory_view), 0):
+ hash_func.update(memory_view[:block])
+ return hash_func.hexdigest()
+
+
def process_checkpoint(in_file, out_file):
checkpoint = torch.load(in_file, map_location='cpu')
# remove optimizer for smaller file size
@@ -22,7 +36,7 @@ def process_checkpoint(in_file, out_file):
# if it is necessary to remove some sensitive data in checkpoint['meta'],
# add the code here.
torch.save(checkpoint, out_file)
- sha = subprocess.check_output(['sha256sum', out_file]).decode()
+ sha = sha256sum(in_file)
final_file = out_file.rstrip('.pth') + f'-{sha[:8]}.pth'
subprocess.Popen(['mv', out_file, final_file])
diff --git a/tools/model_converters/clip2mmseg.py b/tools/model_converters/clip2mmseg.py
new file mode 100644
index 00000000000..9a97e4b04ab
--- /dev/null
+++ b/tools/model_converters/clip2mmseg.py
@@ -0,0 +1,163 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import os.path as osp
+from collections import OrderedDict
+
+import mmengine
+import torch
+from mmengine.runner import CheckpointLoader
+
+
+def convert_vitlayer(paras):
+ new_para_name = ''
+ if paras[0] == 'ln_1':
+ new_para_name = '.'.join(['ln1'] + paras[1:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(['attn.attn'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['ln2'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffn.layers.0.0'] + paras[-1:])
+ else:
+ new_para_name = '.'.join(['ffn.layers.1'] + paras[-1:])
+ else:
+ print(f'Wrong for {paras}')
+ return new_para_name
+
+
+def convert_translayer(paras):
+ new_para_name = ''
+ if paras[0] == 'attn':
+ new_para_name = '.'.join(['attentions.0.attn'] + paras[1:])
+ elif paras[0] == 'ln_1':
+ new_para_name = '.'.join(['norms.0'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['norms.1'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffns.0.layers.0.0'] + paras[2:])
+ elif paras[1] == 'c_proj':
+ new_para_name = '.'.join(['ffns.0.layers.1'] + paras[2:])
+ else:
+ print(f'Wrong for {paras}')
+ else:
+ print(f'Wrong for {paras}')
+ return new_para_name
+
+
+def convert_key_name(ckpt, visual_split):
+ new_ckpt = OrderedDict()
+ for k, v in ckpt.items():
+ key_list = k.split('.')
+ if key_list[0] == 'visual':
+ new_transform_name = 'image_encoder'
+ if key_list[1] == 'class_embedding':
+ new_name = '.'.join([new_transform_name, 'cls_token'])
+ elif key_list[1] == 'positional_embedding':
+ new_name = '.'.join([new_transform_name, 'pos_embed'])
+ elif key_list[1] == 'conv1':
+ new_name = '.'.join([
+ new_transform_name, 'patch_embed.projection', key_list[2]
+ ])
+ elif key_list[1] == 'ln_pre':
+ new_name = '.'.join(
+ [new_transform_name, key_list[1], key_list[2]])
+ elif key_list[1] == 'transformer':
+ new_layer_name = 'layers'
+ layer_index = key_list[3]
+ paras = key_list[4:]
+ if int(layer_index) < visual_split:
+ new_para_name = convert_vitlayer(paras)
+ new_name = '.'.join([
+ new_transform_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ else:
+ new_para_name = convert_translayer(paras)
+ new_transform_name = 'decode_head.rec_with_attnbias'
+ new_layer_name = 'layers'
+ layer_index = str(int(layer_index) - visual_split)
+ new_name = '.'.join([
+ new_transform_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[1] == 'proj':
+ new_name = 'decode_head.rec_with_attnbias.proj.weight'
+ elif key_list[1] == 'ln_post':
+ new_name = k.replace('visual', 'decode_head.rec_with_attnbias')
+ else:
+ print(f'pop parameter: {k}')
+ continue
+ else:
+ text_encoder_name = 'text_encoder'
+ if key_list[0] == 'transformer':
+ layer_name = 'transformer'
+ layer_index = key_list[2]
+ paras = key_list[3:]
+ new_para_name = convert_translayer(paras)
+ new_name = '.'.join([
+ text_encoder_name, layer_name, layer_index, new_para_name
+ ])
+ elif key_list[0] in [
+ 'positional_embedding', 'text_projection', 'bg_embed',
+ 'attn_mask', 'logit_scale', 'token_embedding', 'ln_final'
+ ]:
+ new_name = 'text_encoder.' + k
+ else:
+ print(f'pop parameter: {k}')
+ continue
+ new_ckpt[new_name] = v
+
+ return new_ckpt
+
+
+def convert_tensor(ckpt):
+ cls_token = ckpt['image_encoder.cls_token']
+ new_cls_token = cls_token.unsqueeze(0).unsqueeze(0)
+ ckpt['image_encoder.cls_token'] = new_cls_token
+ pos_embed = ckpt['image_encoder.pos_embed']
+ new_pos_embed = pos_embed.unsqueeze(0)
+ ckpt['image_encoder.pos_embed'] = new_pos_embed
+ proj_weight = ckpt['decode_head.rec_with_attnbias.proj.weight']
+ new_proj_weight = proj_weight.transpose(1, 0)
+ ckpt['decode_head.rec_with_attnbias.proj.weight'] = new_proj_weight
+ return ckpt
+
+
+def main():
+ parser = argparse.ArgumentParser(
+ description='Convert keys in timm pretrained vit models to '
+ 'MMSegmentation style.')
+ parser.add_argument('src', help='src model path or url')
+ # The dst path must be a full path of the new checkpoint.
+ parser.add_argument('dst', help='save path')
+ args = parser.parse_args()
+
+ if any([s in args.src for s in ['B-16', 'b16', 'base_patch16']]):
+ visual_split = 9
+ elif any([s in args.src for s in ['L-14', 'l14', 'large_patch14']]):
+ visual_split = 18
+ else:
+ print('Make sure the clip model is ViT-B/16 or ViT-L/14!')
+ visual_split = -1
+ checkpoint = CheckpointLoader.load_checkpoint(args.src, map_location='cpu')
+ if isinstance(checkpoint, torch.jit.RecursiveScriptModule):
+ state_dict = checkpoint.state_dict()
+ else:
+ if 'state_dict' in checkpoint:
+ # timm checkpoint
+ state_dict = checkpoint['state_dict']
+ elif 'model' in checkpoint:
+ # deit checkpoint
+ state_dict = checkpoint['model']
+ else:
+ state_dict = checkpoint
+ weight = convert_key_name(state_dict, visual_split)
+ weight = convert_tensor(weight)
+ mmengine.mkdir_or_exist(osp.dirname(args.dst))
+ torch.save(weight, args.dst)
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/model_converters/san2mmseg.py b/tools/model_converters/san2mmseg.py
new file mode 100644
index 00000000000..301a46608e0
--- /dev/null
+++ b/tools/model_converters/san2mmseg.py
@@ -0,0 +1,220 @@
+# Copyright (c) OpenMMLab. All rights reserved.
+import argparse
+import os.path as osp
+from collections import OrderedDict
+
+import mmengine
+import torch
+from mmengine.runner import CheckpointLoader
+
+
+def convert_key_name(ckpt):
+ new_ckpt = OrderedDict()
+
+ for k, v in ckpt.items():
+ key_list = k.split('.')
+ if key_list[0] == 'clip_visual_extractor':
+ new_transform_name = 'image_encoder'
+ if key_list[1] == 'class_embedding':
+ new_name = '.'.join([new_transform_name, 'cls_token'])
+ elif key_list[1] == 'positional_embedding':
+ new_name = '.'.join([new_transform_name, 'pos_embed'])
+ elif key_list[1] == 'conv1':
+ new_name = '.'.join([
+ new_transform_name, 'patch_embed.projection', key_list[2]
+ ])
+ elif key_list[1] == 'ln_pre':
+ new_name = '.'.join(
+ [new_transform_name, key_list[1], key_list[2]])
+ elif key_list[1] == 'resblocks':
+ new_layer_name = 'layers'
+ layer_index = key_list[2]
+ paras = key_list[3:]
+ if paras[0] == 'ln_1':
+ new_para_name = '.'.join(['ln1'] + key_list[4:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(['attn.attn'] + key_list[4:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['ln2'] + key_list[4:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffn.layers.0.0'] +
+ key_list[-1:])
+ else:
+ new_para_name = '.'.join(['ffn.layers.1'] +
+ key_list[-1:])
+ new_name = '.'.join([
+ new_transform_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[0] == 'side_adapter_network':
+ decode_head_name = 'decode_head'
+ module_name = 'side_adapter_network'
+ if key_list[1] == 'vit_model':
+ if key_list[2] == 'blocks':
+ layer_name = 'encode_layers'
+ layer_index = key_list[3]
+ paras = key_list[4:]
+ if paras[0] == 'norm1':
+ new_para_name = '.'.join(['ln1'] + key_list[5:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(key_list[4:])
+ new_para_name = new_para_name.replace(
+ 'attn.qkv.', 'attn.attn.in_proj_')
+ new_para_name = new_para_name.replace(
+ 'attn.proj', 'attn.attn.out_proj')
+ elif paras[0] == 'norm2':
+ new_para_name = '.'.join(['ln2'] + key_list[5:])
+ elif paras[0] == 'mlp':
+ new_para_name = '.'.join(['ffn'] + key_list[5:])
+ new_para_name = new_para_name.replace(
+ 'fc1', 'layers.0.0')
+ new_para_name = new_para_name.replace(
+ 'fc2', 'layers.1')
+ else:
+ print(f'Wrong for {k}')
+ new_name = '.'.join([
+ decode_head_name, module_name, layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[2] == 'pos_embed':
+ new_name = '.'.join(
+ [decode_head_name, module_name, 'pos_embed'])
+ elif key_list[2] == 'patch_embed':
+ new_name = '.'.join([
+ decode_head_name, module_name, 'patch_embed',
+ 'projection', key_list[4]
+ ])
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[1] == 'query_embed' or key_list[
+ 1] == 'query_pos_embed':
+ new_name = '.'.join(
+ [decode_head_name, module_name, key_list[1]])
+ elif key_list[1] == 'fusion_layers':
+ layer_name = 'conv_clips'
+ layer_index = key_list[2][-1]
+ paras = '.'.join(key_list[3:])
+ new_para_name = paras.replace('input_proj.0', '0')
+ new_para_name = new_para_name.replace('input_proj.1', '1.conv')
+ new_name = '.'.join([
+ decode_head_name, module_name, layer_name, layer_index,
+ new_para_name
+ ])
+ elif key_list[1] == 'mask_decoder':
+ new_name = 'decode_head.' + k
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[0] == 'clip_rec_head':
+ module_name = 'rec_with_attnbias'
+ if key_list[1] == 'proj':
+ new_name = '.'.join(
+ [decode_head_name, module_name, 'proj.weight'])
+ elif key_list[1] == 'ln_post':
+ new_name = '.'.join(
+ [decode_head_name, module_name, 'ln_post', key_list[2]])
+ elif key_list[1] == 'resblocks':
+ new_layer_name = 'layers'
+ layer_index = key_list[2]
+ paras = key_list[3:]
+ if paras[0] == 'ln_1':
+ new_para_name = '.'.join(['norms.0'] + paras[1:])
+ elif paras[0] == 'attn':
+ new_para_name = '.'.join(['attentions.0.attn'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['norms.1'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffns.0.layers.0.0'] +
+ paras[2:])
+ elif paras[1] == 'c_proj':
+ new_para_name = '.'.join(['ffns.0.layers.1'] +
+ paras[2:])
+ else:
+ print(f'Wrong for {k}')
+ new_name = '.'.join([
+ decode_head_name, module_name, new_layer_name, layer_index,
+ new_para_name
+ ])
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[0] == 'ov_classifier':
+ text_encoder_name = 'text_encoder'
+ if key_list[1] == 'transformer':
+ layer_name = 'transformer'
+ layer_index = key_list[3]
+ paras = key_list[4:]
+ if paras[0] == 'attn':
+ new_para_name = '.'.join(['attentions.0.attn'] + paras[1:])
+ elif paras[0] == 'ln_1':
+ new_para_name = '.'.join(['norms.0'] + paras[1:])
+ elif paras[0] == 'ln_2':
+ new_para_name = '.'.join(['norms.1'] + paras[1:])
+ elif paras[0] == 'mlp':
+ if paras[1] == 'c_fc':
+ new_para_name = '.'.join(['ffns.0.layers.0.0'] +
+ paras[2:])
+ elif paras[1] == 'c_proj':
+ new_para_name = '.'.join(['ffns.0.layers.1'] +
+ paras[2:])
+ else:
+ print(f'Wrong for {k}')
+ else:
+ print(f'Wrong for {k}')
+ new_name = '.'.join([
+ text_encoder_name, layer_name, layer_index, new_para_name
+ ])
+ elif key_list[1] in [
+ 'positional_embedding', 'text_projection', 'bg_embed',
+ 'attn_mask', 'logit_scale', 'token_embedding', 'ln_final'
+ ]:
+ new_name = k.replace('ov_classifier', 'text_encoder')
+ else:
+ print(f'Wrong for {k}')
+ elif key_list[0] == 'criterion':
+ new_name = k
+ else:
+ print(f'Wrong for {k}')
+ new_ckpt[new_name] = v
+ return new_ckpt
+
+
+def convert_tensor(ckpt):
+ cls_token = ckpt['image_encoder.cls_token']
+ new_cls_token = cls_token.unsqueeze(0).unsqueeze(0)
+ ckpt['image_encoder.cls_token'] = new_cls_token
+ pos_embed = ckpt['image_encoder.pos_embed']
+ new_pos_embed = pos_embed.unsqueeze(0)
+ ckpt['image_encoder.pos_embed'] = new_pos_embed
+ proj_weight = ckpt['decode_head.rec_with_attnbias.proj.weight']
+ new_proj_weight = proj_weight.transpose(1, 0)
+ ckpt['decode_head.rec_with_attnbias.proj.weight'] = new_proj_weight
+ return ckpt
+
+
+def main():
+ parser = argparse.ArgumentParser(
+ description='Convert keys in timm pretrained vit models to '
+ 'MMSegmentation style.')
+ parser.add_argument('src', help='src model path or url')
+ # The dst path must be a full path of the new checkpoint.
+ parser.add_argument('dst', help='save path')
+ args = parser.parse_args()
+
+ checkpoint = CheckpointLoader.load_checkpoint(args.src, map_location='cpu')
+ if 'state_dict' in checkpoint:
+ # timm checkpoint
+ state_dict = checkpoint['state_dict']
+ elif 'model' in checkpoint:
+ # deit checkpoint
+ state_dict = checkpoint['model']
+ else:
+ state_dict = checkpoint
+ weight = convert_key_name(state_dict)
+ weight = convert_tensor(weight)
+ mmengine.mkdir_or_exist(osp.dirname(args.dst))
+ torch.save(weight, args.dst)
+
+
+if __name__ == '__main__':
+ main()
diff --git a/tools/test.py b/tools/test.py
index 64da2bc2616..0d7f39b3a8b 100644
--- a/tools/test.py
+++ b/tools/test.py
@@ -47,7 +47,10 @@ def parse_args():
help='job launcher')
parser.add_argument(
'--tta', action='store_true', help='Test time augmentation')
- parser.add_argument('--local_rank', type=int, default=0)
+ # When using PyTorch version >= 2.0.0, the `torch.distributed.launch`
+ # will pass the `--local-rank` parameter to `tools/train.py` instead
+ # of `--local_rank`.
+ parser.add_argument('--local_rank', '--local-rank', type=int, default=0)
args = parser.parse_args()
if 'LOCAL_RANK' not in os.environ:
os.environ['LOCAL_RANK'] = str(args.local_rank)
@@ -65,8 +68,8 @@ def trigger_visualization_hook(cfg, args):
visualization_hook['show'] = True
visualization_hook['wait_time'] = args.wait_time
if args.show_dir:
- visulizer = cfg.visualizer
- visulizer['save_dir'] = args.show_dir
+ visualizer = cfg.visualizer
+ visualizer['save_dir'] = args.show_dir
else:
raise RuntimeError(
'VisualizationHook must be included in default_hooks.'
diff --git a/tools/train.py b/tools/train.py
index 17213066648..10fdaa1874b 100644
--- a/tools/train.py
+++ b/tools/train.py
@@ -40,7 +40,10 @@ def parse_args():
choices=['none', 'pytorch', 'slurm', 'mpi'],
default='none',
help='job launcher')
- parser.add_argument('--local_rank', type=int, default=0)
+ # When using PyTorch version >= 2.0.0, the `torch.distributed.launch`
+ # will pass the `--local-rank` parameter to `tools/train.py` instead
+ # of `--local_rank`.
+ parser.add_argument('--local_rank', '--local-rank', type=int, default=0)
args = parser.parse_args()
if 'LOCAL_RANK' not in os.environ:
os.environ['LOCAL_RANK'] = str(args.local_rank)
 +
+ 

 @@ -45,6 +46,12 @@ English | [简体中文](README_zh-CN.md)
@@ -45,6 +46,12 @@ English | [简体中文](README_zh-CN.md)

 +
+  +
+
+
+  +
+  +
+
+
+ 


 -
- 
 +
我们会在 OpenMMLab 社区为大家
diff --git a/configs/_base_/datasets/bdd100k.py b/configs/_base_/datasets/bdd100k.py
new file mode 100644
index 00000000000..24cec69bfeb
--- /dev/null
+++ b/configs/_base_/datasets/bdd100k.py
@@ -0,0 +1,70 @@
+# dataset settings
+dataset_type = 'BDD100KDataset'
+data_root = 'data/bdd100k/'
+
+crop_size = (512, 1024)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(
+ type='RandomResize',
+ scale=(2048, 1024),
+ ratio_range=(0.5, 2.0),
+ keep_ratio=True),
+ dict(type='RandomCrop', crop_size=crop_size, cat_max_ratio=0.75),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 1024), keep_ratio=True),
+ # add loading annotation after ``Resize`` because ground truth
+ # does not need to do resize data transform
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+img_ratios = [0.5, 0.75, 1.0, 1.25, 1.5, 1.75]
+tta_pipeline = [
+ dict(type='LoadImageFromFile', backend_args=None),
+ dict(
+ type='TestTimeAug',
+ transforms=[
+ [
+ dict(type='Resize', scale_factor=r, keep_ratio=True)
+ for r in img_ratios
+ ],
+ [
+ dict(type='RandomFlip', prob=0., direction='horizontal'),
+ dict(type='RandomFlip', prob=1., direction='horizontal')
+ ], [dict(type='LoadAnnotations')], [dict(type='PackSegInputs')]
+ ])
+]
+train_dataloader = dict(
+ batch_size=2,
+ num_workers=2,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/10k/train',
+ seg_map_path='labels/sem_seg/masks/train'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/10k/val',
+ seg_map_path='labels/sem_seg/masks/val'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU'])
+test_evaluator = val_evaluator
diff --git a/configs/_base_/datasets/coco-stuff164k.py b/configs/_base_/datasets/coco-stuff164k.py
index baf633f9d6f..a9b9d90117a 100644
--- a/configs/_base_/datasets/coco-stuff164k.py
+++ b/configs/_base_/datasets/coco-stuff164k.py
@@ -48,7 +48,7 @@
type=dataset_type,
data_root=data_root,
data_prefix=dict(
- img_path='images/train2017', seg_map_path='annotations/val2017'),
+ img_path='images/train2017', seg_map_path='annotations/train2017'),
pipeline=train_pipeline))
val_dataloader = dict(
batch_size=1,
diff --git a/configs/_base_/datasets/levir_256x256.py b/configs/_base_/datasets/levir_256x256.py
new file mode 100644
index 00000000000..a2a69aa9e9c
--- /dev/null
+++ b/configs/_base_/datasets/levir_256x256.py
@@ -0,0 +1,59 @@
+# dataset settings
+dataset_type = 'LEVIRCDDataset'
+data_root = r'data/LEVIRCD'
+
+albu_train_transforms = [
+ dict(type='RandomBrightnessContrast', p=0.2),
+ dict(type='HorizontalFlip', p=0.5),
+ dict(type='VerticalFlip', p=0.5)
+]
+
+train_pipeline = [
+ dict(type='LoadMultipleRSImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Albu', transforms=albu_train_transforms),
+ dict(type='ConcatCDInput'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadMultipleRSImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='ConcatCDInput'),
+ dict(type='PackSegInputs')
+]
+
+tta_pipeline = [
+ dict(type='LoadMultipleRSImageFromFile'),
+ dict(
+ type='TestTimeAug',
+ transforms=[[dict(type='LoadAnnotations')],
+ [dict(type='ConcatCDInput')],
+ [dict(type='PackSegInputs')]])
+]
+train_dataloader = dict(
+ batch_size=4,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='train/A',
+ img_path2='train/B',
+ seg_map_path='train/label'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='test/A', img_path2='test/B', seg_map_path='test/label'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU'])
+test_evaluator = val_evaluator
diff --git a/configs/_base_/datasets/nyu.py b/configs/_base_/datasets/nyu.py
new file mode 100644
index 00000000000..74d57c5fc50
--- /dev/null
+++ b/configs/_base_/datasets/nyu.py
@@ -0,0 +1,67 @@
+# dataset settings
+dataset_type = 'NYUDataset'
+data_root = 'data/nyu'
+
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3),
+ dict(type='RandomDepthMix', prob=0.25),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='RandomCrop', crop_size=(480, 480)),
+ dict(
+ type='Albu',
+ transforms=[
+ dict(type='RandomBrightnessContrast'),
+ dict(type='RandomGamma'),
+ dict(type='HueSaturationValue'),
+ ]),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id')),
+]
+
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2000, 480), keep_ratio=True),
+ dict(dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3)),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id'))
+]
+
+train_dataloader = dict(
+ batch_size=8,
+ num_workers=8,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/train', depth_map_path='annotations/train'),
+ pipeline=train_pipeline))
+
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ test_mode=True,
+ data_prefix=dict(
+ img_path='images/test', depth_map_path='annotations/test'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(
+ type='DepthMetric',
+ min_depth_eval=0.001,
+ max_depth_eval=10.0,
+ crop_type='nyu_crop')
+test_evaluator = val_evaluator
diff --git a/configs/_base_/datasets/nyu_512x512.py b/configs/_base_/datasets/nyu_512x512.py
new file mode 100644
index 00000000000..88e3878d334
--- /dev/null
+++ b/configs/_base_/datasets/nyu_512x512.py
@@ -0,0 +1,72 @@
+# dataset settings
+dataset_type = 'NYUDataset'
+data_root = 'data/nyu'
+
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3),
+ dict(type='RandomDepthMix', prob=0.25),
+ dict(type='RandomFlip', prob=0.5),
+ dict(
+ type='RandomResize',
+ scale=(768, 512),
+ ratio_range=(0.8, 1.5),
+ keep_ratio=True),
+ dict(type='RandomCrop', crop_size=(512, 512)),
+ dict(
+ type='Albu',
+ transforms=[
+ dict(type='RandomBrightnessContrast'),
+ dict(type='RandomGamma'),
+ dict(type='HueSaturationValue'),
+ ]),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id')),
+]
+
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 512), keep_ratio=True),
+ dict(dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3)),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id'))
+]
+
+train_dataloader = dict(
+ batch_size=8,
+ num_workers=8,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/train', depth_map_path='annotations/train'),
+ pipeline=train_pipeline))
+
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ test_mode=True,
+ data_prefix=dict(
+ img_path='images/test', depth_map_path='annotations/test'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(
+ type='DepthMetric',
+ min_depth_eval=0.001,
+ max_depth_eval=10.0,
+ crop_type='nyu_crop')
+test_evaluator = val_evaluator
diff --git a/configs/_base_/models/fpn_poolformer_s12.py b/configs/_base_/models/fpn_poolformer_s12.py
index b6893f69774..086c804837a 100644
--- a/configs/_base_/models/fpn_poolformer_s12.py
+++ b/configs/_base_/models/fpn_poolformer_s12.py
@@ -1,10 +1,11 @@
# model settings
norm_cfg = dict(type='SyncBN', requires_grad=True)
checkpoint_file = 'https://download.openmmlab.com/mmclassification/v0/poolformer/poolformer-s12_3rdparty_32xb128_in1k_20220414-f8d83051.pth' # noqa
-# TODO: delete custom_imports after mmcls supports auto import
-# please install mmcls>=1.0
-# import mmcls.models to trigger register_module in mmcls
-custom_imports = dict(imports=['mmcls.models'], allow_failed_imports=False)
+# TODO: delete custom_imports after mmpretrain supports auto import
+# please install mmpretrain >= 1.0.0rc7
+# import mmpretrain.models to trigger register_module in mmpretrain
+custom_imports = dict(
+ imports=['mmpretrain.models'], allow_failed_imports=False)
data_preprocessor = dict(
type='SegDataPreProcessor',
mean=[123.675, 116.28, 103.53],
@@ -16,7 +17,7 @@
type='EncoderDecoder',
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.PoolFormer',
+ type='mmpretrain.PoolFormer',
arch='s12',
init_cfg=dict(
type='Pretrained', checkpoint=checkpoint_file, prefix='backbone.'),
diff --git a/configs/_base_/models/san_vit-b16.py b/configs/_base_/models/san_vit-b16.py
new file mode 100644
index 00000000000..96ac41b8dad
--- /dev/null
+++ b/configs/_base_/models/san_vit-b16.py
@@ -0,0 +1,137 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[122.7709, 116.7460, 104.0937],
+ std=[68.5005, 66.6322, 70.3232],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255,
+ size_divisor=640,
+ test_cfg=dict(size_divisor=32))
+
+num_classes = 171
+model = dict(
+ type='MultimodalEncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained='pretrain/clip_vit_base_patch16_224.pth',
+ asymetric_input=True,
+ encoder_resolution=0.5,
+ image_encoder=dict(
+ type='VisionTransformer',
+ img_size=(224, 224),
+ patch_size=16,
+ patch_pad=0,
+ in_channels=3,
+ embed_dims=768,
+ num_layers=9,
+ num_heads=12,
+ mlp_ratio=4,
+ out_origin=True,
+ out_indices=(2, 5, 8),
+ qkv_bias=True,
+ drop_rate=0.0,
+ attn_drop_rate=0.0,
+ drop_path_rate=0.0,
+ with_cls_token=True,
+ output_cls_token=True,
+ patch_bias=False,
+ pre_norm=True,
+ norm_cfg=dict(type='LN', eps=1e-5),
+ act_cfg=dict(type='QuickGELU'),
+ norm_eval=False,
+ interpolate_mode='bicubic',
+ frozen_exclude=['pos_embed']),
+ text_encoder=dict(
+ type='CLIPTextEncoder',
+ dataset_name=None,
+ templates='vild',
+ embed_dims=512,
+ num_layers=12,
+ num_heads=8,
+ mlp_ratio=4,
+ output_dims=512,
+ cache_feature=True,
+ cat_bg=True,
+ norm_cfg=dict(type='LN', eps=1e-5)
+ ),
+ decode_head=dict(
+ type='SideAdapterCLIPHead',
+ num_classes=num_classes,
+ deep_supervision_idxs=[7],
+ san_cfg=dict(
+ in_channels=3,
+ clip_channels=768,
+ embed_dims=240,
+ patch_size=16,
+ patch_bias=True,
+ num_queries=100,
+ cfg_encoder=dict(
+ num_encode_layer=8,
+ num_heads=6,
+ mlp_ratio=4
+ ),
+ fusion_index=[0, 1, 2, 3],
+ cfg_decoder=dict(
+ num_heads=12,
+ num_layers=1,
+ embed_channels=256,
+ mlp_channels=256,
+ num_mlp=3,
+ rescale=True),
+ norm_cfg=dict(type='LN', eps=1e-6),
+ ),
+ maskgen_cfg=dict(
+ sos_token_format='cls_token',
+ sos_token_num=100,
+ cross_attn=False,
+ num_layers=3,
+ embed_dims=768,
+ num_heads=12,
+ mlp_ratio=4,
+ qkv_bias=True,
+ out_dims=512,
+ final_norm=True,
+ act_cfg=dict(type='QuickGELU'),
+ norm_cfg=dict(type='LN', eps=1e-5),
+ frozen_exclude=[]
+ ),
+ align_corners=False,
+ train_cfg=dict(
+ num_points=12544,
+ oversample_ratio=3.0,
+ importance_sample_ratio=0.75,
+ assigner=dict(
+ type='HungarianAssigner',
+ match_costs=[
+ dict(type='ClassificationCost', weight=2.0),
+ dict(
+ type='CrossEntropyLossCost',
+ weight=5.0,
+ use_sigmoid=True),
+ dict(
+ type='DiceCost',
+ weight=5.0,
+ pred_act=True,
+ eps=1.0)
+ ])),
+ loss_decode=[dict(type='CrossEntropyLoss',
+ loss_name='loss_cls_ce',
+ loss_weight=2.0,
+ class_weight=[1.0] * num_classes + [0.1]),
+ dict(type='CrossEntropyLoss',
+ use_sigmoid=True,
+ loss_name='loss_mask_ce',
+ loss_weight=5.0),
+ dict(type='DiceLoss',
+ ignore_index=None,
+ naive_dice=True,
+ eps=1,
+ loss_name='loss_mask_dice',
+ loss_weight=5.0)
+ ]),
+
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole')) # yapf: disable
diff --git a/configs/_base_/models/upernet_convnext.py b/configs/_base_/models/upernet_convnext.py
index 7595295871d..958994c91e6 100644
--- a/configs/_base_/models/upernet_convnext.py
+++ b/configs/_base_/models/upernet_convnext.py
@@ -1,5 +1,5 @@
norm_cfg = dict(type='SyncBN', requires_grad=True)
-custom_imports = dict(imports='mmcls.models', allow_failed_imports=False)
+custom_imports = dict(imports='mmpretrain.models', allow_failed_imports=False)
checkpoint_file = 'https://download.openmmlab.com/mmclassification/v0/convnext/downstream/convnext-base_3rdparty_32xb128-noema_in1k_20220301-2a0ee547.pth' # noqa
data_preprocessor = dict(
type='SegDataPreProcessor',
@@ -13,7 +13,7 @@
data_preprocessor=data_preprocessor,
pretrained=None,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='base',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/_base_/models/vpd_sd.py b/configs/_base_/models/vpd_sd.py
new file mode 100644
index 00000000000..87321e74f04
--- /dev/null
+++ b/configs/_base_/models/vpd_sd.py
@@ -0,0 +1,86 @@
+# model settings
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[127.5, 127.5, 127.5],
+ std=[127.5, 127.5, 127.5],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=0)
+
+# adapted from stable-diffusion/configs/stable-diffusion/v1-inference.yaml
+stable_diffusion_cfg = dict(
+ base_learning_rate=0.0001,
+ target='ldm.models.diffusion.ddpm.LatentDiffusion',
+ checkpoint='https://download.openmmlab.com/mmsegmentation/v0.5/'
+ 'vpd/stable_diffusion_v1-5_pretrain_third_party.pth',
+ params=dict(
+ linear_start=0.00085,
+ linear_end=0.012,
+ num_timesteps_cond=1,
+ log_every_t=200,
+ timesteps=1000,
+ first_stage_key='jpg',
+ cond_stage_key='txt',
+ image_size=64,
+ channels=4,
+ cond_stage_trainable=False,
+ conditioning_key='crossattn',
+ monitor='val/loss_simple_ema',
+ scale_factor=0.18215,
+ use_ema=False,
+ scheduler_config=dict(
+ target='ldm.lr_scheduler.LambdaLinearScheduler',
+ params=dict(
+ warm_up_steps=[10000],
+ cycle_lengths=[10000000000000],
+ f_start=[1e-06],
+ f_max=[1.0],
+ f_min=[1.0])),
+ unet_config=dict(
+ target='ldm.modules.diffusionmodules.openaimodel.UNetModel',
+ params=dict(
+ image_size=32,
+ in_channels=4,
+ out_channels=4,
+ model_channels=320,
+ attention_resolutions=[4, 2, 1],
+ num_res_blocks=2,
+ channel_mult=[1, 2, 4, 4],
+ num_heads=8,
+ use_spatial_transformer=True,
+ transformer_depth=1,
+ context_dim=768,
+ use_checkpoint=True,
+ legacy=False)),
+ first_stage_config=dict(
+ target='ldm.models.autoencoder.AutoencoderKL',
+ params=dict(
+ embed_dim=4,
+ monitor='val/rec_loss',
+ ddconfig=dict(
+ double_z=True,
+ z_channels=4,
+ resolution=256,
+ in_channels=3,
+ out_ch=3,
+ ch=128,
+ ch_mult=[1, 2, 4, 4],
+ num_res_blocks=2,
+ attn_resolutions=[],
+ dropout=0.0),
+ lossconfig=dict(target='torch.nn.Identity'))),
+ cond_stage_config=dict(
+ target='ldm.modules.encoders.modules.AbstractEncoder')))
+
+model = dict(
+ type='DepthEstimator',
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ type='VPD',
+ diffusion_cfg=stable_diffusion_cfg,
+ ),
+)
+
+# some of the parameters in stable-diffusion model will not be updated
+# during training
+find_unused_parameters = True
diff --git a/configs/_base_/schedules/schedule_25k.py b/configs/_base_/schedules/schedule_25k.py
new file mode 100644
index 00000000000..825e141ed12
--- /dev/null
+++ b/configs/_base_/schedules/schedule_25k.py
@@ -0,0 +1,28 @@
+# optimizer
+optimizer = dict(type='AdamW', lr=0.001, weight_decay=0.1)
+optim_wrapper = dict(type='OptimWrapper', optimizer=optimizer, clip_grad=None)
+# learning policy
+param_scheduler = [
+ dict(
+ type='LinearLR', start_factor=3e-2, begin=0, end=12000,
+ by_epoch=False),
+ dict(
+ type='PolyLRRatio',
+ eta_min_ratio=3e-2,
+ power=0.9,
+ begin=12000,
+ end=24000,
+ by_epoch=False),
+ dict(type='ConstantLR', by_epoch=False, factor=1, begin=24000, end=25000)
+]
+# training schedule for 25k
+train_cfg = dict(type='IterBasedTrainLoop', max_iters=25000, val_interval=1000)
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
+default_hooks = dict(
+ timer=dict(type='IterTimerHook'),
+ logger=dict(type='LoggerHook', interval=50, log_metric_by_epoch=False),
+ param_scheduler=dict(type='ParamSchedulerHook'),
+ checkpoint=dict(type='CheckpointHook', by_epoch=False, interval=2000),
+ sampler_seed=dict(type='DistSamplerSeedHook'),
+ visualization=dict(type='SegVisualizationHook'))
diff --git a/configs/ann/README.md b/configs/ann/README.md
index cbc1be2dc15..1281a9ee14f 100644
--- a/configs/ann/README.md
+++ b/configs/ann/README.md
@@ -26,34 +26,34 @@ The non-local module works as a particularly useful technique for semantic segme
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| ANN | R-50-D8 | 512x1024 | 40000 | 6 | 3.71 | V100 | 77.40 | 78.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211-049fc292.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211.log.json) |
-| ANN | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.55 | V100 | 76.55 | 78.85 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243-adf6eece.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243.log.json) |
-| ANN | R-50-D8 | 769x769 | 40000 | 6.8 | 1.70 | V100 | 78.89 | 80.46 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712-2b46b04d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712.log.json) |
-| ANN | R-101-D8 | 769x769 | 40000 | 10.7 | 1.15 | V100 | 79.32 | 80.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720-059bff28.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720.log.json) |
-| ANN | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 77.34 | 78.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911-5a9ad545.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911.log.json) |
-| ANN | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 77.14 | 78.81 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728-aceccc6e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728.log.json) |
-| ANN | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.88 | 80.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426-cc7ff323.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426.log.json) |
-| ANN | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.80 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713-a9d4be8d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| ANN | R-50-D8 | 512x1024 | 40000 | 6 | 3.71 | V100 | 77.40 | 78.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211-049fc292.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211.log.json) |
+| ANN | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.55 | V100 | 76.55 | 78.85 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243-adf6eece.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243.log.json) |
+| ANN | R-50-D8 | 769x769 | 40000 | 6.8 | 1.70 | V100 | 78.89 | 80.46 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712-2b46b04d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712.log.json) |
+| ANN | R-101-D8 | 769x769 | 40000 | 10.7 | 1.15 | V100 | 79.32 | 80.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720-059bff28.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720.log.json) |
+| ANN | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 77.34 | 78.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911-5a9ad545.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911.log.json) |
+| ANN | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 77.14 | 78.81 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728-aceccc6e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728.log.json) |
+| ANN | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.88 | 80.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426-cc7ff323.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426.log.json) |
+| ANN | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.80 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713-a9d4be8d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ----------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| ANN | R-50-D8 | 512x512 | 80000 | 9.1 | 21.01 | V100 | 41.01 | 42.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818-26f75e11.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818.log.json) |
-| ANN | R-101-D8 | 512x512 | 80000 | 12.5 | 14.12 | V100 | 42.94 | 44.18 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818-c0153543.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818.log.json) |
-| ANN | R-50-D8 | 512x512 | 160000 | - | - | V100 | 41.74 | 42.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733-892247bc.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733.log.json) |
-| ANN | R-101-D8 | 512x512 | 160000 | - | - | V100 | 42.94 | 44.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733-955eb1ec.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | -------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| ANN | R-50-D8 | 512x512 | 80000 | 9.1 | 21.01 | V100 | 41.01 | 42.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818-26f75e11.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818.log.json) |
+| ANN | R-101-D8 | 512x512 | 80000 | 12.5 | 14.12 | V100 | 42.94 | 44.18 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818-c0153543.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818.log.json) |
+| ANN | R-50-D8 | 512x512 | 160000 | - | - | V100 | 41.74 | 42.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733-892247bc.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733.log.json) |
+| ANN | R-101-D8 | 512x512 | 160000 | - | - | V100 | 42.94 | 44.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733-955eb1ec.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733.log.json) |
### Pascal VOC 2012 + Aug
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| ANN | R-50-D8 | 512x512 | 20000 | 6 | 20.92 | V100 | 74.86 | 76.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246-dfcb1c62.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
-| ANN | R-101-D8 | 512x512 | 20000 | 9.5 | 13.94 | V100 | 77.47 | 78.70 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246-2fad0042.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
-| ANN | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.56 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314-b5dac322.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
-| ANN | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.70 | 78.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314-bd205bbe.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| ANN | R-50-D8 | 512x512 | 20000 | 6 | 20.92 | V100 | 74.86 | 76.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246-dfcb1c62.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
+| ANN | R-101-D8 | 512x512 | 20000 | 9.5 | 13.94 | V100 | 77.47 | 78.70 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246-2fad0042.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
+| ANN | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.56 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314-b5dac322.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
+| ANN | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.70 | 78.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314-bd205bbe.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
## Citation
diff --git a/configs/apcnet/README.md b/configs/apcnet/README.md
index 0d7f1d66149..9104f3c87f5 100644
--- a/configs/apcnet/README.md
+++ b/configs/apcnet/README.md
@@ -26,25 +26,25 @@ Recent studies witnessed that context features can significantly improve the per
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| APCNet | R-50-D8 | 512x1024 | 40000 | 7.7 | 3.57 | V100 | 78.02 | 79.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes_20201214_115717-5e88fa33.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes-20201214_115717.log.json) |
-| APCNet | R-101-D8 | 512x1024 | 40000 | 11.2 | 2.15 | V100 | 79.08 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes_20201214_115716-abc9d111.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes-20201214_115716.log.json) |
-| APCNet | R-50-D8 | 769x769 | 40000 | 8.7 | 1.52 | V100 | 77.89 | 79.75 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes_20201214_115717-2a2628d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes-20201214_115717.log.json) |
-| APCNet | R-101-D8 | 769x769 | 40000 | 12.7 | 1.03 | V100 | 77.96 | 79.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes_20201214_115718-b650de90.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes-20201214_115718.log.json) |
-| APCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 78.96 | 79.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes_20201214_115716-987f51e3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes-20201214_115716.log.json) |
-| APCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 79.64 | 80.61 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes_20201214_115705-b1ff208a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes-20201214_115705.log.json) |
-| APCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.79 | 80.35 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes_20201214_115718-7ea9fa12.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes-20201214_115718.log.json) |
-| APCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.45 | 79.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes_20201214_115716-a7fbc2ab.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes-20201214_115716.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------------ | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| APCNet | R-50-D8 | 512x1024 | 40000 | 7.7 | 3.57 | V100 | 78.02 | 79.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes_20201214_115717-5e88fa33.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes-20201214_115717.log.json) |
+| APCNet | R-101-D8 | 512x1024 | 40000 | 11.2 | 2.15 | V100 | 79.08 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes_20201214_115716-abc9d111.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes-20201214_115716.log.json) |
+| APCNet | R-50-D8 | 769x769 | 40000 | 8.7 | 1.52 | V100 | 77.89 | 79.75 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes_20201214_115717-2a2628d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes-20201214_115717.log.json) |
+| APCNet | R-101-D8 | 769x769 | 40000 | 12.7 | 1.03 | V100 | 77.96 | 79.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes_20201214_115718-b650de90.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes-20201214_115718.log.json) |
+| APCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 78.96 | 79.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes_20201214_115716-987f51e3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes-20201214_115716.log.json) |
+| APCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 79.64 | 80.61 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes_20201214_115705-b1ff208a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes-20201214_115705.log.json) |
+| APCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.79 | 80.35 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes_20201214_115718-7ea9fa12.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes-20201214_115718.log.json) |
+| APCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.45 | 79.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes_20201214_115716-a7fbc2ab.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes-20201214_115716.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ----------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| APCNet | R-50-D8 | 512x512 | 80000 | 10.1 | 19.61 | V100 | 42.20 | 43.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k_20201214_115705-a8626293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k-20201214_115705.log.json) |
-| APCNet | R-101-D8 | 512x512 | 80000 | 13.6 | 13.10 | V100 | 45.54 | 46.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k_20201214_115704-c656c3fb.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k-20201214_115704.log.json) |
-| APCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 43.40 | 43.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k_20201214_115706-25fb92c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k-20201214_115706.log.json) |
-| APCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 45.41 | 46.63 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k_20201214_115705-73f9a8d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k-20201214_115705.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | -------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| APCNet | R-50-D8 | 512x512 | 80000 | 10.1 | 19.61 | V100 | 42.20 | 43.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k_20201214_115705-a8626293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k-20201214_115705.log.json) |
+| APCNet | R-101-D8 | 512x512 | 80000 | 13.6 | 13.10 | V100 | 45.54 | 46.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k_20201214_115704-c656c3fb.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k-20201214_115704.log.json) |
+| APCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 43.40 | 43.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k_20201214_115706-25fb92c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k-20201214_115706.log.json) |
+| APCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 45.41 | 46.63 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k_20201214_115705-73f9a8d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k-20201214_115705.log.json) |
## Citation
diff --git a/configs/beit/README.md b/configs/beit/README.md
index 8e34e294101..b005c88c501 100644
--- a/configs/beit/README.md
+++ b/configs/beit/README.md
@@ -67,10 +67,10 @@ upernet_beit-large_fp16_8x1_640x640_160k_ade20k-8fc0dd5d.pth $GPUS --eval mIoU
### ADE20K
-| Method | Backbone | Crop Size | pretrain | pretrain img size | Batch Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------- | -------- | --------- | ------------ | ----------------- | ---------- | ------- | -------- | -------------- | ------ | ----- | ------------: | ----------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| UPerNet | BEiT-B | 640x640 | ImageNet-22K | 224x224 | 16 | 160000 | 15.88 | 2.00 | V100 | 53.08 | 53.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/beit/beit-base_upernet_8xb2-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k-eead221d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k.log.json) |
-| UPerNet | BEiT-L | 640x640 | ImageNet-22K | 224x224 | 8 | 320000 | 22.64 | 0.96 | V100 | 56.33 | 56.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/beit/beit-large_upernet_8xb1-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k-8fc0dd5d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k.log.json) |
+| Method | Backbone | Crop Size | pretrain | pretrain img size | Batch Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------- | -------- | --------- | ------------ | ----------------- | ---------- | ------- | -------- | -------------- | ------ | ----- | ------------: | -------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| UPerNet | BEiT-B | 640x640 | ImageNet-22K | 224x224 | 16 | 160000 | 15.88 | 2.00 | V100 | 53.08 | 53.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/beit/beit-base_upernet_8xb2-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k-eead221d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k.log.json) |
+| UPerNet | BEiT-L | 640x640 | ImageNet-22K | 224x224 | 8 | 320000 | 22.64 | 0.96 | V100 | 56.33 | 56.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/beit/beit-large_upernet_8xb1-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k-8fc0dd5d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k.log.json) |
## Citation
diff --git a/configs/bisenetv1/README.md b/configs/bisenetv1/README.md
index c895fdfe63f..a5058957f0c 100644
--- a/configs/bisenetv1/README.md
+++ b/configs/bisenetv1/README.md
@@ -26,24 +26,24 @@ Semantic segmentation requires both rich spatial information and sizeable recept
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| --------- | ---------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | -------------------------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| BiSeNetV1 | R-18-D32 (No Pretrain) | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.44 | 77.05 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239-c55e78e2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239.log.json) |
-| BiSeNetV1 | R-18-D32 | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.37 | 76.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251-8ba80eff.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251.log.json) |
-| BiSeNetV1 | R-18-D32 (4x8) | 1024x1024 | 160000 | 11.17 | 31.77 | V100 | 75.16 | 77.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322-bb8db75f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322.log.json) |
-| BiSeNetV1 | R-50-D32 (No Pretrain) | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 76.92 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639-7b28a2a6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639.log.json) |
-| BiSeNetV1 | R-50-D32 | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 77.68 | 79.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628-8b304447.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| --------- | ---------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ----------------------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| BiSeNetV1 | R-18-D32 (No Pretrain) | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.44 | 77.05 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239-c55e78e2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239.log.json) |
+| BiSeNetV1 | R-18-D32 | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.37 | 76.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251-8ba80eff.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251.log.json) |
+| BiSeNetV1 | R-18-D32 (4x8) | 1024x1024 | 160000 | 11.17 | 31.77 | V100 | 75.16 | 77.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322-bb8db75f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322.log.json) |
+| BiSeNetV1 | R-50-D32 (No Pretrain) | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 76.92 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639-7b28a2a6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639.log.json) |
+| BiSeNetV1 | R-50-D32 | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 77.68 | 79.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628-8b304447.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628.log.json) |
### COCO-Stuff 164k
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| --------- | ----------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ----------------------------------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
-| BiSeNetV1 | R-18-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 25.45 | 26.15 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328-046aa2f2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328.log.json) |
-| BiSeNetV1 | R-18-D32 | 512x512 | 160000 | 6.33 | 74.24 | V100 | 28.55 | 29.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100-f700dbf7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100.log.json) |
-| BiSeNetV1 | R-50-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 29.82 | 30.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616-d2bb0df4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616.log.json) |
-| BiSeNetV1 | R-50-D32 | 512x512 | 160000 | 9.28 | 32.60 | V100 | 34.88 | 35.37 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932-66747911.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932.log.json) |
-| BiSeNetV1 | R-101-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 31.14 | 31.76 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147-c6b32c3b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147.log.json) |
-| BiSeNetV1 | R-101-D32 | 512x512 | 160000 | 10.36 | 25.25 | V100 | 37.38 | 37.99 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r101-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220-28c8f092.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| --------- | ----------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | -------------------------------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| BiSeNetV1 | R-18-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 25.45 | 26.15 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328-046aa2f2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328.log.json) |
+| BiSeNetV1 | R-18-D32 | 512x512 | 160000 | 6.33 | 74.24 | V100 | 28.55 | 29.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100-f700dbf7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100.log.json) |
+| BiSeNetV1 | R-50-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 29.82 | 30.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616-d2bb0df4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616.log.json) |
+| BiSeNetV1 | R-50-D32 | 512x512 | 160000 | 9.28 | 32.60 | V100 | 34.88 | 35.37 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932-66747911.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932.log.json) |
+| BiSeNetV1 | R-101-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 31.14 | 31.76 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147-c6b32c3b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147.log.json) |
+| BiSeNetV1 | R-101-D32 | 512x512 | 160000 | 10.36 | 25.25 | V100 | 37.38 | 37.99 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r101-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220-28c8f092.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220.log.json) |
Note:
diff --git a/configs/bisenetv2/README.md b/configs/bisenetv2/README.md
index 1f44c01f36a..a5871dfeb98 100644
--- a/configs/bisenetv2/README.md
+++ b/configs/bisenetv2/README.md
@@ -26,12 +26,12 @@ The low-level details and high-level semantics are both essential to the semanti
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| --------- | ---------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------------------------ | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
-| BiSeNetV2 | BiSeNetV2 | 1024x1024 | 160000 | 7.64 | 31.77 | V100 | 73.21 | 75.74 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551-bcf10f09.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551.log.json) |
-| BiSeNetV2 | BiSeNetV2 (OHEM) | 1024x1024 | 160000 | 7.64 | - | V100 | 73.57 | 75.80 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb4-ohem-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947-5f8103b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947.log.json) |
-| BiSeNetV2 | BiSeNetV2 (4x8) | 1024x1024 | 160000 | 15.05 | - | V100 | 75.76 | 77.79 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032-e1a2eed6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032.log.json) |
-| BiSeNetV2 | BiSeNetV2 (FP16) | 1024x1024 | 160000 | 5.77 | 36.65 | V100 | 73.07 | 75.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb4-amp-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942-b979777b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| --------- | ---------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| BiSeNetV2 | BiSeNetV2 | 1024x1024 | 160000 | 7.64 | 31.77 | V100 | 73.21 | 75.74 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551-bcf10f09.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551.log.json) |
+| BiSeNetV2 | BiSeNetV2 (OHEM) | 1024x1024 | 160000 | 7.64 | - | V100 | 73.57 | 75.80 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb4-ohem-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947-5f8103b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947.log.json) |
+| BiSeNetV2 | BiSeNetV2 (4x8) | 1024x1024 | 160000 | 15.05 | - | V100 | 75.76 | 77.79 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032-e1a2eed6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032.log.json) |
+| BiSeNetV2 | BiSeNetV2 (FP16) | 1024x1024 | 160000 | 5.77 | 36.65 | V100 | 73.07 | 75.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb4-amp-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942-b979777b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942.log.json) |
Note:
diff --git a/configs/ccnet/README.md b/configs/ccnet/README.md
index c7a0163495d..64dd5f0298a 100644
--- a/configs/ccnet/README.md
+++ b/configs/ccnet/README.md
@@ -26,34 +26,34 @@ Contextual information is vital in visual understanding problems, such as semant
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| CCNet | R-50-D8 | 512x1024 | 40000 | 6 | 3.32 | V100 | 77.76 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517-4123f401.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517.log.json) |
-| CCNet | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.31 | V100 | 76.35 | 78.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540-a3b84ba6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540.log.json) |
-| CCNet | R-50-D8 | 769x769 | 40000 | 6.8 | 1.43 | V100 | 78.46 | 79.93 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125-76d11884.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125.log.json) |
-| CCNet | R-101-D8 | 769x769 | 40000 | 10.7 | 1.01 | V100 | 76.94 | 78.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428-4f57c8d0.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428.log.json) |
-| CCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.03 | 80.16 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421-869a3423.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421.log.json) |
-| CCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 78.87 | 79.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935-ffae8917.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935.log.json) |
-| CCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.29 | 81.08 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421-73eed8ca.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421.log.json) |
-| CCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 79.45 | 80.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502-ad3cd481.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| CCNet | R-50-D8 | 512x1024 | 40000 | 6 | 3.32 | V100 | 77.76 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517-4123f401.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517.log.json) |
+| CCNet | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.31 | V100 | 76.35 | 78.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540-a3b84ba6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540.log.json) |
+| CCNet | R-50-D8 | 769x769 | 40000 | 6.8 | 1.43 | V100 | 78.46 | 79.93 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125-76d11884.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125.log.json) |
+| CCNet | R-101-D8 | 769x769 | 40000 | 10.7 | 1.01 | V100 | 76.94 | 78.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428-4f57c8d0.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428.log.json) |
+| CCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.03 | 80.16 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421-869a3423.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421.log.json) |
+| CCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 78.87 | 79.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935-ffae8917.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935.log.json) |
+| CCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.29 | 81.08 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421-73eed8ca.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421.log.json) |
+| CCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 79.45 | 80.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502-ad3cd481.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| CCNet | R-50-D8 | 512x512 | 80000 | 8.8 | 20.89 | V100 | 41.78 | 42.98 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848-aa37f61e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848.log.json) |
-| CCNet | R-101-D8 | 512x512 | 80000 | 12.2 | 14.11 | V100 | 43.97 | 45.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848-1f4929a3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848.log.json) |
-| CCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.08 | 43.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435-7c97193b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435.log.json) |
-| CCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 43.71 | 45.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644-e849e007.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| CCNet | R-50-D8 | 512x512 | 80000 | 8.8 | 20.89 | V100 | 41.78 | 42.98 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848-aa37f61e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848.log.json) |
+| CCNet | R-101-D8 | 512x512 | 80000 | 12.2 | 14.11 | V100 | 43.97 | 45.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848-1f4929a3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848.log.json) |
+| CCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.08 | 43.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435-7c97193b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435.log.json) |
+| CCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 43.71 | 45.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644-e849e007.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644.log.json) |
### Pascal VOC 2012 + Aug
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| CCNet | R-50-D8 | 512x512 | 20000 | 6 | 20.45 | V100 | 76.17 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212-fad81784.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
-| CCNet | R-101-D8 | 512x512 | 20000 | 9.5 | 13.64 | V100 | 77.27 | 79.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212-0007b61d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
-| CCNet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 75.96 | 77.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127-c2a15f02.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
-| CCNet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 77.87 | 78.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127-c30da577.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| CCNet | R-50-D8 | 512x512 | 20000 | 6 | 20.45 | V100 | 76.17 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212-fad81784.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
+| CCNet | R-101-D8 | 512x512 | 20000 | 9.5 | 13.64 | V100 | 77.27 | 79.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212-0007b61d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
+| CCNet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 75.96 | 77.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127-c2a15f02.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
+| CCNet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 77.87 | 78.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127-c30da577.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
## Citation
diff --git a/configs/cgnet/README.md b/configs/cgnet/README.md
index 60cde582918..96c9fcf515c 100644
--- a/configs/cgnet/README.md
+++ b/configs/cgnet/README.md
@@ -26,10 +26,10 @@ The demand of applying semantic segmentation model on mobile devices has been in
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
-| CGNet | M3N21 | 680x680 | 60000 | 7.5 | 30.51 | V100 | 65.63 | 68.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/cgnet/cgnet_fcn_4xb4-60k_cityscapes-680x680.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes_20201101_110253-4c0b2f2d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes-20201101_110253.log.json) |
-| CGNet | M3N21 | 512x1024 | 60000 | 8.3 | 31.14 | V100 | 68.27 | 70.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/cgnet/cgnet_fcn_4xb8-60k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes_20201101_110254-124ea03b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes-20201101_110254.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| CGNet | M3N21 | 680x680 | 60000 | 7.5 | 30.51 | V100 | 65.63 | 68.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/cgnet/cgnet_fcn_4xb4-60k_cityscapes-680x680.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes_20201101_110253-4c0b2f2d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes-20201101_110253.log.json) |
+| CGNet | M3N21 | 512x1024 | 60000 | 8.3 | 31.14 | V100 | 68.27 | 70.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/cgnet/cgnet_fcn_4xb8-60k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes_20201101_110254-124ea03b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes-20201101_110254.log.json) |
## Citation
diff --git a/configs/convnext/README.md b/configs/convnext/README.md
index 91339377c01..d78fe6ee1bb 100644
--- a/configs/convnext/README.md
+++ b/configs/convnext/README.md
@@ -27,7 +27,7 @@ The "Roaring 20s" of visual recognition began with the introduction of Vision Tr
- ConvNeXt backbone needs to install [MMClassification](https://github.com/open-mmlab/mmclassification) first, which has abundant backbones for downstream tasks.
```shell
-pip install mmcls>=0.20.1
+pip install mmpretrain>=1.0.0rc7
```
### Pre-trained Models
@@ -49,14 +49,14 @@ The pre-trained models on ImageNet-1k or ImageNet-21k are used to fine-tune on t
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------- | ----------- | --------- | ------- | -------- | -------------- | ------ | ----- | ------------- | -------------------------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| UPerNet | ConvNeXt-T | 512x512 | 160000 | 4.23 | 19.90 | V100 | 46.11 | 46.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553-cad485de.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553.log.json) |
-| UPerNet | ConvNeXt-S | 512x512 | 160000 | 5.16 | 15.18 | V100 | 48.56 | 49.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208-1b1e394f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208.log.json) |
-| UPerNet | ConvNeXt-B | 512x512 | 160000 | 6.33 | 14.41 | V100 | 48.71 | 49.54 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227-02a24fc6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227.log.json) |
-| UPerNet | ConvNeXt-B | 640x640 | 160000 | 8.53 | 10.88 | V100 | 52.13 | 52.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859-9280e39b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859.log.json) |
-| UPerNet | ConvNeXt-L | 640x640 | 160000 | 12.08 | 7.69 | V100 | 53.16 | 53.38 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532-e57aa54d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532.log.json) |
-| UPerNet | ConvNeXt-XL | 640x640 | 160000 | 26.16\* | 6.33 | V100 | 53.58 | 54.11 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344-95fc38c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------- | ----------- | --------- | ------- | -------- | -------------- | ------ | ----- | ------------- | ----------------------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| UPerNet | ConvNeXt-T | 512x512 | 160000 | 4.23 | 19.90 | V100 | 46.11 | 46.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553-cad485de.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553.log.json) |
+| UPerNet | ConvNeXt-S | 512x512 | 160000 | 5.16 | 15.18 | V100 | 48.56 | 49.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208-1b1e394f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208.log.json) |
+| UPerNet | ConvNeXt-B | 512x512 | 160000 | 6.33 | 14.41 | V100 | 48.71 | 49.54 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227-02a24fc6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227.log.json) |
+| UPerNet | ConvNeXt-B | 640x640 | 160000 | 8.53 | 10.88 | V100 | 52.13 | 52.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859-9280e39b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859.log.json) |
+| UPerNet | ConvNeXt-L | 640x640 | 160000 | 12.08 | 7.69 | V100 | 53.16 | 53.38 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532-e57aa54d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532.log.json) |
+| UPerNet | ConvNeXt-XL | 640x640 | 160000 | 26.16\* | 6.33 | V100 | 53.58 | 54.11 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344-95fc38c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344.log.json) |
Note:
diff --git a/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py b/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py
index a743e9322ac..06a86431442 100644
--- a/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py
+++ b/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py
@@ -9,7 +9,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='base',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py b/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py
index 6d94989ee1e..2956e86f042 100644
--- a/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py
+++ b/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py
@@ -9,7 +9,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='large',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py b/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py
index 3cbf09902dc..dbe45f10e0b 100644
--- a/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py
+++ b/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py
@@ -8,7 +8,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='small',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.3,
diff --git a/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py b/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py
index 9d4968df600..d2e545a76d0 100644
--- a/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py
+++ b/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py
@@ -8,7 +8,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='tiny',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py b/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py
index 749391cac12..dfad7345215 100644
--- a/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py
+++ b/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py
@@ -9,7 +9,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='xlarge',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/danet/README.md b/configs/danet/README.md
index b5841e23be1..90194f3073e 100644
--- a/configs/danet/README.md
+++ b/configs/danet/README.md
@@ -26,34 +26,34 @@ In this paper, we address the scene segmentation task by capturing rich contextu
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| DANet | R-50-D8 | 512x1024 | 40000 | 7.4 | 2.66 | V100 | 78.74 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324-c0dbfa5f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324.log.json) |
-| DANet | R-101-D8 | 512x1024 | 40000 | 10.9 | 1.99 | V100 | 80.52 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831-c57a7157.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831.log.json) |
-| DANet | R-50-D8 | 769x769 | 40000 | 8.8 | 1.56 | V100 | 78.88 | 80.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703-76681c60.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703.log.json) |
-| DANet | R-101-D8 | 769x769 | 40000 | 12.8 | 1.07 | V100 | 79.88 | 81.47 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717-dcb7fd4e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717.log.json) |
-| DANet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.34 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029-2bfa2293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029.log.json) |
-| DANet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 80.41 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918-955e6350.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918.log.json) |
-| DANet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.27 | 80.96 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954-495689b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954.log.json) |
-| DANet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 80.47 | 82.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918-f3a929e7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| DANet | R-50-D8 | 512x1024 | 40000 | 7.4 | 2.66 | V100 | 78.74 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324-c0dbfa5f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324.log.json) |
+| DANet | R-101-D8 | 512x1024 | 40000 | 10.9 | 1.99 | V100 | 80.52 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831-c57a7157.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831.log.json) |
+| DANet | R-50-D8 | 769x769 | 40000 | 8.8 | 1.56 | V100 | 78.88 | 80.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703-76681c60.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703.log.json) |
+| DANet | R-101-D8 | 769x769 | 40000 | 12.8 | 1.07 | V100 | 79.88 | 81.47 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717-dcb7fd4e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717.log.json) |
+| DANet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.34 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029-2bfa2293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029.log.json) |
+| DANet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 80.41 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918-955e6350.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918.log.json) |
+| DANet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.27 | 80.96 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954-495689b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954.log.json) |
+| DANet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 80.47 | 82.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918-f3a929e7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| DANet | R-50-D8 | 512x512 | 80000 | 11.5 | 21.20 | V100 | 41.66 | 42.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125-edb18e08.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125.log.json) |
-| DANet | R-101-D8 | 512x512 | 80000 | 15 | 14.18 | V100 | 43.64 | 45.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126-d0357c73.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126.log.json) |
-| DANet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.45 | 43.25 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340-9cb35dcd.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340.log.json) |
-| DANet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 44.17 | 45.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348-23bf12f9.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| DANet | R-50-D8 | 512x512 | 80000 | 11.5 | 21.20 | V100 | 41.66 | 42.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125-edb18e08.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125.log.json) |
+| DANet | R-101-D8 | 512x512 | 80000 | 15 | 14.18 | V100 | 43.64 | 45.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126-d0357c73.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126.log.json) |
+| DANet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.45 | 43.25 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340-9cb35dcd.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340.log.json) |
+| DANet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 44.17 | 45.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348-23bf12f9.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348.log.json) |
### Pascal VOC 2012 + Aug
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| DANet | R-50-D8 | 512x512 | 20000 | 6.5 | 20.94 | V100 | 74.45 | 75.69 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026-9e9e3ab3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
-| DANet | R-101-D8 | 512x512 | 20000 | 9.9 | 13.76 | V100 | 76.02 | 77.23 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026-d48d23b2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
-| DANet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.37 | 77.29 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526-426e3a64.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526.log.json) |
-| DANet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.51 | 77.32 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031-788e232a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| DANet | R-50-D8 | 512x512 | 20000 | 6.5 | 20.94 | V100 | 74.45 | 75.69 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026-9e9e3ab3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
+| DANet | R-101-D8 | 512x512 | 20000 | 9.9 | 13.76 | V100 | 76.02 | 77.23 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026-d48d23b2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
+| DANet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.37 | 77.29 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526-426e3a64.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526.log.json) |
+| DANet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.51 | 77.32 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031-788e232a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031.log.json) |
## Citation
diff --git a/configs/ddrnet/README.md b/configs/ddrnet/README.md
new file mode 100644
index 00000000000..ccbfcdff359
--- /dev/null
+++ b/configs/ddrnet/README.md
@@ -0,0 +1,46 @@
+# DDRNet
+
+> [Deep Dual-resolution Networks for Real-time and Accurate Semantic Segmentation of Road Scenes](http://arxiv.org/abs/2101.06085)
+
+## Introduction
+
+
+
+Official Repo
+
+## Abstract
+
+
+
+Semantic segmentation is a key technology for autonomous vehicles to understand the surrounding scenes. The appealing performances of contemporary models usually come at the expense of heavy computations and lengthy inference time, which is intolerable for self-driving. Using light-weight architectures (encoder-decoder or two-pathway) or reasoning on low-resolution images, recent methods realize very fast scene parsing, even running at more than 100 FPS on a single 1080Ti GPU. However, there is still a significant gap in performance between these real-time methods and the models based on dilation backbones. To tackle this problem, we proposed a family of efficient backbones specially designed for real-time semantic segmentation. The proposed deep dual-resolution networks (DDRNets) are composed of two deep branches between which multiple bilateral fusions are performed. Additionally, we design a new contextual information extractor named Deep Aggregation Pyramid Pooling Module (DAPPM) to enlarge effective receptive fields and fuse multi-scale context based on low-resolution feature maps. Our method achieves a new state-of-the-art trade-off between accuracy and speed on both Cityscapes and CamVid dataset. In particular, on a single 2080Ti GPU, DDRNet-23-slim yields 77.4% mIoU at 102 FPS on Cityscapes test set and 74.7% mIoU at 230 FPS on CamVid test set. With widely used test augmentation, our method is superior to most state-of-the-art models and requires much less computation. Codes and trained models are available online.
+
+
+
+
+
我们会在 OpenMMLab 社区为大家
diff --git a/configs/_base_/datasets/bdd100k.py b/configs/_base_/datasets/bdd100k.py
new file mode 100644
index 00000000000..24cec69bfeb
--- /dev/null
+++ b/configs/_base_/datasets/bdd100k.py
@@ -0,0 +1,70 @@
+# dataset settings
+dataset_type = 'BDD100KDataset'
+data_root = 'data/bdd100k/'
+
+crop_size = (512, 1024)
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(
+ type='RandomResize',
+ scale=(2048, 1024),
+ ratio_range=(0.5, 2.0),
+ keep_ratio=True),
+ dict(type='RandomCrop', crop_size=crop_size, cat_max_ratio=0.75),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='PhotoMetricDistortion'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 1024), keep_ratio=True),
+ # add loading annotation after ``Resize`` because ground truth
+ # does not need to do resize data transform
+ dict(type='LoadAnnotations'),
+ dict(type='PackSegInputs')
+]
+img_ratios = [0.5, 0.75, 1.0, 1.25, 1.5, 1.75]
+tta_pipeline = [
+ dict(type='LoadImageFromFile', backend_args=None),
+ dict(
+ type='TestTimeAug',
+ transforms=[
+ [
+ dict(type='Resize', scale_factor=r, keep_ratio=True)
+ for r in img_ratios
+ ],
+ [
+ dict(type='RandomFlip', prob=0., direction='horizontal'),
+ dict(type='RandomFlip', prob=1., direction='horizontal')
+ ], [dict(type='LoadAnnotations')], [dict(type='PackSegInputs')]
+ ])
+]
+train_dataloader = dict(
+ batch_size=2,
+ num_workers=2,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/10k/train',
+ seg_map_path='labels/sem_seg/masks/train'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/10k/val',
+ seg_map_path='labels/sem_seg/masks/val'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU'])
+test_evaluator = val_evaluator
diff --git a/configs/_base_/datasets/coco-stuff164k.py b/configs/_base_/datasets/coco-stuff164k.py
index baf633f9d6f..a9b9d90117a 100644
--- a/configs/_base_/datasets/coco-stuff164k.py
+++ b/configs/_base_/datasets/coco-stuff164k.py
@@ -48,7 +48,7 @@
type=dataset_type,
data_root=data_root,
data_prefix=dict(
- img_path='images/train2017', seg_map_path='annotations/val2017'),
+ img_path='images/train2017', seg_map_path='annotations/train2017'),
pipeline=train_pipeline))
val_dataloader = dict(
batch_size=1,
diff --git a/configs/_base_/datasets/levir_256x256.py b/configs/_base_/datasets/levir_256x256.py
new file mode 100644
index 00000000000..a2a69aa9e9c
--- /dev/null
+++ b/configs/_base_/datasets/levir_256x256.py
@@ -0,0 +1,59 @@
+# dataset settings
+dataset_type = 'LEVIRCDDataset'
+data_root = r'data/LEVIRCD'
+
+albu_train_transforms = [
+ dict(type='RandomBrightnessContrast', p=0.2),
+ dict(type='HorizontalFlip', p=0.5),
+ dict(type='VerticalFlip', p=0.5)
+]
+
+train_pipeline = [
+ dict(type='LoadMultipleRSImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='Albu', transforms=albu_train_transforms),
+ dict(type='ConcatCDInput'),
+ dict(type='PackSegInputs')
+]
+test_pipeline = [
+ dict(type='LoadMultipleRSImageFromFile'),
+ dict(type='LoadAnnotations'),
+ dict(type='ConcatCDInput'),
+ dict(type='PackSegInputs')
+]
+
+tta_pipeline = [
+ dict(type='LoadMultipleRSImageFromFile'),
+ dict(
+ type='TestTimeAug',
+ transforms=[[dict(type='LoadAnnotations')],
+ [dict(type='ConcatCDInput')],
+ [dict(type='PackSegInputs')]])
+]
+train_dataloader = dict(
+ batch_size=4,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='train/A',
+ img_path2='train/B',
+ seg_map_path='train/label'),
+ pipeline=train_pipeline))
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='test/A', img_path2='test/B', seg_map_path='test/label'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+val_evaluator = dict(type='IoUMetric', iou_metrics=['mIoU'])
+test_evaluator = val_evaluator
diff --git a/configs/_base_/datasets/nyu.py b/configs/_base_/datasets/nyu.py
new file mode 100644
index 00000000000..74d57c5fc50
--- /dev/null
+++ b/configs/_base_/datasets/nyu.py
@@ -0,0 +1,67 @@
+# dataset settings
+dataset_type = 'NYUDataset'
+data_root = 'data/nyu'
+
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3),
+ dict(type='RandomDepthMix', prob=0.25),
+ dict(type='RandomFlip', prob=0.5),
+ dict(type='RandomCrop', crop_size=(480, 480)),
+ dict(
+ type='Albu',
+ transforms=[
+ dict(type='RandomBrightnessContrast'),
+ dict(type='RandomGamma'),
+ dict(type='HueSaturationValue'),
+ ]),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id')),
+]
+
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2000, 480), keep_ratio=True),
+ dict(dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3)),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id'))
+]
+
+train_dataloader = dict(
+ batch_size=8,
+ num_workers=8,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/train', depth_map_path='annotations/train'),
+ pipeline=train_pipeline))
+
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ test_mode=True,
+ data_prefix=dict(
+ img_path='images/test', depth_map_path='annotations/test'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(
+ type='DepthMetric',
+ min_depth_eval=0.001,
+ max_depth_eval=10.0,
+ crop_type='nyu_crop')
+test_evaluator = val_evaluator
diff --git a/configs/_base_/datasets/nyu_512x512.py b/configs/_base_/datasets/nyu_512x512.py
new file mode 100644
index 00000000000..88e3878d334
--- /dev/null
+++ b/configs/_base_/datasets/nyu_512x512.py
@@ -0,0 +1,72 @@
+# dataset settings
+dataset_type = 'NYUDataset'
+data_root = 'data/nyu'
+
+train_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3),
+ dict(type='RandomDepthMix', prob=0.25),
+ dict(type='RandomFlip', prob=0.5),
+ dict(
+ type='RandomResize',
+ scale=(768, 512),
+ ratio_range=(0.8, 1.5),
+ keep_ratio=True),
+ dict(type='RandomCrop', crop_size=(512, 512)),
+ dict(
+ type='Albu',
+ transforms=[
+ dict(type='RandomBrightnessContrast'),
+ dict(type='RandomGamma'),
+ dict(type='HueSaturationValue'),
+ ]),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id')),
+]
+
+test_pipeline = [
+ dict(type='LoadImageFromFile'),
+ dict(type='Resize', scale=(2048, 512), keep_ratio=True),
+ dict(dict(type='LoadDepthAnnotation', depth_rescale_factor=1e-3)),
+ dict(
+ type='PackSegInputs',
+ meta_keys=('img_path', 'depth_map_path', 'ori_shape', 'img_shape',
+ 'pad_shape', 'scale_factor', 'flip', 'flip_direction',
+ 'category_id'))
+]
+
+train_dataloader = dict(
+ batch_size=8,
+ num_workers=8,
+ persistent_workers=True,
+ sampler=dict(type='InfiniteSampler', shuffle=True),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ data_prefix=dict(
+ img_path='images/train', depth_map_path='annotations/train'),
+ pipeline=train_pipeline))
+
+val_dataloader = dict(
+ batch_size=1,
+ num_workers=4,
+ persistent_workers=True,
+ sampler=dict(type='DefaultSampler', shuffle=False),
+ dataset=dict(
+ type=dataset_type,
+ data_root=data_root,
+ test_mode=True,
+ data_prefix=dict(
+ img_path='images/test', depth_map_path='annotations/test'),
+ pipeline=test_pipeline))
+test_dataloader = val_dataloader
+
+val_evaluator = dict(
+ type='DepthMetric',
+ min_depth_eval=0.001,
+ max_depth_eval=10.0,
+ crop_type='nyu_crop')
+test_evaluator = val_evaluator
diff --git a/configs/_base_/models/fpn_poolformer_s12.py b/configs/_base_/models/fpn_poolformer_s12.py
index b6893f69774..086c804837a 100644
--- a/configs/_base_/models/fpn_poolformer_s12.py
+++ b/configs/_base_/models/fpn_poolformer_s12.py
@@ -1,10 +1,11 @@
# model settings
norm_cfg = dict(type='SyncBN', requires_grad=True)
checkpoint_file = 'https://download.openmmlab.com/mmclassification/v0/poolformer/poolformer-s12_3rdparty_32xb128_in1k_20220414-f8d83051.pth' # noqa
-# TODO: delete custom_imports after mmcls supports auto import
-# please install mmcls>=1.0
-# import mmcls.models to trigger register_module in mmcls
-custom_imports = dict(imports=['mmcls.models'], allow_failed_imports=False)
+# TODO: delete custom_imports after mmpretrain supports auto import
+# please install mmpretrain >= 1.0.0rc7
+# import mmpretrain.models to trigger register_module in mmpretrain
+custom_imports = dict(
+ imports=['mmpretrain.models'], allow_failed_imports=False)
data_preprocessor = dict(
type='SegDataPreProcessor',
mean=[123.675, 116.28, 103.53],
@@ -16,7 +17,7 @@
type='EncoderDecoder',
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.PoolFormer',
+ type='mmpretrain.PoolFormer',
arch='s12',
init_cfg=dict(
type='Pretrained', checkpoint=checkpoint_file, prefix='backbone.'),
diff --git a/configs/_base_/models/san_vit-b16.py b/configs/_base_/models/san_vit-b16.py
new file mode 100644
index 00000000000..96ac41b8dad
--- /dev/null
+++ b/configs/_base_/models/san_vit-b16.py
@@ -0,0 +1,137 @@
+# model settings
+norm_cfg = dict(type='SyncBN', requires_grad=True)
+
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[122.7709, 116.7460, 104.0937],
+ std=[68.5005, 66.6322, 70.3232],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=255,
+ size_divisor=640,
+ test_cfg=dict(size_divisor=32))
+
+num_classes = 171
+model = dict(
+ type='MultimodalEncoderDecoder',
+ data_preprocessor=data_preprocessor,
+ pretrained='pretrain/clip_vit_base_patch16_224.pth',
+ asymetric_input=True,
+ encoder_resolution=0.5,
+ image_encoder=dict(
+ type='VisionTransformer',
+ img_size=(224, 224),
+ patch_size=16,
+ patch_pad=0,
+ in_channels=3,
+ embed_dims=768,
+ num_layers=9,
+ num_heads=12,
+ mlp_ratio=4,
+ out_origin=True,
+ out_indices=(2, 5, 8),
+ qkv_bias=True,
+ drop_rate=0.0,
+ attn_drop_rate=0.0,
+ drop_path_rate=0.0,
+ with_cls_token=True,
+ output_cls_token=True,
+ patch_bias=False,
+ pre_norm=True,
+ norm_cfg=dict(type='LN', eps=1e-5),
+ act_cfg=dict(type='QuickGELU'),
+ norm_eval=False,
+ interpolate_mode='bicubic',
+ frozen_exclude=['pos_embed']),
+ text_encoder=dict(
+ type='CLIPTextEncoder',
+ dataset_name=None,
+ templates='vild',
+ embed_dims=512,
+ num_layers=12,
+ num_heads=8,
+ mlp_ratio=4,
+ output_dims=512,
+ cache_feature=True,
+ cat_bg=True,
+ norm_cfg=dict(type='LN', eps=1e-5)
+ ),
+ decode_head=dict(
+ type='SideAdapterCLIPHead',
+ num_classes=num_classes,
+ deep_supervision_idxs=[7],
+ san_cfg=dict(
+ in_channels=3,
+ clip_channels=768,
+ embed_dims=240,
+ patch_size=16,
+ patch_bias=True,
+ num_queries=100,
+ cfg_encoder=dict(
+ num_encode_layer=8,
+ num_heads=6,
+ mlp_ratio=4
+ ),
+ fusion_index=[0, 1, 2, 3],
+ cfg_decoder=dict(
+ num_heads=12,
+ num_layers=1,
+ embed_channels=256,
+ mlp_channels=256,
+ num_mlp=3,
+ rescale=True),
+ norm_cfg=dict(type='LN', eps=1e-6),
+ ),
+ maskgen_cfg=dict(
+ sos_token_format='cls_token',
+ sos_token_num=100,
+ cross_attn=False,
+ num_layers=3,
+ embed_dims=768,
+ num_heads=12,
+ mlp_ratio=4,
+ qkv_bias=True,
+ out_dims=512,
+ final_norm=True,
+ act_cfg=dict(type='QuickGELU'),
+ norm_cfg=dict(type='LN', eps=1e-5),
+ frozen_exclude=[]
+ ),
+ align_corners=False,
+ train_cfg=dict(
+ num_points=12544,
+ oversample_ratio=3.0,
+ importance_sample_ratio=0.75,
+ assigner=dict(
+ type='HungarianAssigner',
+ match_costs=[
+ dict(type='ClassificationCost', weight=2.0),
+ dict(
+ type='CrossEntropyLossCost',
+ weight=5.0,
+ use_sigmoid=True),
+ dict(
+ type='DiceCost',
+ weight=5.0,
+ pred_act=True,
+ eps=1.0)
+ ])),
+ loss_decode=[dict(type='CrossEntropyLoss',
+ loss_name='loss_cls_ce',
+ loss_weight=2.0,
+ class_weight=[1.0] * num_classes + [0.1]),
+ dict(type='CrossEntropyLoss',
+ use_sigmoid=True,
+ loss_name='loss_mask_ce',
+ loss_weight=5.0),
+ dict(type='DiceLoss',
+ ignore_index=None,
+ naive_dice=True,
+ eps=1,
+ loss_name='loss_mask_dice',
+ loss_weight=5.0)
+ ]),
+
+ # model training and testing settings
+ train_cfg=dict(),
+ test_cfg=dict(mode='whole')) # yapf: disable
diff --git a/configs/_base_/models/upernet_convnext.py b/configs/_base_/models/upernet_convnext.py
index 7595295871d..958994c91e6 100644
--- a/configs/_base_/models/upernet_convnext.py
+++ b/configs/_base_/models/upernet_convnext.py
@@ -1,5 +1,5 @@
norm_cfg = dict(type='SyncBN', requires_grad=True)
-custom_imports = dict(imports='mmcls.models', allow_failed_imports=False)
+custom_imports = dict(imports='mmpretrain.models', allow_failed_imports=False)
checkpoint_file = 'https://download.openmmlab.com/mmclassification/v0/convnext/downstream/convnext-base_3rdparty_32xb128-noema_in1k_20220301-2a0ee547.pth' # noqa
data_preprocessor = dict(
type='SegDataPreProcessor',
@@ -13,7 +13,7 @@
data_preprocessor=data_preprocessor,
pretrained=None,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='base',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/_base_/models/vpd_sd.py b/configs/_base_/models/vpd_sd.py
new file mode 100644
index 00000000000..87321e74f04
--- /dev/null
+++ b/configs/_base_/models/vpd_sd.py
@@ -0,0 +1,86 @@
+# model settings
+data_preprocessor = dict(
+ type='SegDataPreProcessor',
+ mean=[127.5, 127.5, 127.5],
+ std=[127.5, 127.5, 127.5],
+ bgr_to_rgb=True,
+ pad_val=0,
+ seg_pad_val=0)
+
+# adapted from stable-diffusion/configs/stable-diffusion/v1-inference.yaml
+stable_diffusion_cfg = dict(
+ base_learning_rate=0.0001,
+ target='ldm.models.diffusion.ddpm.LatentDiffusion',
+ checkpoint='https://download.openmmlab.com/mmsegmentation/v0.5/'
+ 'vpd/stable_diffusion_v1-5_pretrain_third_party.pth',
+ params=dict(
+ linear_start=0.00085,
+ linear_end=0.012,
+ num_timesteps_cond=1,
+ log_every_t=200,
+ timesteps=1000,
+ first_stage_key='jpg',
+ cond_stage_key='txt',
+ image_size=64,
+ channels=4,
+ cond_stage_trainable=False,
+ conditioning_key='crossattn',
+ monitor='val/loss_simple_ema',
+ scale_factor=0.18215,
+ use_ema=False,
+ scheduler_config=dict(
+ target='ldm.lr_scheduler.LambdaLinearScheduler',
+ params=dict(
+ warm_up_steps=[10000],
+ cycle_lengths=[10000000000000],
+ f_start=[1e-06],
+ f_max=[1.0],
+ f_min=[1.0])),
+ unet_config=dict(
+ target='ldm.modules.diffusionmodules.openaimodel.UNetModel',
+ params=dict(
+ image_size=32,
+ in_channels=4,
+ out_channels=4,
+ model_channels=320,
+ attention_resolutions=[4, 2, 1],
+ num_res_blocks=2,
+ channel_mult=[1, 2, 4, 4],
+ num_heads=8,
+ use_spatial_transformer=True,
+ transformer_depth=1,
+ context_dim=768,
+ use_checkpoint=True,
+ legacy=False)),
+ first_stage_config=dict(
+ target='ldm.models.autoencoder.AutoencoderKL',
+ params=dict(
+ embed_dim=4,
+ monitor='val/rec_loss',
+ ddconfig=dict(
+ double_z=True,
+ z_channels=4,
+ resolution=256,
+ in_channels=3,
+ out_ch=3,
+ ch=128,
+ ch_mult=[1, 2, 4, 4],
+ num_res_blocks=2,
+ attn_resolutions=[],
+ dropout=0.0),
+ lossconfig=dict(target='torch.nn.Identity'))),
+ cond_stage_config=dict(
+ target='ldm.modules.encoders.modules.AbstractEncoder')))
+
+model = dict(
+ type='DepthEstimator',
+ data_preprocessor=data_preprocessor,
+ backbone=dict(
+ type='VPD',
+ diffusion_cfg=stable_diffusion_cfg,
+ ),
+)
+
+# some of the parameters in stable-diffusion model will not be updated
+# during training
+find_unused_parameters = True
diff --git a/configs/_base_/schedules/schedule_25k.py b/configs/_base_/schedules/schedule_25k.py
new file mode 100644
index 00000000000..825e141ed12
--- /dev/null
+++ b/configs/_base_/schedules/schedule_25k.py
@@ -0,0 +1,28 @@
+# optimizer
+optimizer = dict(type='AdamW', lr=0.001, weight_decay=0.1)
+optim_wrapper = dict(type='OptimWrapper', optimizer=optimizer, clip_grad=None)
+# learning policy
+param_scheduler = [
+ dict(
+ type='LinearLR', start_factor=3e-2, begin=0, end=12000,
+ by_epoch=False),
+ dict(
+ type='PolyLRRatio',
+ eta_min_ratio=3e-2,
+ power=0.9,
+ begin=12000,
+ end=24000,
+ by_epoch=False),
+ dict(type='ConstantLR', by_epoch=False, factor=1, begin=24000, end=25000)
+]
+# training schedule for 25k
+train_cfg = dict(type='IterBasedTrainLoop', max_iters=25000, val_interval=1000)
+val_cfg = dict(type='ValLoop')
+test_cfg = dict(type='TestLoop')
+default_hooks = dict(
+ timer=dict(type='IterTimerHook'),
+ logger=dict(type='LoggerHook', interval=50, log_metric_by_epoch=False),
+ param_scheduler=dict(type='ParamSchedulerHook'),
+ checkpoint=dict(type='CheckpointHook', by_epoch=False, interval=2000),
+ sampler_seed=dict(type='DistSamplerSeedHook'),
+ visualization=dict(type='SegVisualizationHook'))
diff --git a/configs/ann/README.md b/configs/ann/README.md
index cbc1be2dc15..1281a9ee14f 100644
--- a/configs/ann/README.md
+++ b/configs/ann/README.md
@@ -26,34 +26,34 @@ The non-local module works as a particularly useful technique for semantic segme
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| ANN | R-50-D8 | 512x1024 | 40000 | 6 | 3.71 | V100 | 77.40 | 78.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211-049fc292.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211.log.json) |
-| ANN | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.55 | V100 | 76.55 | 78.85 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243-adf6eece.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243.log.json) |
-| ANN | R-50-D8 | 769x769 | 40000 | 6.8 | 1.70 | V100 | 78.89 | 80.46 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712-2b46b04d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712.log.json) |
-| ANN | R-101-D8 | 769x769 | 40000 | 10.7 | 1.15 | V100 | 79.32 | 80.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720-059bff28.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720.log.json) |
-| ANN | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 77.34 | 78.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911-5a9ad545.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911.log.json) |
-| ANN | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 77.14 | 78.81 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728-aceccc6e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728.log.json) |
-| ANN | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.88 | 80.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426-cc7ff323.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426.log.json) |
-| ANN | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.80 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713-a9d4be8d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| ANN | R-50-D8 | 512x1024 | 40000 | 6 | 3.71 | V100 | 77.40 | 78.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211-049fc292.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_40k_cityscapes/ann_r50-d8_512x1024_40k_cityscapes_20200605_095211.log.json) |
+| ANN | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.55 | V100 | 76.55 | 78.85 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243-adf6eece.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_40k_cityscapes/ann_r101-d8_512x1024_40k_cityscapes_20200605_095243.log.json) |
+| ANN | R-50-D8 | 769x769 | 40000 | 6.8 | 1.70 | V100 | 78.89 | 80.46 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712-2b46b04d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_40k_cityscapes/ann_r50-d8_769x769_40k_cityscapes_20200530_025712.log.json) |
+| ANN | R-101-D8 | 769x769 | 40000 | 10.7 | 1.15 | V100 | 79.32 | 80.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720-059bff28.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_40k_cityscapes/ann_r101-d8_769x769_40k_cityscapes_20200530_025720.log.json) |
+| ANN | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 77.34 | 78.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911-5a9ad545.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x1024_80k_cityscapes/ann_r50-d8_512x1024_80k_cityscapes_20200607_101911.log.json) |
+| ANN | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 77.14 | 78.81 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728-aceccc6e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x1024_80k_cityscapes/ann_r101-d8_512x1024_80k_cityscapes_20200607_013728.log.json) |
+| ANN | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.88 | 80.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426-cc7ff323.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_769x769_80k_cityscapes/ann_r50-d8_769x769_80k_cityscapes_20200607_044426.log.json) |
+| ANN | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.80 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713-a9d4be8d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_769x769_80k_cityscapes/ann_r101-d8_769x769_80k_cityscapes_20200607_013713.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ----------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| ANN | R-50-D8 | 512x512 | 80000 | 9.1 | 21.01 | V100 | 41.01 | 42.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818-26f75e11.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818.log.json) |
-| ANN | R-101-D8 | 512x512 | 80000 | 12.5 | 14.12 | V100 | 42.94 | 44.18 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818-c0153543.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818.log.json) |
-| ANN | R-50-D8 | 512x512 | 160000 | - | - | V100 | 41.74 | 42.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733-892247bc.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733.log.json) |
-| ANN | R-101-D8 | 512x512 | 160000 | - | - | V100 | 42.94 | 44.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733-955eb1ec.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | -------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| ANN | R-50-D8 | 512x512 | 80000 | 9.1 | 21.01 | V100 | 41.01 | 42.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818-26f75e11.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_80k_ade20k/ann_r50-d8_512x512_80k_ade20k_20200615_014818.log.json) |
+| ANN | R-101-D8 | 512x512 | 80000 | 12.5 | 14.12 | V100 | 42.94 | 44.18 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818-c0153543.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_80k_ade20k/ann_r101-d8_512x512_80k_ade20k_20200615_014818.log.json) |
+| ANN | R-50-D8 | 512x512 | 160000 | - | - | V100 | 41.74 | 42.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733-892247bc.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_160k_ade20k/ann_r50-d8_512x512_160k_ade20k_20200615_231733.log.json) |
+| ANN | R-101-D8 | 512x512 | 160000 | - | - | V100 | 42.94 | 44.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733-955eb1ec.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_160k_ade20k/ann_r101-d8_512x512_160k_ade20k_20200615_231733.log.json) |
### Pascal VOC 2012 + Aug
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| ANN | R-50-D8 | 512x512 | 20000 | 6 | 20.92 | V100 | 74.86 | 76.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246-dfcb1c62.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
-| ANN | R-101-D8 | 512x512 | 20000 | 9.5 | 13.94 | V100 | 77.47 | 78.70 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246-2fad0042.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
-| ANN | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.56 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314-b5dac322.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
-| ANN | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.70 | 78.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ann/ann_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314-bd205bbe.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| ANN | R-50-D8 | 512x512 | 20000 | 6 | 20.92 | V100 | 74.86 | 76.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246-dfcb1c62.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_20k_voc12aug/ann_r50-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
+| ANN | R-101-D8 | 512x512 | 20000 | 9.5 | 13.94 | V100 | 77.47 | 78.70 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246-2fad0042.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_20k_voc12aug/ann_r101-d8_512x512_20k_voc12aug_20200617_222246.log.json) |
+| ANN | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.56 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314-b5dac322.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r50-d8_512x512_40k_voc12aug/ann_r50-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
+| ANN | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.70 | 78.06 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ann/ann_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314-bd205bbe.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ann/ann_r101-d8_512x512_40k_voc12aug/ann_r101-d8_512x512_40k_voc12aug_20200613_231314.log.json) |
## Citation
diff --git a/configs/apcnet/README.md b/configs/apcnet/README.md
index 0d7f1d66149..9104f3c87f5 100644
--- a/configs/apcnet/README.md
+++ b/configs/apcnet/README.md
@@ -26,25 +26,25 @@ Recent studies witnessed that context features can significantly improve the per
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| APCNet | R-50-D8 | 512x1024 | 40000 | 7.7 | 3.57 | V100 | 78.02 | 79.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes_20201214_115717-5e88fa33.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes-20201214_115717.log.json) |
-| APCNet | R-101-D8 | 512x1024 | 40000 | 11.2 | 2.15 | V100 | 79.08 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes_20201214_115716-abc9d111.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes-20201214_115716.log.json) |
-| APCNet | R-50-D8 | 769x769 | 40000 | 8.7 | 1.52 | V100 | 77.89 | 79.75 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes_20201214_115717-2a2628d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes-20201214_115717.log.json) |
-| APCNet | R-101-D8 | 769x769 | 40000 | 12.7 | 1.03 | V100 | 77.96 | 79.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes_20201214_115718-b650de90.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes-20201214_115718.log.json) |
-| APCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 78.96 | 79.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes_20201214_115716-987f51e3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes-20201214_115716.log.json) |
-| APCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 79.64 | 80.61 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes_20201214_115705-b1ff208a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes-20201214_115705.log.json) |
-| APCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.79 | 80.35 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes_20201214_115718-7ea9fa12.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes-20201214_115718.log.json) |
-| APCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.45 | 79.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes_20201214_115716-a7fbc2ab.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes-20201214_115716.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------------ | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| APCNet | R-50-D8 | 512x1024 | 40000 | 7.7 | 3.57 | V100 | 78.02 | 79.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes_20201214_115717-5e88fa33.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_40k_cityscapes/apcnet_r50-d8_512x1024_40k_cityscapes-20201214_115717.log.json) |
+| APCNet | R-101-D8 | 512x1024 | 40000 | 11.2 | 2.15 | V100 | 79.08 | 80.34 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes_20201214_115716-abc9d111.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_40k_cityscapes/apcnet_r101-d8_512x1024_40k_cityscapes-20201214_115716.log.json) |
+| APCNet | R-50-D8 | 769x769 | 40000 | 8.7 | 1.52 | V100 | 77.89 | 79.75 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes_20201214_115717-2a2628d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_40k_cityscapes/apcnet_r50-d8_769x769_40k_cityscapes-20201214_115717.log.json) |
+| APCNet | R-101-D8 | 769x769 | 40000 | 12.7 | 1.03 | V100 | 77.96 | 79.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes_20201214_115718-b650de90.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_40k_cityscapes/apcnet_r101-d8_769x769_40k_cityscapes-20201214_115718.log.json) |
+| APCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 78.96 | 79.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes_20201214_115716-987f51e3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x1024_80k_cityscapes/apcnet_r50-d8_512x1024_80k_cityscapes-20201214_115716.log.json) |
+| APCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 79.64 | 80.61 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes_20201214_115705-b1ff208a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x1024_80k_cityscapes/apcnet_r101-d8_512x1024_80k_cityscapes-20201214_115705.log.json) |
+| APCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 78.79 | 80.35 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes_20201214_115718-7ea9fa12.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_769x769_80k_cityscapes/apcnet_r50-d8_769x769_80k_cityscapes-20201214_115718.log.json) |
+| APCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 78.45 | 79.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes_20201214_115716-a7fbc2ab.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_769x769_80k_cityscapes/apcnet_r101-d8_769x769_80k_cityscapes-20201214_115716.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ----------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| APCNet | R-50-D8 | 512x512 | 80000 | 10.1 | 19.61 | V100 | 42.20 | 43.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k_20201214_115705-a8626293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k-20201214_115705.log.json) |
-| APCNet | R-101-D8 | 512x512 | 80000 | 13.6 | 13.10 | V100 | 45.54 | 46.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k_20201214_115704-c656c3fb.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k-20201214_115704.log.json) |
-| APCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 43.40 | 43.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k_20201214_115706-25fb92c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k-20201214_115706.log.json) |
-| APCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 45.41 | 46.63 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/apcnet/apcnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k_20201214_115705-73f9a8d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k-20201214_115705.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | -------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| APCNet | R-50-D8 | 512x512 | 80000 | 10.1 | 19.61 | V100 | 42.20 | 43.30 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k_20201214_115705-a8626293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_80k_ade20k/apcnet_r50-d8_512x512_80k_ade20k-20201214_115705.log.json) |
+| APCNet | R-101-D8 | 512x512 | 80000 | 13.6 | 13.10 | V100 | 45.54 | 46.65 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k_20201214_115704-c656c3fb.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_80k_ade20k/apcnet_r101-d8_512x512_80k_ade20k-20201214_115704.log.json) |
+| APCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 43.40 | 43.94 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k_20201214_115706-25fb92c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r50-d8_512x512_160k_ade20k/apcnet_r50-d8_512x512_160k_ade20k-20201214_115706.log.json) |
+| APCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 45.41 | 46.63 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/apcnet/apcnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k_20201214_115705-73f9a8d7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/apcnet/apcnet_r101-d8_512x512_160k_ade20k/apcnet_r101-d8_512x512_160k_ade20k-20201214_115705.log.json) |
## Citation
diff --git a/configs/beit/README.md b/configs/beit/README.md
index 8e34e294101..b005c88c501 100644
--- a/configs/beit/README.md
+++ b/configs/beit/README.md
@@ -67,10 +67,10 @@ upernet_beit-large_fp16_8x1_640x640_160k_ade20k-8fc0dd5d.pth $GPUS --eval mIoU
### ADE20K
-| Method | Backbone | Crop Size | pretrain | pretrain img size | Batch Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------- | -------- | --------- | ------------ | ----------------- | ---------- | ------- | -------- | -------------- | ------ | ----- | ------------: | ----------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| UPerNet | BEiT-B | 640x640 | ImageNet-22K | 224x224 | 16 | 160000 | 15.88 | 2.00 | V100 | 53.08 | 53.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/beit/beit-base_upernet_8xb2-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k-eead221d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k.log.json) |
-| UPerNet | BEiT-L | 640x640 | ImageNet-22K | 224x224 | 8 | 320000 | 22.64 | 0.96 | V100 | 56.33 | 56.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/beit/beit-large_upernet_8xb1-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k-8fc0dd5d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k.log.json) |
+| Method | Backbone | Crop Size | pretrain | pretrain img size | Batch Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------- | -------- | --------- | ------------ | ----------------- | ---------- | ------- | -------- | -------------- | ------ | ----- | ------------: | -------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| UPerNet | BEiT-B | 640x640 | ImageNet-22K | 224x224 | 16 | 160000 | 15.88 | 2.00 | V100 | 53.08 | 53.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/beit/beit-base_upernet_8xb2-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k-eead221d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-base_8x2_640x640_160k_ade20k/upernet_beit-base_8x2_640x640_160k_ade20k.log.json) |
+| UPerNet | BEiT-L | 640x640 | ImageNet-22K | 224x224 | 8 | 320000 | 22.64 | 0.96 | V100 | 56.33 | 56.84 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/beit/beit-large_upernet_8xb1-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k-8fc0dd5d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/beit/upernet_beit-large_fp16_8x1_640x640_160k_ade20k/upernet_beit-large_fp16_8x1_640x640_160k_ade20k.log.json) |
## Citation
diff --git a/configs/bisenetv1/README.md b/configs/bisenetv1/README.md
index c895fdfe63f..a5058957f0c 100644
--- a/configs/bisenetv1/README.md
+++ b/configs/bisenetv1/README.md
@@ -26,24 +26,24 @@ Semantic segmentation requires both rich spatial information and sizeable recept
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| --------- | ---------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | -------------------------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| BiSeNetV1 | R-18-D32 (No Pretrain) | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.44 | 77.05 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239-c55e78e2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239.log.json) |
-| BiSeNetV1 | R-18-D32 | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.37 | 76.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251-8ba80eff.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251.log.json) |
-| BiSeNetV1 | R-18-D32 (4x8) | 1024x1024 | 160000 | 11.17 | 31.77 | V100 | 75.16 | 77.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322-bb8db75f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322.log.json) |
-| BiSeNetV1 | R-50-D32 (No Pretrain) | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 76.92 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639-7b28a2a6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639.log.json) |
-| BiSeNetV1 | R-50-D32 | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 77.68 | 79.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628-8b304447.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| --------- | ---------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ----------------------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| BiSeNetV1 | R-18-D32 (No Pretrain) | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.44 | 77.05 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239-c55e78e2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_4x4_1024x1024_160k_cityscapes_20210922_172239.log.json) |
+| BiSeNetV1 | R-18-D32 | 1024x1024 | 160000 | 5.69 | 31.77 | V100 | 74.37 | 76.91 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251-8ba80eff.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210905_220251.log.json) |
+| BiSeNetV1 | R-18-D32 (4x8) | 1024x1024 | 160000 | 11.17 | 31.77 | V100 | 75.16 | 77.24 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322-bb8db75f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes/bisenetv1_r18-d32_in1k-pre_4x8_1024x1024_160k_cityscapes_20210905_220322.log.json) |
+| BiSeNetV1 | R-50-D32 (No Pretrain) | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 76.92 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639-7b28a2a6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_4x4_1024x1024_160k_cityscapes_20210923_222639.log.json) |
+| BiSeNetV1 | R-50-D32 | 1024x1024 | 160000 | 15.39 | 7.71 | V100 | 77.68 | 79.57 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628-8b304447.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes/bisenetv1_r50-d32_in1k-pre_4x4_1024x1024_160k_cityscapes_20210917_234628.log.json) |
### COCO-Stuff 164k
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| --------- | ----------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ----------------------------------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
-| BiSeNetV1 | R-18-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 25.45 | 26.15 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328-046aa2f2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328.log.json) |
-| BiSeNetV1 | R-18-D32 | 512x512 | 160000 | 6.33 | 74.24 | V100 | 28.55 | 29.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100-f700dbf7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100.log.json) |
-| BiSeNetV1 | R-50-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 29.82 | 30.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616-d2bb0df4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616.log.json) |
-| BiSeNetV1 | R-50-D32 | 512x512 | 160000 | 9.28 | 32.60 | V100 | 34.88 | 35.37 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932-66747911.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932.log.json) |
-| BiSeNetV1 | R-101-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 31.14 | 31.76 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147-c6b32c3b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147.log.json) |
-| BiSeNetV1 | R-101-D32 | 512x512 | 160000 | 10.36 | 25.25 | V100 | 37.38 | 37.99 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv1/bisenetv1_r101-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220-28c8f092.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| --------- | ----------------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | -------------------------------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| BiSeNetV1 | R-18-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 25.45 | 26.15 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328-046aa2f2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211022_054328.log.json) |
+| BiSeNetV1 | R-18-D32 | 512x512 | 160000 | 6.33 | 74.24 | V100 | 28.55 | 29.26 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r18-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100-f700dbf7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r18-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211023_013100.log.json) |
+| BiSeNetV1 | R-50-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 29.82 | 30.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616-d2bb0df4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_040616.log.json) |
+| BiSeNetV1 | R-50-D32 | 512x512 | 160000 | 9.28 | 32.60 | V100 | 34.88 | 35.37 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932-66747911.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r50-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_181932.log.json) |
+| BiSeNetV1 | R-101-D32 (No Pretrain) | 512x512 | 160000 | - | - | V100 | 31.14 | 31.76 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r50-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147-c6b32c3b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211102_164147.log.json) |
+| BiSeNetV1 | R-101-D32 | 512x512 | 160000 | 10.36 | 25.25 | V100 | 37.38 | 37.99 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv1/bisenetv1_r101-d32-in1k-pre_4xb4-160k_coco-stuff164k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220-28c8f092.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv1/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k/bisenetv1_r101-d32_in1k-pre_lr5e-3_4x4_512x512_160k_coco-stuff164k_20211101_225220.log.json) |
Note:
diff --git a/configs/bisenetv2/README.md b/configs/bisenetv2/README.md
index 1f44c01f36a..a5871dfeb98 100644
--- a/configs/bisenetv2/README.md
+++ b/configs/bisenetv2/README.md
@@ -26,12 +26,12 @@ The low-level details and high-level semantics are both essential to the semanti
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| --------- | ---------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------------------------ | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
-| BiSeNetV2 | BiSeNetV2 | 1024x1024 | 160000 | 7.64 | 31.77 | V100 | 73.21 | 75.74 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551-bcf10f09.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551.log.json) |
-| BiSeNetV2 | BiSeNetV2 (OHEM) | 1024x1024 | 160000 | 7.64 | - | V100 | 73.57 | 75.80 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb4-ohem-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947-5f8103b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947.log.json) |
-| BiSeNetV2 | BiSeNetV2 (4x8) | 1024x1024 | 160000 | 15.05 | - | V100 | 75.76 | 77.79 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032-e1a2eed6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032.log.json) |
-| BiSeNetV2 | BiSeNetV2 (FP16) | 1024x1024 | 160000 | 5.77 | 36.65 | V100 | 73.07 | 75.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/bisenetv2/bisenetv2_fcn_4xb4-amp-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942-b979777b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| --------- | ---------------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| BiSeNetV2 | BiSeNetV2 | 1024x1024 | 160000 | 7.64 | 31.77 | V100 | 73.21 | 75.74 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb4-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551-bcf10f09.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_4x4_1024x1024_160k_cityscapes_20210902_015551.log.json) |
+| BiSeNetV2 | BiSeNetV2 (OHEM) | 1024x1024 | 160000 | 7.64 | - | V100 | 73.57 | 75.80 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb4-ohem-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947-5f8103b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_ohem_4x4_1024x1024_160k_cityscapes_20210902_112947.log.json) |
+| BiSeNetV2 | BiSeNetV2 (4x8) | 1024x1024 | 160000 | 15.05 | - | V100 | 75.76 | 77.79 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb8-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032-e1a2eed6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes/bisenetv2_fcn_4x8_1024x1024_160k_cityscapes_20210903_000032.log.json) |
+| BiSeNetV2 | BiSeNetV2 (FP16) | 1024x1024 | 160000 | 5.77 | 36.65 | V100 | 73.07 | 75.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/bisenetv2/bisenetv2_fcn_4xb4-amp-160k_cityscapes-1024x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942-b979777b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/bisenetv2/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes/bisenetv2_fcn_fp16_4x4_1024x1024_160k_cityscapes_20210902_045942.log.json) |
Note:
diff --git a/configs/ccnet/README.md b/configs/ccnet/README.md
index c7a0163495d..64dd5f0298a 100644
--- a/configs/ccnet/README.md
+++ b/configs/ccnet/README.md
@@ -26,34 +26,34 @@ Contextual information is vital in visual understanding problems, such as semant
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| CCNet | R-50-D8 | 512x1024 | 40000 | 6 | 3.32 | V100 | 77.76 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517-4123f401.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517.log.json) |
-| CCNet | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.31 | V100 | 76.35 | 78.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540-a3b84ba6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540.log.json) |
-| CCNet | R-50-D8 | 769x769 | 40000 | 6.8 | 1.43 | V100 | 78.46 | 79.93 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125-76d11884.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125.log.json) |
-| CCNet | R-101-D8 | 769x769 | 40000 | 10.7 | 1.01 | V100 | 76.94 | 78.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428-4f57c8d0.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428.log.json) |
-| CCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.03 | 80.16 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421-869a3423.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421.log.json) |
-| CCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 78.87 | 79.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935-ffae8917.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935.log.json) |
-| CCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.29 | 81.08 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421-73eed8ca.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421.log.json) |
-| CCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 79.45 | 80.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502-ad3cd481.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| CCNet | R-50-D8 | 512x1024 | 40000 | 6 | 3.32 | V100 | 77.76 | 78.87 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517-4123f401.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_40k_cityscapes/ccnet_r50-d8_512x1024_40k_cityscapes_20200616_142517.log.json) |
+| CCNet | R-101-D8 | 512x1024 | 40000 | 9.5 | 2.31 | V100 | 76.35 | 78.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540-a3b84ba6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_40k_cityscapes/ccnet_r101-d8_512x1024_40k_cityscapes_20200616_142540.log.json) |
+| CCNet | R-50-D8 | 769x769 | 40000 | 6.8 | 1.43 | V100 | 78.46 | 79.93 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125-76d11884.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_40k_cityscapes/ccnet_r50-d8_769x769_40k_cityscapes_20200616_145125.log.json) |
+| CCNet | R-101-D8 | 769x769 | 40000 | 10.7 | 1.01 | V100 | 76.94 | 78.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428-4f57c8d0.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_40k_cityscapes/ccnet_r101-d8_769x769_40k_cityscapes_20200617_101428.log.json) |
+| CCNet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.03 | 80.16 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421-869a3423.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x1024_80k_cityscapes/ccnet_r50-d8_512x1024_80k_cityscapes_20200617_010421.log.json) |
+| CCNet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 78.87 | 79.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935-ffae8917.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x1024_80k_cityscapes/ccnet_r101-d8_512x1024_80k_cityscapes_20200617_203935.log.json) |
+| CCNet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.29 | 81.08 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421-73eed8ca.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_769x769_80k_cityscapes/ccnet_r50-d8_769x769_80k_cityscapes_20200617_010421.log.json) |
+| CCNet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 79.45 | 80.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502-ad3cd481.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_769x769_80k_cityscapes/ccnet_r101-d8_769x769_80k_cityscapes_20200618_011502.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| CCNet | R-50-D8 | 512x512 | 80000 | 8.8 | 20.89 | V100 | 41.78 | 42.98 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848-aa37f61e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848.log.json) |
-| CCNet | R-101-D8 | 512x512 | 80000 | 12.2 | 14.11 | V100 | 43.97 | 45.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848-1f4929a3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848.log.json) |
-| CCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.08 | 43.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435-7c97193b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435.log.json) |
-| CCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 43.71 | 45.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644-e849e007.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| CCNet | R-50-D8 | 512x512 | 80000 | 8.8 | 20.89 | V100 | 41.78 | 42.98 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848-aa37f61e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_80k_ade20k/ccnet_r50-d8_512x512_80k_ade20k_20200615_014848.log.json) |
+| CCNet | R-101-D8 | 512x512 | 80000 | 12.2 | 14.11 | V100 | 43.97 | 45.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848-1f4929a3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_80k_ade20k/ccnet_r101-d8_512x512_80k_ade20k_20200615_014848.log.json) |
+| CCNet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.08 | 43.13 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435-7c97193b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_160k_ade20k/ccnet_r50-d8_512x512_160k_ade20k_20200616_084435.log.json) |
+| CCNet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 43.71 | 45.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644-e849e007.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_160k_ade20k/ccnet_r101-d8_512x512_160k_ade20k_20200616_000644.log.json) |
### Pascal VOC 2012 + Aug
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| CCNet | R-50-D8 | 512x512 | 20000 | 6 | 20.45 | V100 | 76.17 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212-fad81784.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
-| CCNet | R-101-D8 | 512x512 | 20000 | 9.5 | 13.64 | V100 | 77.27 | 79.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212-0007b61d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
-| CCNet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 75.96 | 77.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127-c2a15f02.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
-| CCNet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 77.87 | 78.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/ccnet/ccnet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127-c30da577.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| CCNet | R-50-D8 | 512x512 | 20000 | 6 | 20.45 | V100 | 76.17 | 77.51 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212-fad81784.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_20k_voc12aug/ccnet_r50-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
+| CCNet | R-101-D8 | 512x512 | 20000 | 9.5 | 13.64 | V100 | 77.27 | 79.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212-0007b61d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_20k_voc12aug/ccnet_r101-d8_512x512_20k_voc12aug_20200617_193212.log.json) |
+| CCNet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 75.96 | 77.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127-c2a15f02.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r50-d8_512x512_40k_voc12aug/ccnet_r50-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
+| CCNet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 77.87 | 78.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/ccnet/ccnet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127-c30da577.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/ccnet/ccnet_r101-d8_512x512_40k_voc12aug/ccnet_r101-d8_512x512_40k_voc12aug_20200613_232127.log.json) |
## Citation
diff --git a/configs/cgnet/README.md b/configs/cgnet/README.md
index 60cde582918..96c9fcf515c 100644
--- a/configs/cgnet/README.md
+++ b/configs/cgnet/README.md
@@ -26,10 +26,10 @@ The demand of applying semantic segmentation model on mobile devices has been in
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
-| CGNet | M3N21 | 680x680 | 60000 | 7.5 | 30.51 | V100 | 65.63 | 68.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/cgnet/cgnet_fcn_4xb4-60k_cityscapes-680x680.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes_20201101_110253-4c0b2f2d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes-20201101_110253.log.json) |
-| CGNet | M3N21 | 512x1024 | 60000 | 8.3 | 31.14 | V100 | 68.27 | 70.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/cgnet/cgnet_fcn_4xb8-60k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes_20201101_110254-124ea03b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes-20201101_110254.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | ------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------ |
+| CGNet | M3N21 | 680x680 | 60000 | 7.5 | 30.51 | V100 | 65.63 | 68.04 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/cgnet/cgnet_fcn_4xb4-60k_cityscapes-680x680.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes_20201101_110253-4c0b2f2d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_680x680_60k_cityscapes/cgnet_680x680_60k_cityscapes-20201101_110253.log.json) |
+| CGNet | M3N21 | 512x1024 | 60000 | 8.3 | 31.14 | V100 | 68.27 | 70.33 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/cgnet/cgnet_fcn_4xb8-60k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes_20201101_110254-124ea03b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/cgnet/cgnet_512x1024_60k_cityscapes/cgnet_512x1024_60k_cityscapes-20201101_110254.log.json) |
## Citation
diff --git a/configs/convnext/README.md b/configs/convnext/README.md
index 91339377c01..d78fe6ee1bb 100644
--- a/configs/convnext/README.md
+++ b/configs/convnext/README.md
@@ -27,7 +27,7 @@ The "Roaring 20s" of visual recognition began with the introduction of Vision Tr
- ConvNeXt backbone needs to install [MMClassification](https://github.com/open-mmlab/mmclassification) first, which has abundant backbones for downstream tasks.
```shell
-pip install mmcls>=0.20.1
+pip install mmpretrain>=1.0.0rc7
```
### Pre-trained Models
@@ -49,14 +49,14 @@ The pre-trained models on ImageNet-1k or ImageNet-21k are used to fine-tune on t
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------- | ----------- | --------- | ------- | -------- | -------------- | ------ | ----- | ------------- | -------------------------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| UPerNet | ConvNeXt-T | 512x512 | 160000 | 4.23 | 19.90 | V100 | 46.11 | 46.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553-cad485de.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553.log.json) |
-| UPerNet | ConvNeXt-S | 512x512 | 160000 | 5.16 | 15.18 | V100 | 48.56 | 49.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208-1b1e394f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208.log.json) |
-| UPerNet | ConvNeXt-B | 512x512 | 160000 | 6.33 | 14.41 | V100 | 48.71 | 49.54 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227-02a24fc6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227.log.json) |
-| UPerNet | ConvNeXt-B | 640x640 | 160000 | 8.53 | 10.88 | V100 | 52.13 | 52.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859-9280e39b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859.log.json) |
-| UPerNet | ConvNeXt-L | 640x640 | 160000 | 12.08 | 7.69 | V100 | 53.16 | 53.38 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532-e57aa54d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532.log.json) |
-| UPerNet | ConvNeXt-XL | 640x640 | 160000 | 26.16\* | 6.33 | V100 | 53.58 | 54.11 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344-95fc38c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------- | ----------- | --------- | ------- | -------- | -------------- | ------ | ----- | ------------- | ----------------------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| UPerNet | ConvNeXt-T | 512x512 | 160000 | 4.23 | 19.90 | V100 | 46.11 | 46.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553-cad485de.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_tiny_fp16_512x512_160k_ade20k/upernet_convnext_tiny_fp16_512x512_160k_ade20k_20220227_124553.log.json) |
+| UPerNet | ConvNeXt-S | 512x512 | 160000 | 5.16 | 15.18 | V100 | 48.56 | 49.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208-1b1e394f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_small_fp16_512x512_160k_ade20k/upernet_convnext_small_fp16_512x512_160k_ade20k_20220227_131208.log.json) |
+| UPerNet | ConvNeXt-B | 512x512 | 160000 | 6.33 | 14.41 | V100 | 48.71 | 49.54 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227-02a24fc6.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_512x512_160k_ade20k/upernet_convnext_base_fp16_512x512_160k_ade20k_20220227_181227.log.json) |
+| UPerNet | ConvNeXt-B | 640x640 | 160000 | 8.53 | 10.88 | V100 | 52.13 | 52.66 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859-9280e39b.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_base_fp16_640x640_160k_ade20k/upernet_convnext_base_fp16_640x640_160k_ade20k_20220227_182859.log.json) |
+| UPerNet | ConvNeXt-L | 640x640 | 160000 | 12.08 | 7.69 | V100 | 53.16 | 53.38 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532-e57aa54d.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_large_fp16_640x640_160k_ade20k/upernet_convnext_large_fp16_640x640_160k_ade20k_20220226_040532.log.json) |
+| UPerNet | ConvNeXt-XL | 640x640 | 160000 | 26.16\* | 6.33 | V100 | 53.58 | 54.11 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344-95fc38c2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/convnext/upernet_convnext_xlarge_fp16_640x640_160k_ade20k/upernet_convnext_xlarge_fp16_640x640_160k_ade20k_20220226_080344.log.json) |
Note:
diff --git a/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py b/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py
index a743e9322ac..06a86431442 100644
--- a/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py
+++ b/configs/convnext/convnext-base_upernet_8xb2-amp-160k_ade20k-640x640.py
@@ -9,7 +9,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='base',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py b/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py
index 6d94989ee1e..2956e86f042 100644
--- a/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py
+++ b/configs/convnext/convnext-large_upernet_8xb2-amp-160k_ade20k-640x640.py
@@ -9,7 +9,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='large',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py b/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py
index 3cbf09902dc..dbe45f10e0b 100644
--- a/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py
+++ b/configs/convnext/convnext-small_upernet_8xb2-amp-160k_ade20k-512x512.py
@@ -8,7 +8,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='small',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.3,
diff --git a/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py b/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py
index 9d4968df600..d2e545a76d0 100644
--- a/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py
+++ b/configs/convnext/convnext-tiny_upernet_8xb2-amp-160k_ade20k-512x512.py
@@ -8,7 +8,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='tiny',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py b/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py
index 749391cac12..dfad7345215 100644
--- a/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py
+++ b/configs/convnext/convnext-xlarge_upernet_8xb2-amp-160k_ade20k-640x640.py
@@ -9,7 +9,7 @@
model = dict(
data_preprocessor=data_preprocessor,
backbone=dict(
- type='mmcls.ConvNeXt',
+ type='mmpretrain.ConvNeXt',
arch='xlarge',
out_indices=[0, 1, 2, 3],
drop_path_rate=0.4,
diff --git a/configs/danet/README.md b/configs/danet/README.md
index b5841e23be1..90194f3073e 100644
--- a/configs/danet/README.md
+++ b/configs/danet/README.md
@@ -26,34 +26,34 @@ In this paper, we address the scene segmentation task by capturing rich contextu
### Cityscapes
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ------------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| DANet | R-50-D8 | 512x1024 | 40000 | 7.4 | 2.66 | V100 | 78.74 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324-c0dbfa5f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324.log.json) |
-| DANet | R-101-D8 | 512x1024 | 40000 | 10.9 | 1.99 | V100 | 80.52 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831-c57a7157.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831.log.json) |
-| DANet | R-50-D8 | 769x769 | 40000 | 8.8 | 1.56 | V100 | 78.88 | 80.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703-76681c60.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703.log.json) |
-| DANet | R-101-D8 | 769x769 | 40000 | 12.8 | 1.07 | V100 | 79.88 | 81.47 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717-dcb7fd4e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717.log.json) |
-| DANet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.34 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029-2bfa2293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029.log.json) |
-| DANet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 80.41 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918-955e6350.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918.log.json) |
-| DANet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.27 | 80.96 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954-495689b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954.log.json) |
-| DANet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 80.47 | 82.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918-f3a929e7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------- | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| DANet | R-50-D8 | 512x1024 | 40000 | 7.4 | 2.66 | V100 | 78.74 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324-c0dbfa5f.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_40k_cityscapes/danet_r50-d8_512x1024_40k_cityscapes_20200605_191324.log.json) |
+| DANet | R-101-D8 | 512x1024 | 40000 | 10.9 | 1.99 | V100 | 80.52 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831-c57a7157.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_40k_cityscapes/danet_r101-d8_512x1024_40k_cityscapes_20200605_200831.log.json) |
+| DANet | R-50-D8 | 769x769 | 40000 | 8.8 | 1.56 | V100 | 78.88 | 80.62 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703-76681c60.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_40k_cityscapes/danet_r50-d8_769x769_40k_cityscapes_20200530_025703.log.json) |
+| DANet | R-101-D8 | 769x769 | 40000 | 12.8 | 1.07 | V100 | 79.88 | 81.47 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-40k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717-dcb7fd4e.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_40k_cityscapes/danet_r101-d8_769x769_40k_cityscapes_20200530_025717.log.json) |
+| DANet | R-50-D8 | 512x1024 | 80000 | - | - | V100 | 79.34 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029-2bfa2293.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x1024_80k_cityscapes/danet_r50-d8_512x1024_80k_cityscapes_20200607_133029.log.json) |
+| DANet | R-101-D8 | 512x1024 | 80000 | - | - | V100 | 80.41 | - | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-512x1024.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918-955e6350.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x1024_80k_cityscapes/danet_r101-d8_512x1024_80k_cityscapes_20200607_132918.log.json) |
+| DANet | R-50-D8 | 769x769 | 80000 | - | - | V100 | 79.27 | 80.96 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954-495689b4.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_769x769_80k_cityscapes/danet_r50-d8_769x769_80k_cityscapes_20200607_132954.log.json) |
+| DANet | R-101-D8 | 769x769 | 80000 | - | - | V100 | 80.47 | 82.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb2-80k_cityscapes-769x769.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918-f3a929e7.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_769x769_80k_cityscapes/danet_r101-d8_769x769_80k_cityscapes_20200607_132918.log.json) |
### ADE20K
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | --------------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| DANet | R-50-D8 | 512x512 | 80000 | 11.5 | 21.20 | V100 | 41.66 | 42.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125-edb18e08.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125.log.json) |
-| DANet | R-101-D8 | 512x512 | 80000 | 15 | 14.18 | V100 | 43.64 | 45.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126-d0357c73.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126.log.json) |
-| DANet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.45 | 43.25 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340-9cb35dcd.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340.log.json) |
-| DANet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 44.17 | 45.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348-23bf12f9.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------ | ---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| DANet | R-50-D8 | 512x512 | 80000 | 11.5 | 21.20 | V100 | 41.66 | 42.90 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125-edb18e08.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_80k_ade20k/danet_r50-d8_512x512_80k_ade20k_20200615_015125.log.json) |
+| DANet | R-101-D8 | 512x512 | 80000 | 15 | 14.18 | V100 | 43.64 | 45.19 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-80k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126-d0357c73.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_80k_ade20k/danet_r101-d8_512x512_80k_ade20k_20200615_015126.log.json) |
+| DANet | R-50-D8 | 512x512 | 160000 | - | - | V100 | 42.45 | 43.25 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340-9cb35dcd.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_160k_ade20k/danet_r50-d8_512x512_160k_ade20k_20200616_082340.log.json) |
+| DANet | R-101-D8 | 512x512 | 160000 | - | - | V100 | 44.17 | 45.02 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-160k_ade20k-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348-23bf12f9.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_160k_ade20k/danet_r101-d8_512x512_160k_ade20k_20200616_082348.log.json) |
### Pascal VOC 2012 + Aug
-| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
-| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ---------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
-| DANet | R-50-D8 | 512x512 | 20000 | 6.5 | 20.94 | V100 | 74.45 | 75.69 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026-9e9e3ab3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
-| DANet | R-101-D8 | 512x512 | 20000 | 9.9 | 13.76 | V100 | 76.02 | 77.23 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026-d48d23b2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
-| DANet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.37 | 77.29 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526-426e3a64.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526.log.json) |
-| DANet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.51 | 77.32 | [config](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/danet/danet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031-788e232a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031.log.json) |
+| Method | Backbone | Crop Size | Lr schd | Mem (GB) | Inf time (fps) | Device | mIoU | mIoU(ms+flip) | config | download |
+| ------ | -------- | --------- | ------: | -------- | -------------- | ------ | ----: | ------------: | ------------------------------------------------------------------------------------------------------------------------- | -------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------- |
+| DANet | R-50-D8 | 512x512 | 20000 | 6.5 | 20.94 | V100 | 74.45 | 75.69 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026-9e9e3ab3.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_20k_voc12aug/danet_r50-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
+| DANet | R-101-D8 | 512x512 | 20000 | 9.9 | 13.76 | V100 | 76.02 | 77.23 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-20k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026-d48d23b2.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_20k_voc12aug/danet_r101-d8_512x512_20k_voc12aug_20200618_070026.log.json) |
+| DANet | R-50-D8 | 512x512 | 40000 | - | - | V100 | 76.37 | 77.29 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r50-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526-426e3a64.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r50-d8_512x512_40k_voc12aug/danet_r50-d8_512x512_40k_voc12aug_20200613_235526.log.json) |
+| DANet | R-101-D8 | 512x512 | 40000 | - | - | V100 | 76.51 | 77.32 | [config](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/danet/danet_r101-d8_4xb4-40k_voc12aug-512x512.py) | [model](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031-788e232a.pth) \| [log](https://download.openmmlab.com/mmsegmentation/v0.5/danet/danet_r101-d8_512x512_40k_voc12aug/danet_r101-d8_512x512_40k_voc12aug_20200613_223031.log.json) |
## Citation
diff --git a/configs/ddrnet/README.md b/configs/ddrnet/README.md
new file mode 100644
index 00000000000..ccbfcdff359
--- /dev/null
+++ b/configs/ddrnet/README.md
@@ -0,0 +1,46 @@
+# DDRNet
+
+> [Deep Dual-resolution Networks for Real-time and Accurate Semantic Segmentation of Road Scenes](http://arxiv.org/abs/2101.06085)
+
+## Introduction
+
+
+
+Official Repo
+
+## Abstract
+
+
+
+Semantic segmentation is a key technology for autonomous vehicles to understand the surrounding scenes. The appealing performances of contemporary models usually come at the expense of heavy computations and lengthy inference time, which is intolerable for self-driving. Using light-weight architectures (encoder-decoder or two-pathway) or reasoning on low-resolution images, recent methods realize very fast scene parsing, even running at more than 100 FPS on a single 1080Ti GPU. However, there is still a significant gap in performance between these real-time methods and the models based on dilation backbones. To tackle this problem, we proposed a family of efficient backbones specially designed for real-time semantic segmentation. The proposed deep dual-resolution networks (DDRNets) are composed of two deep branches between which multiple bilateral fusions are performed. Additionally, we design a new contextual information extractor named Deep Aggregation Pyramid Pooling Module (DAPPM) to enlarge effective receptive fields and fuse multi-scale context based on low-resolution feature maps. Our method achieves a new state-of-the-art trade-off between accuracy and speed on both Cityscapes and CamVid dataset. In particular, on a single 2080Ti GPU, DDRNet-23-slim yields 77.4% mIoU at 102 FPS on Cityscapes test set and 74.7% mIoU at 230 FPS on CamVid test set. With widely used test augmentation, our method is superior to most state-of-the-art models and requires much less computation. Codes and trained models are available online.
+
+
+
+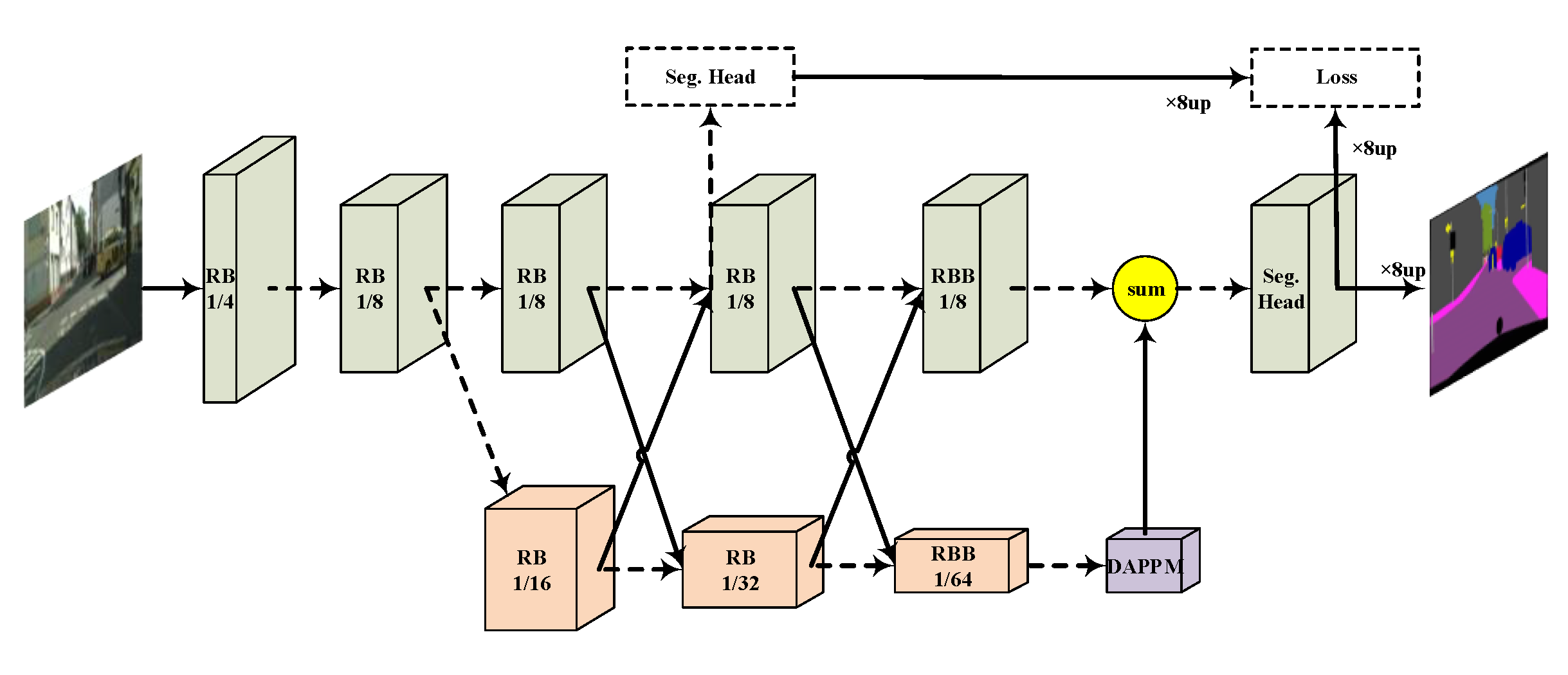 +
+ +
+ -
-  -
- 
 -
-  -
- 
 diff --git a/docs/zh_cn/advanced_guides/add_datasets.md b/docs/zh_cn/advanced_guides/add_datasets.md
index 4ea14934ed2..22fbf3462fc 100644
--- a/docs/zh_cn/advanced_guides/add_datasets.md
+++ b/docs/zh_cn/advanced_guides/add_datasets.md
@@ -1,4 +1,62 @@
-# 新增自定义数据集(待更新)
+# 新增自定义数据集
+
+## 新增自定义数据集
+
+在这里,我们展示如何构建一个新的数据集。
+
+1. 创建一个新文件 `mmseg/datasets/example.py`
+
+ ```python
+ from mmseg.registry import DATASETS
+ from .basesegdataset import BaseSegDataset
+
+
+ @DATASETS.register_module()
+ class ExampleDataset(BaseSegDataset):
+
+ METAINFO = dict(
+ classes=('xxx', 'xxx', ...),
+ palette=[[x, x, x], [x, x, x], ...])
+
+ def __init__(self, aeg1, arg2):
+ pass
+ ```
+
+2. 在 `mmseg/datasets/__init__.py` 中导入模块
+
+ ```python
+ from .example import ExampleDataset
+ ```
+
+3. 通过创建一个新的数据集配置文件 `configs/_base_/datasets/example_dataset.py` 来使用它
+
+ ```python
+ dataset_type = 'ExampleDataset'
+ data_root = 'data/example/'
+ ...
+ ```
+
+4. 在 `mmseg/utils/class_names.py` 中补充数据集元信息
+
+ ```python
+ def example_classes():
+ return [
+ 'xxx', 'xxx',
+ ...
+ ]
+
+ def example_palette():
+ return [
+ [x, x, x], [x, x, x],
+ ...
+ ]
+ dataset_aliases ={
+ 'example': ['example', ...],
+ ...
+ }
+ ```
+
+**注意:** 如果新数据集不满足 mmseg 的要求,则需要在 `tools/dataset_converters/` 中准备一个数据集预处理脚本
## 通过重新组织数据来定制数据集
@@ -26,30 +84,17 @@
一个训练对将由 img_dir/ann_dir 里同样首缀的文件组成。
-如果给定 `split` 参数,只有部分在 img_dir/ann_dir 里的文件会被加载。
-我们可以对被包括在 split 文本里的文件指定前缀。
+有些数据集不会发布测试集或测试集的标注,如果没有测试集的标注,我们就无法在本地进行评估模型,因此我们在配置文件中将验证集设置为默认测试集。
-除此以外,一个 split 文本如下所示:
-
-```none
-xxx
-zzz
-```
+关于如何构建自己的数据集或实现新的数据集类,请参阅[数据集指南](./datasets.md)以获取更多详细信息。
-只有
-
-`data/my_dataset/img_dir/train/xxx{img_suffix}`,
-`data/my_dataset/img_dir/train/zzz{img_suffix}`,
-`data/my_dataset/ann_dir/train/xxx{seg_map_suffix}`,
-`data/my_dataset/ann_dir/train/zzz{seg_map_suffix}` 将被加载。
-
-注意:标注是跟图像同样的形状 (H, W),其中的像素值的范围是 `[0, num_classes - 1]`。
+**注意:** 标注是跟图像同样的形状 (H, W),其中的像素值的范围是 `[0, num_classes - 1]`。
您也可以使用 [pillow](https://pillow.readthedocs.io/en/stable/handbook/concepts.html#palette) 的 `'P'` 模式去创建包含颜色的标注。
## 通过混合数据去定制数据集
MMSegmentation 同样支持混合数据集去训练。
-当前它支持拼接 (concat), 重复 (repeat) 和多图混合 (multi-image mix)数据集。
+当前它支持拼接 (concat), 重复 (repeat) 和多图混合 (multi-image mix) 数据集。
### 重复数据集
@@ -58,79 +103,29 @@ MMSegmentation 同样支持混合数据集去训练。
```python
dataset_A_train = dict(
- type='RepeatDataset',
- times=N,
- dataset=dict( # 这是 Dataset_A 数据集的原始配置
- type='Dataset_A',
- ...
- pipeline=train_pipeline
- )
+ type='RepeatDataset',
+ times=N,
+ dataset=dict( # 这是 Dataset_A 数据集的原始配置
+ type='Dataset_A',
+ ...
+ pipeline=train_pipeline
)
+)
```
### 拼接数据集
-有2种方式去拼接数据集。
-
-1. 如果您想拼接的数据集是同样的类型,但有不同的标注文件,
- 您可以按如下操作去拼接数据集的配置文件:
-
- 1. 您也许可以拼接两个标注文件夹 `ann_dir`
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = ['anno_dir_1', 'anno_dir_2'],
- pipeline=train_pipeline
- )
- ```
-
- 2. 您也可以去拼接两个 `split` 文件列表
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = 'anno_dir',
- split = ['split_1.txt', 'split_2.txt'],
- pipeline=train_pipeline
- )
- ```
+如果要拼接不同的数据集,可以按如下方式连接数据集配置。
- 3. 您也可以同时拼接 `ann_dir` 文件夹和 `split` 文件列表
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = ['anno_dir_1', 'anno_dir_2'],
- split = ['split_1.txt', 'split_2.txt'],
- pipeline=train_pipeline
- )
- ```
-
- 在这样的情况下, `ann_dir_1` 和 `ann_dir_2` 分别对应于 `split_1.txt` 和 `split_2.txt`
-
-2. 如果您想拼接不同的数据集,您可以如下去拼接数据集的配置文件:
-
- ```python
- dataset_A_train = dict()
- dataset_B_train = dict()
-
- data = dict(
- imgs_per_gpu=2,
- workers_per_gpu=2,
- train = [
- dataset_A_train,
- dataset_B_train
- ],
- val = dataset_A_val,
- test = dataset_A_test
- )
- ```
+```python
+dataset_A_train = dict()
+dataset_B_train = dict()
+concatenate_dataset = dict(
+ type='ConcatDataset',
+ datasets=[dataset_A_train, dataset_B_train])
+```
-一个更复杂的例子如下:分别重复 `Dataset_A` 和 `Dataset_B` N 次和 M 次,然后再去拼接重复后的数据集
+下面是一个更复杂的示例,它分别重复 `Dataset_A` 和 `Dataset_B` N 次和 M 次,然后连接重复的数据集。
```python
dataset_A_train = dict(
@@ -159,41 +154,36 @@ dataset_B_train = dict(
pipeline=train_pipeline
)
)
-data = dict(
- imgs_per_gpu=2,
- workers_per_gpu=2,
- train = [
- dataset_A_train,
- dataset_B_train
- ],
- val = dataset_A_val,
- test = dataset_A_test
-)
+train_dataloader = dict(
+ dataset=dict(
+ type='ConcatDataset',
+ datasets=[dataset_A_train, dataset_B_train]))
+
+val_dataloader = dict(dataset=dataset_A_val)
+test_dataloader = dict(dataset=dataset_A_test)
```
+您可以参考 mmengine 的基础数据集[教程](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/basedataset.html)以了解更多详细信息
+
### 多图混合集
-我们使用 `MultiImageMixDataset` 作为包装(wrapper)去混合多个数据集的图片。
-`MultiImageMixDataset`可以被类似mosaic和mixup的多图混合数据増广使用。
+我们使用 `MultiImageMixDataset` 作为包装(wrapper)去混合多个数据集的图片。
+`MultiImageMixDataset`可以被类似 mosaic 和 mixup 的多图混合数据増广使用。
-`MultiImageMixDataset`与`Mosaic`数据増广一起使用的例子:
+`MultiImageMixDataset` 与 `Mosaic` 数据増广一起使用的例子:
```python
train_pipeline = [
dict(type='RandomMosaic', prob=1),
dict(type='Resize', img_scale=(1024, 512), keep_ratio=True),
dict(type='RandomFlip', prob=0.5),
- dict(type='Normalize', **img_norm_cfg),
- dict(type='DefaultFormatBundle'),
- dict(type='Collect', keys=['img', 'gt_semantic_seg']),
+ dict(type='PackSegInputs')
]
train_dataset = dict(
type='MultiImageMixDataset',
dataset=dict(
- classes=classes,
- palette=palette,
type=dataset_type,
reduce_zero_label=False,
img_dir=data_root + "images/train",
diff --git a/docs/zh_cn/advanced_guides/add_metrics.md b/docs/zh_cn/advanced_guides/add_metrics.md
index 3a371e357e8..0637b447284 100644
--- a/docs/zh_cn/advanced_guides/add_metrics.md
+++ b/docs/zh_cn/advanced_guides/add_metrics.md
@@ -1 +1,81 @@
-# 新增评测指标 (待更新)
+# 新增评测指标
+
+## 使用 MMSegmentation 的源代码进行开发
+
+在这里,我们用 `CustomMetric` 作为例子来展示如何开发一个新的评测指标。
+
+1. 创建一个新文件 `mmseg/evaluation/metrics/custom_metric.py`。
+
+ ```python
+ from typing import List, Sequence
+
+ from mmengine.evaluator import BaseMetric
+
+ from mmseg.registry import METRICS
+
+
+ @METRICS.register_module()
+ class CustomMetric(BaseMetric):
+
+ def __init__(self, arg1, arg2):
+ """
+ The metric first processes each batch of data_samples and predictions,
+ and appends the processed results to the results list. Then it
+ collects all results together from all ranks if distributed training
+ is used. Finally, it computes the metrics of the entire dataset.
+ """
+
+ def process(self, data_batch: dict, data_samples: Sequence[dict]) -> None:
+ pass
+
+ def compute_metrics(self, results: list) -> dict:
+ pass
+
+ def evaluate(self, size: int) -> dict:
+ pass
+ ```
+
+ 在上面的示例中,`CustomMetric` 是 `BaseMetric` 的子类。它有三个方法:`process`,`compute_metrics` 和 `evaluate`。
+
+ - `process()` 处理一批数据样本和预测。处理后的结果需要显示地传给 `self.results` ,将在处理所有数据样本后用于计算指标。更多细节请参考 [MMEngine 文档](https://github.com/open-mmlab/mmengine/blob/main/docs/zh_cn/design/evaluation.md)
+
+ - `compute_metrics()` 用于从处理后的结果中计算指标。
+
+ - `evaluate()` 是一个接口,用于计算指标并返回结果。它将由 `ValLoop` 或 `TestLoop` 在 `Runner` 中调用。在大多数情况下,您不需要重写此方法,但如果您想做一些额外的工作,可以重写它。
+
+ **注意:** 您可以在[这里](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/loops.py#L366) 找到 `Runner` 调用 `evaluate()` 方法的过程。`Runner` 是训练和测试过程的执行器,您可以在[训练引擎文档](./engine.md)中找到有关它的详细信息。
+
+2. 在 `mmseg/evaluation/metrics/__init__.py` 中导入新的指标。
+
+ ```python
+ from .custom_metric import CustomMetric
+ __all__ = ['CustomMetric', ...]
+ ```
+
+3. 在配置文件中设置新的评测指标
+
+ ```python
+ val_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ test_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ ```
+
+## 使用发布版本的 MMSegmentation 进行开发
+
+上面的示例展示了如何使用 MMSegmentation 的源代码开发新指标。如果您想使用 MMSegmentation 的发布版本开发新指标,可以按照以下步骤操作。
+
+1. 创建一个新文件 `/Path/to/metrics/custom_metric.py`,实现 `process`,`compute_metrics` 和 `evaluate` 方法,`evaluate` 方法是可选的。
+
+2. 在代码或配置文件中导入新的指标。
+
+ ```python
+ from path.to.metrics import CustomMetric
+ ```
+
+ 或者
+
+ ```python
+ custom_imports = dict(imports=['/Path/to/metrics'], allow_failed_imports=False)
+
+ val_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ test_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ ```
diff --git a/docs/zh_cn/advanced_guides/add_models.md b/docs/zh_cn/advanced_guides/add_models.md
index 2f0a5af0d18..e05c07c8bac 100644
--- a/docs/zh_cn/advanced_guides/add_models.md
+++ b/docs/zh_cn/advanced_guides/add_models.md
@@ -166,7 +166,7 @@ loss_decode=dict(type='MyLoss', loss_weight=1.0))
### 添加新的数据预处理器(data preprocessor)
-在 MMSegmentation 1.x 版本中,我们使用 [SegDataPreProcessor](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/data_preprocessor.py#L13) 将数据复制到目标设备,并将数据预处理为默认的模型输入格式。这里我们将展示如何开发一个新的数据预处理器。
+在 MMSegmentation 1.x 版本中,我们使用 [SegDataPreProcessor](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/data_preprocessor.py#L13) 将数据复制到目标设备,并将数据预处理为默认的模型输入格式。这里我们将展示如何开发一个新的数据预处理器。
1. 创建一个新文件 `mmseg/models/my_datapreprocessor.py`。
diff --git a/docs/zh_cn/advanced_guides/contribute_dataset.md b/docs/zh_cn/advanced_guides/contribute_dataset.md
new file mode 100644
index 00000000000..4222de32a6c
--- /dev/null
+++ b/docs/zh_cn/advanced_guides/contribute_dataset.md
@@ -0,0 +1,461 @@
+# 在 mmsegmentation projects 中贡献一个标准格式的数据集
+
+- 在开始您的贡献流程前,请先阅读[《OpenMMLab 贡献代码指南》](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html),以详细的了解 OpenMMLab 代码库的代码贡献流程。
+- 该教程以 [Gaofen Image Dataset (GID)](https://www.sciencedirect.com/science/article/pii/S0034425719303414) 高分 2 号卫星所拍摄的遥感图像语义分割数据集作为样例,来演示在 mmsegmentation 中的数据集贡献流程。
+
+## 步骤 1: 配置 mmsegmentation 开发所需必要环境
+
+- 开发所必需的环境安装请参考[中文快速入门指南](https://github.com/open-mmlab/mmsegmentation/blob/main/docs/zh_cn/get_started.md)或[英文 get_started](https://github.com/open-mmlab/mmsegmentation/blob/main/docs/en/get_started.md)。
+
+- 如果您已安装了最新版的 pytorch、mmcv、mmengine,那么您可以跳过步骤 1 至[步骤 2](<#[步骤-2](#%E6%AD%A5%E9%AA%A4-2%E4%BB%A3%E7%A0%81%E8%B4%A1%E7%8C%AE%E5%89%8D%E7%9A%84%E5%87%86%E5%A4%87%E5%B7%A5%E4%BD%9C)>)。
+
+- **注:** 在此处无需安装 mmsegmentation,只需安装开发 mmsegmentation 所必需的 pytorch、mmcv、mmengine 等即可。
+
+**新建虚拟环境(如已有合适的开发环境,可跳过)**
+
+- 从[官方网站](https://docs.conda.io/en/latest/miniconda.html)下载并安装 Miniconda
+- 创建一个 conda 环境,并激活
+
+```shell
+conda create --name openmmlab python=3.8 -y
+conda activate openmmlab
+```
+
+**安装 pytorch (如环境下已安装 pytorch,可跳过)**
+
+- 参考 [official instructions](https://pytorch.org/get-started/locally/) 安装 **PyTorch**
+
+**使用 mim 安装 mmcv、mmengine**
+
+- 使用 [MIM](https://github.com/open-mmlab/mim) 安装 [MMCV](https://github.com/open-mmlab/mmcv)
+
+```shell
+pip install -U openmim
+mim install mmengine
+mim install "mmcv>=2.0.0"
+```
+
+## 步骤 2:代码贡献前的准备工作
+
+### 2.1 Fork mmsegmentation 仓库
+
+- 通过浏览器打开[mmsegmentation 官方仓库](https://github.com/open-mmlab/mmsegmentation/tree/main)。
+- 登录您的 GitHub 账户,以下步骤均需在 GitHub 登录的情况下进行。
+- Fork mmsegmentation 仓库
+ 
+- Fork 之后,mmsegmentation 仓库将会出现在您的个人仓库中。
+
+### 2.2 在您的代码编写软件中 git clone mmsegmentation
+
+这里以 VSCODE 为例
+
+- 打开 VSCODE,新建终端窗口并激活您在[步骤 1 ](#%E6%AD%A5%E9%AA%A4-1-%E9%85%8D%E7%BD%AE-mmsegmentation-%E5%BC%80%E5%8F%91%E6%89%80%E9%9C%80%E5%BF%85%E8%A6%81%E7%8E%AF%E5%A2%83)中所安装的虚拟环境。
+- 在您 GitHub 的个人仓库中找到您 Fork 的 mmsegmentation 仓库,复制其链接。
+ 
+- 在终端中执行命令
+ ```bash
+ git clone {您所复制的个人仓库的链接}
+ ```
+ 
+ **注:** 如提示以下信息,请在 GitHub 中添加 [SSH 秘钥](https://docs.github.com/en/authentication/connecting-to-github-with-ssh/generating-a-new-ssh-key-and-adding-it-to-the-ssh-agent)
+ 
+- 进入 mmsegmentation 目录(之后的操作均在 mmsegmentation 目录下)。
+ ```bash
+ cd mmsegmentation
+ ```
+- 在终端中执行以下命令,添加官方仓库为上游仓库。
+ ```bash
+ git remote add upstream git@github.com:open-mmlab/mmsegmentation.git
+ ```
+- 使用以下命令检查 remote 是否添加成功。
+ ```bash
+ git remote -v
+ ```
+ 
+
+### 2.3 切换目录至 mmsegmentation 并从源码安装mmsegmentation
+
+在`mmsegmentation`目录下执行`pip install -v -e .`,通过源码构建方式安装 mmsegmentaion 库。
+安装完成后,您将能看到如下图所示的文件树。
+
diff --git a/docs/zh_cn/advanced_guides/add_datasets.md b/docs/zh_cn/advanced_guides/add_datasets.md
index 4ea14934ed2..22fbf3462fc 100644
--- a/docs/zh_cn/advanced_guides/add_datasets.md
+++ b/docs/zh_cn/advanced_guides/add_datasets.md
@@ -1,4 +1,62 @@
-# 新增自定义数据集(待更新)
+# 新增自定义数据集
+
+## 新增自定义数据集
+
+在这里,我们展示如何构建一个新的数据集。
+
+1. 创建一个新文件 `mmseg/datasets/example.py`
+
+ ```python
+ from mmseg.registry import DATASETS
+ from .basesegdataset import BaseSegDataset
+
+
+ @DATASETS.register_module()
+ class ExampleDataset(BaseSegDataset):
+
+ METAINFO = dict(
+ classes=('xxx', 'xxx', ...),
+ palette=[[x, x, x], [x, x, x], ...])
+
+ def __init__(self, aeg1, arg2):
+ pass
+ ```
+
+2. 在 `mmseg/datasets/__init__.py` 中导入模块
+
+ ```python
+ from .example import ExampleDataset
+ ```
+
+3. 通过创建一个新的数据集配置文件 `configs/_base_/datasets/example_dataset.py` 来使用它
+
+ ```python
+ dataset_type = 'ExampleDataset'
+ data_root = 'data/example/'
+ ...
+ ```
+
+4. 在 `mmseg/utils/class_names.py` 中补充数据集元信息
+
+ ```python
+ def example_classes():
+ return [
+ 'xxx', 'xxx',
+ ...
+ ]
+
+ def example_palette():
+ return [
+ [x, x, x], [x, x, x],
+ ...
+ ]
+ dataset_aliases ={
+ 'example': ['example', ...],
+ ...
+ }
+ ```
+
+**注意:** 如果新数据集不满足 mmseg 的要求,则需要在 `tools/dataset_converters/` 中准备一个数据集预处理脚本
## 通过重新组织数据来定制数据集
@@ -26,30 +84,17 @@
一个训练对将由 img_dir/ann_dir 里同样首缀的文件组成。
-如果给定 `split` 参数,只有部分在 img_dir/ann_dir 里的文件会被加载。
-我们可以对被包括在 split 文本里的文件指定前缀。
+有些数据集不会发布测试集或测试集的标注,如果没有测试集的标注,我们就无法在本地进行评估模型,因此我们在配置文件中将验证集设置为默认测试集。
-除此以外,一个 split 文本如下所示:
-
-```none
-xxx
-zzz
-```
+关于如何构建自己的数据集或实现新的数据集类,请参阅[数据集指南](./datasets.md)以获取更多详细信息。
-只有
-
-`data/my_dataset/img_dir/train/xxx{img_suffix}`,
-`data/my_dataset/img_dir/train/zzz{img_suffix}`,
-`data/my_dataset/ann_dir/train/xxx{seg_map_suffix}`,
-`data/my_dataset/ann_dir/train/zzz{seg_map_suffix}` 将被加载。
-
-注意:标注是跟图像同样的形状 (H, W),其中的像素值的范围是 `[0, num_classes - 1]`。
+**注意:** 标注是跟图像同样的形状 (H, W),其中的像素值的范围是 `[0, num_classes - 1]`。
您也可以使用 [pillow](https://pillow.readthedocs.io/en/stable/handbook/concepts.html#palette) 的 `'P'` 模式去创建包含颜色的标注。
## 通过混合数据去定制数据集
MMSegmentation 同样支持混合数据集去训练。
-当前它支持拼接 (concat), 重复 (repeat) 和多图混合 (multi-image mix)数据集。
+当前它支持拼接 (concat), 重复 (repeat) 和多图混合 (multi-image mix) 数据集。
### 重复数据集
@@ -58,79 +103,29 @@ MMSegmentation 同样支持混合数据集去训练。
```python
dataset_A_train = dict(
- type='RepeatDataset',
- times=N,
- dataset=dict( # 这是 Dataset_A 数据集的原始配置
- type='Dataset_A',
- ...
- pipeline=train_pipeline
- )
+ type='RepeatDataset',
+ times=N,
+ dataset=dict( # 这是 Dataset_A 数据集的原始配置
+ type='Dataset_A',
+ ...
+ pipeline=train_pipeline
)
+)
```
### 拼接数据集
-有2种方式去拼接数据集。
-
-1. 如果您想拼接的数据集是同样的类型,但有不同的标注文件,
- 您可以按如下操作去拼接数据集的配置文件:
-
- 1. 您也许可以拼接两个标注文件夹 `ann_dir`
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = ['anno_dir_1', 'anno_dir_2'],
- pipeline=train_pipeline
- )
- ```
-
- 2. 您也可以去拼接两个 `split` 文件列表
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = 'anno_dir',
- split = ['split_1.txt', 'split_2.txt'],
- pipeline=train_pipeline
- )
- ```
+如果要拼接不同的数据集,可以按如下方式连接数据集配置。
- 3. 您也可以同时拼接 `ann_dir` 文件夹和 `split` 文件列表
-
- ```python
- dataset_A_train = dict(
- type='Dataset_A',
- img_dir = 'img_dir',
- ann_dir = ['anno_dir_1', 'anno_dir_2'],
- split = ['split_1.txt', 'split_2.txt'],
- pipeline=train_pipeline
- )
- ```
-
- 在这样的情况下, `ann_dir_1` 和 `ann_dir_2` 分别对应于 `split_1.txt` 和 `split_2.txt`
-
-2. 如果您想拼接不同的数据集,您可以如下去拼接数据集的配置文件:
-
- ```python
- dataset_A_train = dict()
- dataset_B_train = dict()
-
- data = dict(
- imgs_per_gpu=2,
- workers_per_gpu=2,
- train = [
- dataset_A_train,
- dataset_B_train
- ],
- val = dataset_A_val,
- test = dataset_A_test
- )
- ```
+```python
+dataset_A_train = dict()
+dataset_B_train = dict()
+concatenate_dataset = dict(
+ type='ConcatDataset',
+ datasets=[dataset_A_train, dataset_B_train])
+```
-一个更复杂的例子如下:分别重复 `Dataset_A` 和 `Dataset_B` N 次和 M 次,然后再去拼接重复后的数据集
+下面是一个更复杂的示例,它分别重复 `Dataset_A` 和 `Dataset_B` N 次和 M 次,然后连接重复的数据集。
```python
dataset_A_train = dict(
@@ -159,41 +154,36 @@ dataset_B_train = dict(
pipeline=train_pipeline
)
)
-data = dict(
- imgs_per_gpu=2,
- workers_per_gpu=2,
- train = [
- dataset_A_train,
- dataset_B_train
- ],
- val = dataset_A_val,
- test = dataset_A_test
-)
+train_dataloader = dict(
+ dataset=dict(
+ type='ConcatDataset',
+ datasets=[dataset_A_train, dataset_B_train]))
+
+val_dataloader = dict(dataset=dataset_A_val)
+test_dataloader = dict(dataset=dataset_A_test)
```
+您可以参考 mmengine 的基础数据集[教程](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/basedataset.html)以了解更多详细信息
+
### 多图混合集
-我们使用 `MultiImageMixDataset` 作为包装(wrapper)去混合多个数据集的图片。
-`MultiImageMixDataset`可以被类似mosaic和mixup的多图混合数据増广使用。
+我们使用 `MultiImageMixDataset` 作为包装(wrapper)去混合多个数据集的图片。
+`MultiImageMixDataset`可以被类似 mosaic 和 mixup 的多图混合数据増广使用。
-`MultiImageMixDataset`与`Mosaic`数据増广一起使用的例子:
+`MultiImageMixDataset` 与 `Mosaic` 数据増广一起使用的例子:
```python
train_pipeline = [
dict(type='RandomMosaic', prob=1),
dict(type='Resize', img_scale=(1024, 512), keep_ratio=True),
dict(type='RandomFlip', prob=0.5),
- dict(type='Normalize', **img_norm_cfg),
- dict(type='DefaultFormatBundle'),
- dict(type='Collect', keys=['img', 'gt_semantic_seg']),
+ dict(type='PackSegInputs')
]
train_dataset = dict(
type='MultiImageMixDataset',
dataset=dict(
- classes=classes,
- palette=palette,
type=dataset_type,
reduce_zero_label=False,
img_dir=data_root + "images/train",
diff --git a/docs/zh_cn/advanced_guides/add_metrics.md b/docs/zh_cn/advanced_guides/add_metrics.md
index 3a371e357e8..0637b447284 100644
--- a/docs/zh_cn/advanced_guides/add_metrics.md
+++ b/docs/zh_cn/advanced_guides/add_metrics.md
@@ -1 +1,81 @@
-# 新增评测指标 (待更新)
+# 新增评测指标
+
+## 使用 MMSegmentation 的源代码进行开发
+
+在这里,我们用 `CustomMetric` 作为例子来展示如何开发一个新的评测指标。
+
+1. 创建一个新文件 `mmseg/evaluation/metrics/custom_metric.py`。
+
+ ```python
+ from typing import List, Sequence
+
+ from mmengine.evaluator import BaseMetric
+
+ from mmseg.registry import METRICS
+
+
+ @METRICS.register_module()
+ class CustomMetric(BaseMetric):
+
+ def __init__(self, arg1, arg2):
+ """
+ The metric first processes each batch of data_samples and predictions,
+ and appends the processed results to the results list. Then it
+ collects all results together from all ranks if distributed training
+ is used. Finally, it computes the metrics of the entire dataset.
+ """
+
+ def process(self, data_batch: dict, data_samples: Sequence[dict]) -> None:
+ pass
+
+ def compute_metrics(self, results: list) -> dict:
+ pass
+
+ def evaluate(self, size: int) -> dict:
+ pass
+ ```
+
+ 在上面的示例中,`CustomMetric` 是 `BaseMetric` 的子类。它有三个方法:`process`,`compute_metrics` 和 `evaluate`。
+
+ - `process()` 处理一批数据样本和预测。处理后的结果需要显示地传给 `self.results` ,将在处理所有数据样本后用于计算指标。更多细节请参考 [MMEngine 文档](https://github.com/open-mmlab/mmengine/blob/main/docs/zh_cn/design/evaluation.md)
+
+ - `compute_metrics()` 用于从处理后的结果中计算指标。
+
+ - `evaluate()` 是一个接口,用于计算指标并返回结果。它将由 `ValLoop` 或 `TestLoop` 在 `Runner` 中调用。在大多数情况下,您不需要重写此方法,但如果您想做一些额外的工作,可以重写它。
+
+ **注意:** 您可以在[这里](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/loops.py#L366) 找到 `Runner` 调用 `evaluate()` 方法的过程。`Runner` 是训练和测试过程的执行器,您可以在[训练引擎文档](./engine.md)中找到有关它的详细信息。
+
+2. 在 `mmseg/evaluation/metrics/__init__.py` 中导入新的指标。
+
+ ```python
+ from .custom_metric import CustomMetric
+ __all__ = ['CustomMetric', ...]
+ ```
+
+3. 在配置文件中设置新的评测指标
+
+ ```python
+ val_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ test_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ ```
+
+## 使用发布版本的 MMSegmentation 进行开发
+
+上面的示例展示了如何使用 MMSegmentation 的源代码开发新指标。如果您想使用 MMSegmentation 的发布版本开发新指标,可以按照以下步骤操作。
+
+1. 创建一个新文件 `/Path/to/metrics/custom_metric.py`,实现 `process`,`compute_metrics` 和 `evaluate` 方法,`evaluate` 方法是可选的。
+
+2. 在代码或配置文件中导入新的指标。
+
+ ```python
+ from path.to.metrics import CustomMetric
+ ```
+
+ 或者
+
+ ```python
+ custom_imports = dict(imports=['/Path/to/metrics'], allow_failed_imports=False)
+
+ val_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ test_evaluator = dict(type='CustomMetric', arg1=xxx, arg2=xxx)
+ ```
diff --git a/docs/zh_cn/advanced_guides/add_models.md b/docs/zh_cn/advanced_guides/add_models.md
index 2f0a5af0d18..e05c07c8bac 100644
--- a/docs/zh_cn/advanced_guides/add_models.md
+++ b/docs/zh_cn/advanced_guides/add_models.md
@@ -166,7 +166,7 @@ loss_decode=dict(type='MyLoss', loss_weight=1.0))
### 添加新的数据预处理器(data preprocessor)
-在 MMSegmentation 1.x 版本中,我们使用 [SegDataPreProcessor](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/data_preprocessor.py#L13) 将数据复制到目标设备,并将数据预处理为默认的模型输入格式。这里我们将展示如何开发一个新的数据预处理器。
+在 MMSegmentation 1.x 版本中,我们使用 [SegDataPreProcessor](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/data_preprocessor.py#L13) 将数据复制到目标设备,并将数据预处理为默认的模型输入格式。这里我们将展示如何开发一个新的数据预处理器。
1. 创建一个新文件 `mmseg/models/my_datapreprocessor.py`。
diff --git a/docs/zh_cn/advanced_guides/contribute_dataset.md b/docs/zh_cn/advanced_guides/contribute_dataset.md
new file mode 100644
index 00000000000..4222de32a6c
--- /dev/null
+++ b/docs/zh_cn/advanced_guides/contribute_dataset.md
@@ -0,0 +1,461 @@
+# 在 mmsegmentation projects 中贡献一个标准格式的数据集
+
+- 在开始您的贡献流程前,请先阅读[《OpenMMLab 贡献代码指南》](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html),以详细的了解 OpenMMLab 代码库的代码贡献流程。
+- 该教程以 [Gaofen Image Dataset (GID)](https://www.sciencedirect.com/science/article/pii/S0034425719303414) 高分 2 号卫星所拍摄的遥感图像语义分割数据集作为样例,来演示在 mmsegmentation 中的数据集贡献流程。
+
+## 步骤 1: 配置 mmsegmentation 开发所需必要环境
+
+- 开发所必需的环境安装请参考[中文快速入门指南](https://github.com/open-mmlab/mmsegmentation/blob/main/docs/zh_cn/get_started.md)或[英文 get_started](https://github.com/open-mmlab/mmsegmentation/blob/main/docs/en/get_started.md)。
+
+- 如果您已安装了最新版的 pytorch、mmcv、mmengine,那么您可以跳过步骤 1 至[步骤 2](<#[步骤-2](#%E6%AD%A5%E9%AA%A4-2%E4%BB%A3%E7%A0%81%E8%B4%A1%E7%8C%AE%E5%89%8D%E7%9A%84%E5%87%86%E5%A4%87%E5%B7%A5%E4%BD%9C)>)。
+
+- **注:** 在此处无需安装 mmsegmentation,只需安装开发 mmsegmentation 所必需的 pytorch、mmcv、mmengine 等即可。
+
+**新建虚拟环境(如已有合适的开发环境,可跳过)**
+
+- 从[官方网站](https://docs.conda.io/en/latest/miniconda.html)下载并安装 Miniconda
+- 创建一个 conda 环境,并激活
+
+```shell
+conda create --name openmmlab python=3.8 -y
+conda activate openmmlab
+```
+
+**安装 pytorch (如环境下已安装 pytorch,可跳过)**
+
+- 参考 [official instructions](https://pytorch.org/get-started/locally/) 安装 **PyTorch**
+
+**使用 mim 安装 mmcv、mmengine**
+
+- 使用 [MIM](https://github.com/open-mmlab/mim) 安装 [MMCV](https://github.com/open-mmlab/mmcv)
+
+```shell
+pip install -U openmim
+mim install mmengine
+mim install "mmcv>=2.0.0"
+```
+
+## 步骤 2:代码贡献前的准备工作
+
+### 2.1 Fork mmsegmentation 仓库
+
+- 通过浏览器打开[mmsegmentation 官方仓库](https://github.com/open-mmlab/mmsegmentation/tree/main)。
+- 登录您的 GitHub 账户,以下步骤均需在 GitHub 登录的情况下进行。
+- Fork mmsegmentation 仓库
+ 
+- Fork 之后,mmsegmentation 仓库将会出现在您的个人仓库中。
+
+### 2.2 在您的代码编写软件中 git clone mmsegmentation
+
+这里以 VSCODE 为例
+
+- 打开 VSCODE,新建终端窗口并激活您在[步骤 1 ](#%E6%AD%A5%E9%AA%A4-1-%E9%85%8D%E7%BD%AE-mmsegmentation-%E5%BC%80%E5%8F%91%E6%89%80%E9%9C%80%E5%BF%85%E8%A6%81%E7%8E%AF%E5%A2%83)中所安装的虚拟环境。
+- 在您 GitHub 的个人仓库中找到您 Fork 的 mmsegmentation 仓库,复制其链接。
+ 
+- 在终端中执行命令
+ ```bash
+ git clone {您所复制的个人仓库的链接}
+ ```
+ 
+ **注:** 如提示以下信息,请在 GitHub 中添加 [SSH 秘钥](https://docs.github.com/en/authentication/connecting-to-github-with-ssh/generating-a-new-ssh-key-and-adding-it-to-the-ssh-agent)
+ 
+- 进入 mmsegmentation 目录(之后的操作均在 mmsegmentation 目录下)。
+ ```bash
+ cd mmsegmentation
+ ```
+- 在终端中执行以下命令,添加官方仓库为上游仓库。
+ ```bash
+ git remote add upstream git@github.com:open-mmlab/mmsegmentation.git
+ ```
+- 使用以下命令检查 remote 是否添加成功。
+ ```bash
+ git remote -v
+ ```
+ 
+
+### 2.3 切换目录至 mmsegmentation 并从源码安装mmsegmentation
+
+在`mmsegmentation`目录下执行`pip install -v -e .`,通过源码构建方式安装 mmsegmentaion 库。
+安装完成后,您将能看到如下图所示的文件树。
+ +
+### 2.4 切换分支为 dev-1.x
+
+正如您在[ mmsegmentation 官网](https://github.com/open-mmlab/mmsegmentation/tree/main)所见,该仓库有许多分支,默认分支`main`为稳定的发行版本,以及用于贡献者进行开发的`dev-1.x`分支。`dev-1.x`分支是贡献者们用来提交创意和 PR 的分支,`dev-1.x`分支的内容会被周期性的合入到`main`分支。
+
+
+回到 VSCODE 中,在终端执行命令
+
+```bash
+git checkout dev-1.x
+```
+
+### 2.5 创新属于自己的新分支
+
+在基于`dev-1.x`分支下,使用如下命令,创建属于您自己的分支。
+
+```bash
+# git checkout -b 您的GitHubID/您的分支想要实现的功能的名字
+# git checkout -b AI-Tianlong/support_GID_dataset
+git checkout -b {您的GitHubID/您的分支想要实现的功能的名字}
+```
+
+### 2.6 配置 pre-commit
+
+OpenMMLab 仓库对代码质量有着较高的要求,所有提交的 PR 必须要通过代码格式检查。pre-commit 详细配置参阅[配置 pre-commit](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html#pre-commit)。
+
+## 步骤 3:在`mmsegmentation/projects`下贡献您的代码
+
+**先对 GID 数据集进行分析**
+
+这里以贡献高分 2 号遥感图像语义分割数据集 GID 为例,GID 数据集是由我国自主研发的高分 2 号卫星所拍摄的光学遥感图像所创建,经图像预处理后共提供了 150 张 6800x7200 像素的 RGB 三通道遥感图像。并提供了两种不同类别数的数据标注,一种是包含 5 类有效物体的 RGB 标签,另一种是包含 15 类有效物体的 RGB 标签。本教程将针对 5 类标签进行数据集贡献流程讲解。
+
+GID 的 5 类有效标签分别为:0-背景-\[0,0,0\](mask 标签值-标签名称-RGB 标签值)、1-建筑-\[255,0,0\]、2-农田-\[0,255,0\]、3-森林-\[0,0,255\]、4-草地-\[255,255,0\]、5-水-\[0,0,255\]。在语义分割任务中,标签是与原图尺寸一致的单通道图像,标签图像中的像素值为真实样本图像中对应像素所包含的物体的类别。GID 数据集提供的是具有 RGB 三通道的彩色标签,为了模型的训练需要将 RGB 标签转换为 mask 标签。并且由于图像尺寸为 6800x7200 像素,对于神经网络的训练来有些过大,所以将每张图像裁切成了没有重叠的 512x512 的图像以便进行训练。
+
+
+### 2.4 切换分支为 dev-1.x
+
+正如您在[ mmsegmentation 官网](https://github.com/open-mmlab/mmsegmentation/tree/main)所见,该仓库有许多分支,默认分支`main`为稳定的发行版本,以及用于贡献者进行开发的`dev-1.x`分支。`dev-1.x`分支是贡献者们用来提交创意和 PR 的分支,`dev-1.x`分支的内容会被周期性的合入到`main`分支。
+
+
+回到 VSCODE 中,在终端执行命令
+
+```bash
+git checkout dev-1.x
+```
+
+### 2.5 创新属于自己的新分支
+
+在基于`dev-1.x`分支下,使用如下命令,创建属于您自己的分支。
+
+```bash
+# git checkout -b 您的GitHubID/您的分支想要实现的功能的名字
+# git checkout -b AI-Tianlong/support_GID_dataset
+git checkout -b {您的GitHubID/您的分支想要实现的功能的名字}
+```
+
+### 2.6 配置 pre-commit
+
+OpenMMLab 仓库对代码质量有着较高的要求,所有提交的 PR 必须要通过代码格式检查。pre-commit 详细配置参阅[配置 pre-commit](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html#pre-commit)。
+
+## 步骤 3:在`mmsegmentation/projects`下贡献您的代码
+
+**先对 GID 数据集进行分析**
+
+这里以贡献高分 2 号遥感图像语义分割数据集 GID 为例,GID 数据集是由我国自主研发的高分 2 号卫星所拍摄的光学遥感图像所创建,经图像预处理后共提供了 150 张 6800x7200 像素的 RGB 三通道遥感图像。并提供了两种不同类别数的数据标注,一种是包含 5 类有效物体的 RGB 标签,另一种是包含 15 类有效物体的 RGB 标签。本教程将针对 5 类标签进行数据集贡献流程讲解。
+
+GID 的 5 类有效标签分别为:0-背景-\[0,0,0\](mask 标签值-标签名称-RGB 标签值)、1-建筑-\[255,0,0\]、2-农田-\[0,255,0\]、3-森林-\[0,0,255\]、4-草地-\[255,255,0\]、5-水-\[0,0,255\]。在语义分割任务中,标签是与原图尺寸一致的单通道图像,标签图像中的像素值为真实样本图像中对应像素所包含的物体的类别。GID 数据集提供的是具有 RGB 三通道的彩色标签,为了模型的训练需要将 RGB 标签转换为 mask 标签。并且由于图像尺寸为 6800x7200 像素,对于神经网络的训练来有些过大,所以将每张图像裁切成了没有重叠的 512x512 的图像以便进行训练。
+ +
+### 3.1 在`mmsegmentation/projects`下创建新的项目文件夹
+
+在`mmsegmentation/projects`下创建文件夹`gid_dataset`
+
+
+### 3.2 贡献您的数据集代码
+
+为了最终能将您在 projects 中贡献的代码更加顺畅的移入核心库中(对代码要求质量更高),非常建议按照核心库的目录来编辑您的数据集文件。
+关于数据集有 4 个必要的文件:
+
+- **1** `mmseg/datasets/gid.py` 定义了数据集的尾缀、CLASSES、PALETTE、reduce_zero_label等
+- **2** `configs/_base_/gid.py` GID 数据集的配置文件,定义了数据集的`dataset_type`(数据集类型,`mmseg/datasets/gid.py`中注册的数据集的类名)、`data_root`(数据集所在的根目录,建议将数据集通过软连接的方式将数据集放至`mmsegmentation/data`)、`train_pipline`(训练的数据流)、`test_pipline`(测试和验证时的数据流)、`img_rations`(多尺度预测时的多尺度配置)、`tta_pipeline`(多尺度预测)、`train_dataloader`(训练集的数据加载器)、`val_dataloader`(验证集的数据加载器)、`test_dataloader`(测试集的数据加载器)、`val_evaluator`(验证集的评估器)、`test_evaluator`(测试集的评估器)。
+- **3** 使用了 GID 数据集的模型训练配置文件
+ 这个是可选的,但是强烈建议您添加。在核心库中,所贡献的数据集需要和参考文献中所提出的结果精度对齐,为了后期将您贡献的代码合并入核心库。如您的算力充足,最好能提供对应的模型配置文件在您贡献的数据集上所验证的结果以及相应的权重文件,并撰写较为详细的README.md文档。[示例参考结果](https://github.com/open-mmlab/mmsegmentation/tree/main/configs/deeplabv3plus#mapillary-vistas-v12)
+ 
+- **4** 使用如下命令格式: 撰写`docs/zh_cn/user_guides/2_dataset_prepare.md`来添加您的数据集介绍,包括但不限于数据集的下载方式,数据集目录结构、数据集生成等一些必要性的文字性描述和运行命令。以更好地帮助用户能更快的实现数据集的准备工作。
+
+### 3.3 贡献`tools/dataset_converters/gid.py`
+
+由于 GID 数据集是由未经过切分的 6800x7200 图像所构成的数据集,并且没有划分训练集、验证集与测试集。以及其标签为 RGB 彩色标签,需要将标签转换为单通道的 mask label。为了方便训练,首先将 GID 数据集进行裁切和标签转换,并进行数据集划分,构建为 mmsegmentation 所支持的格式。
+
+```python
+# tools/dataset_converters/gid.py
+import argparse
+import glob
+import math
+import os
+import os.path as osp
+from PIL import Image
+
+import mmcv
+import numpy as np
+from mmengine.utils import ProgressBar, mkdir_or_exist
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='Convert GID dataset to mmsegmentation format')
+ parser.add_argument('dataset_img_path', help='GID images folder path')
+ parser.add_argument('dataset_label_path', help='GID labels folder path')
+ parser.add_argument('--tmp_dir', help='path of the temporary directory')
+ parser.add_argument('-o', '--out_dir', help='output path', default='data/gid')
+ parser.add_argument(
+ '--clip_size',
+ type=int,
+ help='clipped size of image after preparation',
+ default=256)
+ parser.add_argument(
+ '--stride_size',
+ type=int,
+ help='stride of clipping original images',
+ default=256)
+ args = parser.parse_args()
+ return args
+
+GID_COLORMAP = dict(
+ Background=(0, 0, 0), #0-背景-黑色
+ Building=(255, 0, 0), #1-建筑-红色
+ Farmland=(0, 255, 0), #2-农田-绿色
+ Forest=(0, 0, 255), #3-森林-蓝色
+ Meadow=(255, 255, 0),#4-草地-黄色
+ Water=(0, 0, 255)#5-水-蓝色
+)
+palette = list(GID_COLORMAP.values())
+classes = list(GID_COLORMAP.keys())
+
+#############用列表来存一个 RGB 和一个类别的对应################
+def colormap2label(palette):
+ colormap2label_list = np.zeros(256**3, dtype = np.longlong)
+ for i, colormap in enumerate(palette):
+ colormap2label_list[(colormap[0] * 256 + colormap[1])*256+colormap[2]] = i
+ return colormap2label_list
+
+#############给定那个列表,和vis_png然后生成masks_png################
+def label_indices(RGB_label, colormap2label_list):
+ RGB_label = RGB_label.astype('int32')
+ idx = (RGB_label[:, :, 0] * 256 + RGB_label[:, :, 1]) * 256 + RGB_label[:, :, 2]
+ # print(idx.shape)
+ return colormap2label_list[idx]
+
+def RGB2mask(RGB_label, colormap2label_list):
+ # RGB_label = np.array(Image.open(RGB_label).convert('RGB')) #打开RGB_png
+ mask_label = label_indices(RGB_label, colormap2label_list) # .numpy()
+ return mask_label
+
+colormap2label_list = colormap2label(palette)
+
+def clip_big_image(image_path, clip_save_dir, args, to_label=False):
+ """
+ Original image of GID dataset is very large, thus pre-processing
+ of them is adopted. Given fixed clip size and stride size to generate
+ clipped image, the intersection of width and height is determined.
+ For example, given one 6800 x 7200 original image, the clip size is
+ 256 and stride size is 256, thus it would generate 29 x 27 = 783 images
+ whose size are all 256 x 256.
+
+ """
+
+ image = mmcv.imread(image_path, channel_order='rgb')
+ # image = mmcv.bgr2gray(image)
+
+ h, w, c = image.shape
+ clip_size = args.clip_size
+ stride_size = args.stride_size
+
+ num_rows = math.ceil((h - clip_size) / stride_size) if math.ceil(
+ (h - clip_size) /
+ stride_size) * stride_size + clip_size >= h else math.ceil(
+ (h - clip_size) / stride_size) + 1
+ num_cols = math.ceil((w - clip_size) / stride_size) if math.ceil(
+ (w - clip_size) /
+ stride_size) * stride_size + clip_size >= w else math.ceil(
+ (w - clip_size) / stride_size) + 1
+
+ x, y = np.meshgrid(np.arange(num_cols + 1), np.arange(num_rows + 1))
+ xmin = x * clip_size
+ ymin = y * clip_size
+
+ xmin = xmin.ravel()
+ ymin = ymin.ravel()
+ xmin_offset = np.where(xmin + clip_size > w, w - xmin - clip_size,
+ np.zeros_like(xmin))
+ ymin_offset = np.where(ymin + clip_size > h, h - ymin - clip_size,
+ np.zeros_like(ymin))
+ boxes = np.stack([
+ xmin + xmin_offset, ymin + ymin_offset,
+ np.minimum(xmin + clip_size, w),
+ np.minimum(ymin + clip_size, h)
+ ], axis=1)
+
+ if to_label:
+ image = RGB2mask(image, colormap2label_list) #这里得改一下
+
+ for count, box in enumerate(boxes):
+ start_x, start_y, end_x, end_y = box
+ clipped_image = image[start_y:end_y,
+ start_x:end_x] if to_label else image[
+ start_y:end_y, start_x:end_x, :]
+ img_name = osp.basename(image_path).replace('.tif', '')
+ img_name = img_name.replace('_label', '')
+ if count % 3 == 0:
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(
+ clip_save_dir.replace('train', 'val'),
+ f'{img_name}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+ else:
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(
+ clip_save_dir,
+ f'{img_name}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+ count += 1
+
+def main():
+ args = parse_args()
+
+ """
+ According to this paper: https://ieeexplore.ieee.org/document/9343296/
+ select 15 images contained in GID, , which cover the whole six
+ categories, to generate train set and validation set.
+
+ According to Paper: https://ieeexplore.ieee.org/document/9343296/
+
+ """
+
+ if args.out_dir is None:
+ out_dir = osp.join('data', 'gid')
+ else:
+ out_dir = args.out_dir
+
+ print('Making directories...')
+ mkdir_or_exist(osp.join(out_dir, 'img_dir', 'train'))
+ mkdir_or_exist(osp.join(out_dir, 'img_dir', 'val'))
+ mkdir_or_exist(osp.join(out_dir, 'ann_dir', 'train'))
+ mkdir_or_exist(osp.join(out_dir, 'ann_dir', 'val'))
+
+ src_path_list = glob.glob(os.path.join(args.dataset_img_path, '*.tif'))
+ print(f'Find {len(src_path_list)} pictures')
+
+ prog_bar = ProgressBar(len(src_path_list))
+
+ dst_img_dir = osp.join(out_dir, 'img_dir', 'train')
+ dst_label_dir = osp.join(out_dir, 'ann_dir', 'train')
+
+ for i, img_path in enumerate(src_path_list):
+ label_path = osp.join(args.dataset_label_path, osp.basename(img_path.replace('.tif', '_label.tif')))
+
+ clip_big_image(img_path, dst_img_dir, args, to_label=False)
+ clip_big_image(label_path, dst_label_dir, args, to_label=True)
+ prog_bar.update()
+
+ print('Done!')
+
+if __name__ == '__main__':
+ main()
+```
+
+### 3.4 贡献`mmseg/datasets/gid.py`
+
+可参考[`projects/mapillary_dataset/mmseg/datasets/mapillary.py`](https://github.com/open-mmlab/mmsegmentation/blob/main/projects/mapillary_dataset/mmseg/datasets/mapillary.py)并在此基础上修改相应变量以适配您的数据集。
+
+```python
+# mmseg/datasets/gid.py
+# Copyright (c) OpenMMLab. All rights reserved.
+from mmseg.datasets.basesegdataset import BaseSegDataset
+from mmseg.registry import DATASETS
+
+# 注册数据集类
+@DATASETS.register_module()
+class GID_Dataset(BaseSegDataset):
+ """Gaofen Image Dataset (GID)
+
+ Dataset paper link:
+ https://www.sciencedirect.com/science/article/pii/S0034425719303414
+ https://x-ytong.github.io/project/GID.html
+
+ GID 6 classes: background(others), built-up, farmland, forest, meadow, water
+
+ In This example, select 10 images from GID dataset as training set,
+ and select 5 images as validation set.
+ The selected images are listed as follows:
+
+ GF2_PMS1__L1A0000647767-MSS1
+ GF2_PMS1__L1A0001064454-MSS1
+ GF2_PMS1__L1A0001348919-MSS1
+ GF2_PMS1__L1A0001680851-MSS1
+ GF2_PMS1__L1A0001680853-MSS1
+ GF2_PMS1__L1A0001680857-MSS1
+ GF2_PMS1__L1A0001757429-MSS1
+ GF2_PMS2__L1A0000607681-MSS2
+ GF2_PMS2__L1A0000635115-MSS2
+ GF2_PMS2__L1A0000658637-MSS2
+ GF2_PMS2__L1A0001206072-MSS2
+ GF2_PMS2__L1A0001471436-MSS2
+ GF2_PMS2__L1A0001642620-MSS2
+ GF2_PMS2__L1A0001787089-MSS2
+ GF2_PMS2__L1A0001838560-MSS2
+
+ The ``img_suffix`` is fixed to '.tif' and ``seg_map_suffix`` is
+ fixed to '.tif' for GID.
+ """
+ METAINFO = dict(
+ classes=('Others', 'Built-up', 'Farmland', 'Forest',
+ 'Meadow', 'Water'),
+
+ palette=[[0, 0, 0], [255, 0, 0], [0, 255, 0], [0, 255, 255],
+ [255, 255, 0], [0, 0, 255]])
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=None,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
+```
+
+### 3.5 贡献使用 GID 的训练 config file
+
+```python
+_base_ = [
+ '../../../configs/_base_/models/deeplabv3plus_r50-d8.py',
+ './_base_/datasets/gid.py',
+ '../../../configs/_base_/default_runtime.py',
+ '../../../configs/_base_/schedules/schedule_240k.py'
+]
+custom_imports = dict(
+ imports=['projects.gid_dataset.mmseg.datasets.gid'])
+
+crop_size = (256, 256)
+data_preprocessor = dict(size=crop_size)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ pretrained='open-mmlab://resnet101_v1c',
+ backbone=dict(depth=101),
+ decode_head=dict(num_classes=6),
+ auxiliary_head=dict(num_classes=6))
+
+```
+
+### 3.6 撰写`docs/zh_cn/user_guides/2_dataset_prepare.md`
+
+**Gaofen Image Dataset (GID)**
+
+- GID 数据集可在[此处](https://x-ytong.github.io/project/Five-Billion-Pixels.html)进行下载。
+- GID 数据集包含 150 张 6800x7200 的大尺寸图像,标签为 RGB 标签。
+- 此处选择 15 张图像生成训练集和验证集,该 15 张图像包含了所有六类信息。所选的图像名称如下:
+
+```None
+ GF2_PMS1__L1A0000647767-MSS1
+ GF2_PMS1__L1A0001064454-MSS1
+ GF2_PMS1__L1A0001348919-MSS1
+ GF2_PMS1__L1A0001680851-MSS1
+ GF2_PMS1__L1A0001680853-MSS1
+ GF2_PMS1__L1A0001680857-MSS1
+ GF2_PMS1__L1A0001757429-MSS1
+ GF2_PMS2__L1A0000607681-MSS2
+ GF2_PMS2__L1A0000635115-MSS2
+ GF2_PMS2__L1A0000658637-MSS2
+ GF2_PMS2__L1A0001206072-MSS2
+ GF2_PMS2__L1A0001471436-MSS2
+ GF2_PMS2__L1A0001642620-MSS2
+ GF2_PMS2__L1A0001787089-MSS2
+ GF2_PMS2__L1A0001838560-MSS2
+```
+
+执行以下命令进行裁切及标签的转换,需要修改为您所存储 15 张图像及标签的路径。
+
+```
+python projects/gid_dataset/tools/dataset_converters/gid.py [15 张图像的路径] [15 张标签的路径]
+```
+
+完成裁切后的 GID 数据结构如下:
+
+```none
+mmsegmentation
+├── mmseg
+├── tools
+├── configs
+├── data
+│ ├── gid
+│ │ ├── ann_dir
+| │ │ │ ├── train
+| │ │ │ ├── val
+│ │ ├── img_dir
+| │ │ │ ├── train
+| │ │ │ ├── val
+
+```
+
+### 3.7 贡献的代码及文档通过`pre-commit`检查
+
+使用命令
+
+```bash
+git add .
+git commit -m "添加描述"
+git push
+```
+
+### 3.8 在 GitHub 中向 mmsegmentation 提交 PR
+
+具体步骤可见[《OpenMMLab 贡献代码指南》](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html)
diff --git a/docs/zh_cn/advanced_guides/data_flow.md b/docs/zh_cn/advanced_guides/data_flow.md
index 0716d36d1b4..20dbe07e75d 100644
--- a/docs/zh_cn/advanced_guides/data_flow.md
+++ b/docs/zh_cn/advanced_guides/data_flow.md
@@ -16,7 +16,7 @@ val_cfg = dict(type='ValLoop')
test_cfg = dict(type='TestLoop')
```
-在上图中,红色线表示 [train_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#train_step) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#train_step))*** ,在每次训练迭代中,数据加载器(dataloader)从存储中加载图像并传输到数据预处理器(data preprocessor),数据预处理器会将图像放到特定的设备上,并将数据堆叠到批处理中,之后模型接受批处理数据作为输入,最后将模型的输出发送给优化器(optimizer)。蓝色线表示 [val_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#val_step) 和 [test_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#test_step) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#test_step))*** 。这两个过程的数据流除了模型输出与 `train_step` 不同外,其余均和 `train_step` 类似。由于在评估时模型参数会被冻结,因此模型的输出将被传递给 [Evaluator](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/evaluation.md#ioumetric) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/evaluation.md#ioumetric))***
+在上图中,红色线表示 [train_step](./models.md#train_step),在每次训练迭代中,数据加载器(dataloader)从存储中加载图像并传输到数据预处理器(data preprocessor),数据预处理器会将图像放到特定的设备上,并将数据堆叠到批处理中,之后模型接受批处理数据作为输入,最后将模型的输出发送给优化器(optimizer)。蓝色线表示 [val_step](./models.md#val_step) 和 [test_step](./models.md#test_step)。这两个过程的数据流除了模型输出与 `train_step` 不同外,其余均和 `train_step` 类似。由于在评估时模型参数会被冻结,因此模型的输出将被传递给 [Evaluator](./evaluation.md#ioumetric)。
来计算指标。
## MMSegmentation 中的数据流约定
@@ -28,7 +28,7 @@ test_cfg = dict(type='TestLoop')
数据加载器(DataLoader)是 MMEngine 的训练和测试流程中的一个重要组件。
从概念上讲,它源于 [PyTorch](https://pytorch.org/) 并保持一致。DataLoader 从文件系统加载数据,原始数据通过数据准备流程后被发送给数据预处理器。
-MMSegmentation 在 [PackSegInputs](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/transforms/formatting.py#L12) 中定义了默认数据格式, 它是 `train_pipeline` 和 `test_pipeline` 的最后一个组件。有关数据转换 `pipeline` 的更多信息,请参阅[数据转换文档](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/transforms.html)。 ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/transforms.html))***
+MMSegmentation 在 [PackSegInputs](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/transforms/formatting.py#L12) 中定义了默认数据格式, 它是 `train_pipeline` 和 `test_pipeline` 的最后一个组件。有关数据转换 `pipeline` 的更多信息,请参阅[数据转换文档](./transforms.md)。
在没有任何修改的情况下,PackSegInputs 的返回值通常是一个包含 `inputs` 和 `data_samples` 的 `dict`。以下伪代码展示了 mmseg 中数据加载器输出的数据类型,它是从数据集中获取的一批数据样本,数据加载器将它们打包成一个字典列表。`inputs` 是输入进模型的张量列表,`data_samples` 包含了输入图像的 meta information 和相应的 ground truth。
@@ -39,11 +39,11 @@ dict(
)
```
-**注意:** [SegDataSample](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/structures/seg_data_sample.py) 是 MMSegmentation 的数据结构接口,用于连接不同组件。`SegDataSample` 实现了抽象数据元素 `mmengine.structures.BaseDataElement`,更多信息请在 [MMEngine](https://github.com/open-mmlab/mmengine) 中参阅 [SegDataSample 文档](https://mmsegmentation.readthedocs.io/zh_CN/1.x/advanced_guides/structures.html)和[数据元素文档](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/data_element.html)。
+**注意:** [SegDataSample](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/structures/seg_data_sample.py) 是 MMSegmentation 的数据结构接口,用于连接不同组件。`SegDataSample` 实现了抽象数据元素 `mmengine.structures.BaseDataElement`,更多信息请在 [MMEngine](https://github.com/open-mmlab/mmengine) 中参阅 [SegDataSample 文档](./structures.md)和[数据元素文档](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/data_element.html)。
### 数据预处理器到模型
-虽然在[上面的图](##数据流概述)中分开绘制了数据预处理器和模型,但数据预处理器是模型的一部分,因此可以在[模型教程](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/models.html)中找到数据预处理器章节。 ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/models.html))***
+虽然在[上面的图](##数据流概述)中分开绘制了数据预处理器和模型,但数据预处理器是模型的一部分,因此可以在[模型教程](./models.md)中找到数据预处理器章节。
数据预处理器的返回值是一个包含 `inputs` 和 `data_samples` 的字典,其中 `inputs` 是批处理图像的 4D 张量,`data_samples` 中添加了一些用于数据预处理的额外元信息。当传递给网络时,字典将被解包为两个值。 以下伪代码展示了数据预处理器的返回值和模型的输入值。
@@ -61,22 +61,22 @@ class Network(BaseSegmentor):
pass
```
-**注意:** 模型的前向传播有 3 种模式,由输入参数 mode 控制,更多信息请参阅[模型教程](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md)。 ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md))***
+**注意:** 模型的前向传播有 3 种模式,由输入参数 mode 控制,更多信息请参阅[模型教程](./models.md)。
### 模型输出
-如[模型教程](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#forward) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#forward))*** 所提到的 3 种前向传播具有 3 种输出。
+如[模型教程](./models.md#forward) ***([中文链接待更新](./models.md#forward))*** 所提到的 3 种前向传播具有 3 种输出。
`train_step` 和 `test_step`(或 `val_step`)分别对应于 `'loss'` 和 `'predict'`。
-在 `test_step` 或 `val_step` 中,推理结果会被传递给 `Evaluator` 。您可以参阅[评估文档](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/evaluation.html) ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/evaluation.html))*** 来获取更多关于 `Evaluator` 的信息。
+在 `test_step` 或 `val_step` 中,推理结果会被传递给 `Evaluator` 。您可以参阅[评估文档](./evaluation.md)来获取更多关于 `Evaluator` 的信息。
-在推理后,MMSegmentation 中的 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/segmentors/base.py#L15) 会对推理结果进行简单的后处理以打包推理结果。神经网络生成的分割 logits,经过 `argmax` 操作后的分割 mask 和 ground truth(如果存在)将被打包到类似 `SegDataSample` 的实例。 [postprocess_result](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/segmentors/base.py#L132) 的返回值是一个 **`SegDataSample`的`List`**。下图显示了这些 `SegDataSample` 实例的关键属性。
+在推理后,MMSegmentation 中的 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/segmentors/base.py#L15) 会对推理结果进行简单的后处理以打包推理结果。神经网络生成的分割 logits,经过 `argmax` 操作后的分割 mask 和 ground truth(如果存在)将被打包到类似 `SegDataSample` 的实例。 [postprocess_result](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/segmentors/base.py#L132) 的返回值是一个 **`SegDataSample`的`List`**。下图显示了这些 `SegDataSample` 实例的关键属性。

-与数据预处理器一致,损失函数也是模型的一部分,它是[解码头](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/decode_heads/decode_head.py#L142)的属性之一。
+与数据预处理器一致,损失函数也是模型的一部分,它是[解码头](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/decode_heads/decode_head.py#L142)的属性之一。
-在 MMSegmentation 中,`decode_head` 的 [loss_by_feat](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/decode_heads/decode_head.py#L291) 方法是用于计算损失的统一接口。
+在 MMSegmentation 中,`decode_head` 的 [loss_by_feat](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/decode_heads/decode_head.py#L291) 方法是用于计算损失的统一接口。
参数:
@@ -87,4 +87,4 @@ class Network(BaseSegmentor):
- dict\[str, Tensor\]:一个损失组件的字典
-**注意:** `train_step` 将损失传递进 OptimWrapper 以更新模型中的权重,更多信息请参阅 [train_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#train_step)。 ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#train_step))***
+**注意:** `train_step` 将损失传递进 OptimWrapper 以更新模型中的权重,更多信息请参阅 [train_step](./models.md#train_step)。
diff --git a/docs/zh_cn/advanced_guides/datasets.md b/docs/zh_cn/advanced_guides/datasets.md
index 546e97f70d6..b45f2d22bb6 100644
--- a/docs/zh_cn/advanced_guides/datasets.md
+++ b/docs/zh_cn/advanced_guides/datasets.md
@@ -1,10 +1,10 @@
# 数据集
-在 MMSegmentation 算法库中, 所有 Dataset 类的功能有两个: 加载[预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/2_dataset_prepare.md) 之后的数据集的信息, 和将数据送入[数据集变换流水线](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/basesegdataset.py#L141) 中, 进行[数据变换操作](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/transforms.md). 加载的数据集信息包括两类: 元信息 (meta information), 数据集本身的信息, 例如数据集总共的类别, 和它们对应调色盘信息: 数据信息 (data information) 是指每组数据中图片和对应标签的路径. 下文中介绍了 MMSegmentation 1.x 中数据集的常用接口, 和 mmseg 数据集基类中数据信息加载与修改数据集类别的逻辑, 以及数据集与数据变换流水线 (pipeline) 的关系.
+在 MMSegmentation 算法库中, 所有 Dataset 类的功能有两个: 加载[预处理](../user_guides/2_dataset_prepare.md) 之后的数据集的信息, 和将数据送入[数据集变换流水线](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L141) 中, 进行[数据变换操作](./transforms.md). 加载的数据集信息包括两类: 元信息 (meta information), 数据集本身的信息, 例如数据集总共的类别, 和它们对应调色盘信息: 数据信息 (data information) 是指每组数据中图片和对应标签的路径. 下文中介绍了 MMSegmentation 1.x 中数据集的常用接口, 和 mmseg 数据集基类中数据信息加载与修改数据集类别的逻辑, 以及数据集与数据变换流水线 (pipeline) 的关系.
## 常用接口
-以 Cityscapes 为例, 介绍数据集常用接口. 如需运行以下示例, 请在当前工作目录下的 `data` 目录下载并[预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/2_dataset_prepare.md#cityscapes) Cityscapes 数据集.
+以 Cityscapes 为例, 介绍数据集常用接口. 如需运行以下示例, 请在当前工作目录下的 `data` 目录下载并[预处理](../user_guides/2_dataset_prepare.md#cityscapes) Cityscapes 数据集.
实例化 Cityscapes 训练数据集:
@@ -96,7 +96,7 @@ print(dataset.metainfo)
'reduce_zero_label': False}
```
-数据集 `__getitem__` 方法的返回值, 是经过数据增强的样本数据的输出, 同样也是一个字典, 包括两个字段, `'inputs'` 字段是当前样本经过数据增强操作的图像, 类型为 torch.Tensor, `'data_samples'` 字段存放的数据类型是 MMSegmentation 1.x 新添加的数据结构 [`Segdatasample`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/structures.md), 其中`gt_sem_seg` 字段是经过数据增强的标签数据.
+数据集 `__getitem__` 方法的返回值, 是经过数据增强的样本数据的输出, 同样也是一个字典, 包括两个字段, `'inputs'` 字段是当前样本经过数据增强操作的图像, 类型为 torch.Tensor, `'data_samples'` 字段存放的数据类型是 MMSegmentation 1.x 新添加的数据结构 [`Segdatasample`](./structures.md), 其中`gt_sem_seg` 字段是经过数据增强的标签数据.
```python
print(dataset[0])
@@ -166,13 +166,13 @@ print(dataset[0])
## BaseSegDataset
-由于 MMSegmentation 中的所有数据集的基本功能均包括(1) 加载[数据集预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/user_guides/2_dataset_prepare.md) 之后的数据信息和 (2) 将数据送入数据变换流水线中进行数据变换, 因此在 MMSegmentation 中将其中的共同接口抽象成 [`BaseSegDataset`](https://mmsegmentation.readthedocs.io/en/dev-1.x/api.html?highlight=BaseSegDataset#mmseg.datasets.BaseSegDataset),它继承自 [MMEngine 的 `BaseDataset`](https://github.com/open-mmlab/mmengine/blob/main/docs/en/advanced_tutorials/basedataset.md), 遵循 OpenMMLab 数据集初始化统一流程, 支持高效的内部数据存储格式, 支持数据集拼接、数据集重复采样等功能.
+由于 MMSegmentation 中的所有数据集的基本功能均包括(1) 加载[数据集预处理](../user_guides/2_dataset_prepare.md) 之后的数据信息和 (2) 将数据送入数据变换流水线中进行数据变换, 因此在 MMSegmentation 中将其中的共同接口抽象成 [`BaseSegDataset`](https://mmsegmentation.readthedocs.io/zh_CN/latest/api.html?highlight=BaseSegDataset#mmseg.datasets.BaseSegDataset),它继承自 [MMEngine 的 `BaseDataset`](https://github.com/open-mmlab/mmengine/blob/main/docs/en/advanced_tutorials/basedataset.md), 遵循 OpenMMLab 数据集初始化统一流程, 支持高效的内部数据存储格式, 支持数据集拼接、数据集重复采样等功能.
在 MMSegmentation BaseSegDataset 中重新定义了**数据信息加载方法**(`load_data_list`)和并新增了 `get_label_map` 方法用来**修改数据集的类别信息**.
### 数据信息加载
-数据信息加载的内容是样本数据的图片路径和标签路径, 具体实现在 MMSegmentation 的 BaseSegDataset 的 [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/163277bfe0fa8fefb63ee5137917fafada1b301c/mmseg/datasets/basesegdataset.py#L231) 中.
-主要有两种获取图片和标签的路径方法, 如果当数据集目录按以下目录结构组织, [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/163277bfe0fa8fefb63ee5137917fafada1b301c/mmseg/datasets/basesegdataset.py#L231)) 会根据数据路径和后缀来解析.
+数据信息加载的内容是样本数据的图片路径和标签路径, 具体实现在 MMSegmentation 的 BaseSegDataset 的 [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L231) 中.
+主要有两种获取图片和标签的路径方法, 如果当数据集目录按以下目录结构组织, [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L231)) 会根据数据路径和后缀来解析.
```
├── data
@@ -322,7 +322,7 @@ print(dataset.metainfo)
'reduce_zero_label': False}
```
-可以看到, 数据集元信息的类别和默认 Cityscapes 不同. 并且, 定义了标签重映射的字段 `label_map` 用来修改每个分割掩膜上的像素的类别索引, 分割标签类别会根据 `label_map`, 将类别重映射, [具体实现](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/basesegdataset.py#L151):
+可以看到, 数据集元信息的类别和默认 Cityscapes 不同. 并且, 定义了标签重映射的字段 `label_map` 用来修改每个分割掩膜上的像素的类别索引, 分割标签类别会根据 `label_map`, 将类别重映射, [具体实现](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L151):
```python
gt_semantic_seg_copy = gt_semantic_seg.copy()
diff --git a/docs/zh_cn/advanced_guides/engine.md b/docs/zh_cn/advanced_guides/engine.md
index a5746fcec89..79b4c8d2296 100644
--- a/docs/zh_cn/advanced_guides/engine.md
+++ b/docs/zh_cn/advanced_guides/engine.md
@@ -61,21 +61,21 @@ OpenMMLab 将模型训练和测试过程抽象为 `Runner`, 插入钩子可以
- 默认钩子 (default hooks)
-它们实现了训练时所必需的功能, 在配置文件中用 `default_hooks` 定义传给 `Runner`, `Runner` 通过 [`register_default_hooks`](https://github.com/open-mmlab/mmengine/blob/090104df21acd05a8aadae5a0d743a7da3314f6f/mmengine/runner/runner.py#L1780) 方法注册.
+它们实现了训练时所必需的功能, 在配置文件中用 `default_hooks` 定义传给 `Runner`, `Runner` 通过 [`register_default_hooks`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/runner.py#L1780) 方法注册.
钩子有对应的优先级, 优先级越高, 越早被执行器调用. 如果优先级一样, 被调用的顺序和钩子注册的顺序一致.
不建议用户修改默认钩子的优先级, 可以参考 [mmengine hooks 文档](https://github.com/open-mmlab/mmengine/blob/main/docs/zh_cn/tutorials/hook.md) 了解钩子优先级的定义.
下面是 MMSegmentation 中所用到的默认钩子:
-| 钩子 | 功能 | 优先级 |
-| :-----------------------------------------------------------------------------------------------------------------------: | :---------------------------------------------------------------------------------------------: | :---------------: |
-| [IterTimerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/iter_timer_hook.py) | 记录 iteration 花费的时间. | NORMAL (50) |
-| [LoggerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/logger_hook.py) | 从 `Runner` 里不同的组件中收集日志记录, 并将其输出到终端, JSON 文件, tensorboard, wandb 等下游. | BELOW_NORMAL (60) |
-| [ParamSchedulerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/param_scheduler_hook.py) | 更新优化器里面的一些超参数, 例如学习率的动量. | LOW (70) |
-| [CheckpointHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py) | 规律性地保存 checkpoint 文件. | VERY_LOW (90) |
-| [DistSamplerSeedHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/sampler_seed_hook.py) | 确保分布式采样器 shuffle 是打开的. | NORMAL (50) |
-| [SegVisualizationHook](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/visualization/local_visualizer.py) | 可视化验证和测试过程里的预测结果. | NORMAL (50) |
+| 钩子 | 功能 | 优先级 |
+| :--------------------------------------------------------------------------------------------------------------------: | :---------------------------------------------------------------------------------------------: | :---------------: |
+| [IterTimerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/iter_timer_hook.py) | 记录 iteration 花费的时间. | NORMAL (50) |
+| [LoggerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/logger_hook.py) | 从 `Runner` 里不同的组件中收集日志记录, 并将其输出到终端, JSON 文件, tensorboard, wandb 等下游. | BELOW_NORMAL (60) |
+| [ParamSchedulerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/param_scheduler_hook.py) | 更新优化器里面的一些超参数, 例如学习率的动量. | LOW (70) |
+| [CheckpointHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py) | 规律性地保存 checkpoint 文件. | VERY_LOW (90) |
+| [DistSamplerSeedHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/sampler_seed_hook.py) | 确保分布式采样器 shuffle 是打开的. | NORMAL (50) |
+| [SegVisualizationHook](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/visualization/local_visualizer.py) | 可视化验证和测试过程里的预测结果. | NORMAL (50) |
-MMSegmentation 会在 [`defualt_hooks`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/schedules/schedule_160k.py#L19-L25) 里面注册一些训练所必需功能的钩子::
+MMSegmentation 会在 [`defualt_hooks`](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/schedules/schedule_160k.py#L19-L25) 里面注册一些训练所必需功能的钩子::
```python
default_hooks = dict(
@@ -94,6 +94,7 @@ default_hooks = dict(
以 `default_hooks` 里面的 `logger` 和 `checkpoint` 为例, 我们来介绍如何修改 `default_hooks` 中默认的钩子.
(1) 模型保存配置
+
`default_hooks` 使用 `checkpoint` 字段来初始化[模型保存钩子 (CheckpointHook)](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py#L19).
```python
@@ -104,6 +105,7 @@ checkpoint = dict(type='CheckpointHook', interval=1)
更多相关参数的细节可以参考[这里](https://mmengine.readthedocs.io/zh_CN/latest/api/generated/mmengine.hooks.CheckpointHook.html#checkpointhook).
(2) 日志配置
+
`日志钩子 (LoggerHook)` 被用来收集 `执行器 (Runner)` 里面不同组件的日志信息然后写入终端, JSON 文件, tensorboard 和 wandb 等地方.
```python
@@ -126,7 +128,7 @@ visualizer = dict(
- 自定义钩子 (custom hooks)
-自定义钩子在配置通过 `custom_hooks` 定义, `Runner` 通过 [`register_custom_hooks`](https://github.com/open-mmlab/mmengine/blob/090104df21acd05a8aadae5a0d743a7da3314f6f/mmengine/runner/runner.py#L1852) 方法注册.
+自定义钩子在配置通过 `custom_hooks` 定义, `Runner` 通过 [`register_custom_hooks`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/runner.py#L1820) 方法注册.
自定义钩子优先级需要在配置文件里设置, 如果没有设置, 则会被默认设置为 `NORMAL`. 下面是部分 MMEngine 中实现的自定义钩子:
| 钩子 | 用法 |
@@ -145,7 +147,7 @@ custom_hooks = [
### SegVisualizationHook
-MMSegmentation 实现了 [`SegVisualizationHook`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/engine/hooks/visualization_hook.py#L17), 用来在验证和测试时可视化预测结果.
+MMSegmentation 实现了 [`SegVisualizationHook`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/engine/hooks/visualization_hook.py#L17), 用来在验证和测试时可视化预测结果.
`SegVisualizationHook` 重写了基类 `Hook` 中的 `_after_iter` 方法, 在验证或测试时, 根据指定的迭代次数间隔调用 `visualizer` 的 `add_datasample` 方法绘制语义分割结果, 具体实现如下:
```python
@@ -181,7 +183,7 @@ class SegVisualizationHook(Hook):
```
-关于可视化更多的细节可以查看[这里](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/visualization.md).
+关于可视化更多的细节可以查看[这里](../user_guides/visualization.md).
## 优化器
@@ -234,7 +236,7 @@ optim_wrapper = dict(type='AmpOptimWrapper', optimizer=optimizer)
在模型训练中, 如果想在优化器里为不同参数分别设置优化策略, 例如设置不同的学习率、权重衰减等超参数, 可以通过设置配置文件里 `optim_wrapper` 中的 `paramwise_cfg` 来实现.
-下面的配置文件以 [ViT `optim_wrapper`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/vit/vit_vit-b16-ln_mln_upernet_8xb2-160k_ade20k-512x512.py#L15-L27) 为例介绍 `paramwise_cfg` 参数使用.
+下面的配置文件以 [ViT `optim_wrapper`](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/vit/vit_vit-b16-ln_mln_upernet_8xb2-160k_ade20k-512x512.py#L15-L27) 为例介绍 `paramwise_cfg` 参数使用.
训练时将 `pos_embed`, `mask_token`, `norm` 模块的 weight decay 参数的系数设置成 0.
即: 在训练时, 这些模块的 weight decay 将被变为 `weight_decay * decay_mult`=0.
@@ -257,9 +259,9 @@ optim_wrapper = dict(
### 优化器封装构造器
-默认的优化器封装构造器 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/376251961da47ea8254ab808ae5c51e1430f18dc/mmengine/optim/optimizer/default_constructor.py#L19) 根据输入的 `optim_wrapper` 和 `optim_wrapper` 中定义的 `paramwise_cfg` 来构建训练中使用的优化器. 当 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/376251961da47ea8254ab808ae5c51e1430f18dc/mmengine/optim/optimizer/default_constructor.py#L19) 功能不能满足需求时, 可以自定义优化器封装构造器来实现超参数的配置.
+默认的优化器封装构造器 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/optim/optimizer/default_constructor.py#L19) 根据输入的 `optim_wrapper` 和 `optim_wrapper` 中定义的 `paramwise_cfg` 来构建训练中使用的优化器. 当 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/optim/optimizer/default_constructor.py#L19) 功能不能满足需求时, 可以自定义优化器封装构造器来实现超参数的配置.
-MMSegmentation 中的实现了 [`LearningRateDecayOptimizerConstructor`](https://github.com/open-mmlab/mmsegmentation/blob/b21df463d47447f33c28d9a4f46136ad64d34a40/mmseg/engine/optimizers/layer_decay_optimizer_constructor.py#L104), 可以对以 ConvNeXt, BEiT 和 MAE 为骨干网络的模型训练时, 骨干网络的模型参数的学习率按照定义的衰减比例(`decay_rate`)逐层递减, 在配置文件中的配置如下:
+MMSegmentation 中的实现了 [`LearningRateDecayOptimizerConstructor`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/engine/optimizers/layer_decay_optimizer_constructor.py#L104), 可以对以 ConvNeXt, BEiT 和 MAE 为骨干网络的模型训练时, 骨干网络的模型参数的学习率按照定义的衰减比例(`decay_rate`)逐层递减, 在配置文件中的配置如下:
```python
optim_wrapper = dict(
diff --git a/docs/zh_cn/advanced_guides/models.md b/docs/zh_cn/advanced_guides/models.md
index 408a57863c4..6eb22517a46 100644
--- a/docs/zh_cn/advanced_guides/models.md
+++ b/docs/zh_cn/advanced_guides/models.md
@@ -30,17 +30,17 @@
## 基本接口
-MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmenter](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/models/segmentors/base.py#L15) 类,主要提供 `forward`、`train_step`、`val_step` 和 `test_step` 接口。接下来将详细介绍这些接口。
+MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/models/segmentors/base.py#L15) 类,主要提供 `forward`、`train_step`、`val_step` 和 `test_step` 接口。接下来将详细介绍这些接口。
### forward
+
+### 3.1 在`mmsegmentation/projects`下创建新的项目文件夹
+
+在`mmsegmentation/projects`下创建文件夹`gid_dataset`
+
+
+### 3.2 贡献您的数据集代码
+
+为了最终能将您在 projects 中贡献的代码更加顺畅的移入核心库中(对代码要求质量更高),非常建议按照核心库的目录来编辑您的数据集文件。
+关于数据集有 4 个必要的文件:
+
+- **1** `mmseg/datasets/gid.py` 定义了数据集的尾缀、CLASSES、PALETTE、reduce_zero_label等
+- **2** `configs/_base_/gid.py` GID 数据集的配置文件,定义了数据集的`dataset_type`(数据集类型,`mmseg/datasets/gid.py`中注册的数据集的类名)、`data_root`(数据集所在的根目录,建议将数据集通过软连接的方式将数据集放至`mmsegmentation/data`)、`train_pipline`(训练的数据流)、`test_pipline`(测试和验证时的数据流)、`img_rations`(多尺度预测时的多尺度配置)、`tta_pipeline`(多尺度预测)、`train_dataloader`(训练集的数据加载器)、`val_dataloader`(验证集的数据加载器)、`test_dataloader`(测试集的数据加载器)、`val_evaluator`(验证集的评估器)、`test_evaluator`(测试集的评估器)。
+- **3** 使用了 GID 数据集的模型训练配置文件
+ 这个是可选的,但是强烈建议您添加。在核心库中,所贡献的数据集需要和参考文献中所提出的结果精度对齐,为了后期将您贡献的代码合并入核心库。如您的算力充足,最好能提供对应的模型配置文件在您贡献的数据集上所验证的结果以及相应的权重文件,并撰写较为详细的README.md文档。[示例参考结果](https://github.com/open-mmlab/mmsegmentation/tree/main/configs/deeplabv3plus#mapillary-vistas-v12)
+ 
+- **4** 使用如下命令格式: 撰写`docs/zh_cn/user_guides/2_dataset_prepare.md`来添加您的数据集介绍,包括但不限于数据集的下载方式,数据集目录结构、数据集生成等一些必要性的文字性描述和运行命令。以更好地帮助用户能更快的实现数据集的准备工作。
+
+### 3.3 贡献`tools/dataset_converters/gid.py`
+
+由于 GID 数据集是由未经过切分的 6800x7200 图像所构成的数据集,并且没有划分训练集、验证集与测试集。以及其标签为 RGB 彩色标签,需要将标签转换为单通道的 mask label。为了方便训练,首先将 GID 数据集进行裁切和标签转换,并进行数据集划分,构建为 mmsegmentation 所支持的格式。
+
+```python
+# tools/dataset_converters/gid.py
+import argparse
+import glob
+import math
+import os
+import os.path as osp
+from PIL import Image
+
+import mmcv
+import numpy as np
+from mmengine.utils import ProgressBar, mkdir_or_exist
+
+def parse_args():
+ parser = argparse.ArgumentParser(
+ description='Convert GID dataset to mmsegmentation format')
+ parser.add_argument('dataset_img_path', help='GID images folder path')
+ parser.add_argument('dataset_label_path', help='GID labels folder path')
+ parser.add_argument('--tmp_dir', help='path of the temporary directory')
+ parser.add_argument('-o', '--out_dir', help='output path', default='data/gid')
+ parser.add_argument(
+ '--clip_size',
+ type=int,
+ help='clipped size of image after preparation',
+ default=256)
+ parser.add_argument(
+ '--stride_size',
+ type=int,
+ help='stride of clipping original images',
+ default=256)
+ args = parser.parse_args()
+ return args
+
+GID_COLORMAP = dict(
+ Background=(0, 0, 0), #0-背景-黑色
+ Building=(255, 0, 0), #1-建筑-红色
+ Farmland=(0, 255, 0), #2-农田-绿色
+ Forest=(0, 0, 255), #3-森林-蓝色
+ Meadow=(255, 255, 0),#4-草地-黄色
+ Water=(0, 0, 255)#5-水-蓝色
+)
+palette = list(GID_COLORMAP.values())
+classes = list(GID_COLORMAP.keys())
+
+#############用列表来存一个 RGB 和一个类别的对应################
+def colormap2label(palette):
+ colormap2label_list = np.zeros(256**3, dtype = np.longlong)
+ for i, colormap in enumerate(palette):
+ colormap2label_list[(colormap[0] * 256 + colormap[1])*256+colormap[2]] = i
+ return colormap2label_list
+
+#############给定那个列表,和vis_png然后生成masks_png################
+def label_indices(RGB_label, colormap2label_list):
+ RGB_label = RGB_label.astype('int32')
+ idx = (RGB_label[:, :, 0] * 256 + RGB_label[:, :, 1]) * 256 + RGB_label[:, :, 2]
+ # print(idx.shape)
+ return colormap2label_list[idx]
+
+def RGB2mask(RGB_label, colormap2label_list):
+ # RGB_label = np.array(Image.open(RGB_label).convert('RGB')) #打开RGB_png
+ mask_label = label_indices(RGB_label, colormap2label_list) # .numpy()
+ return mask_label
+
+colormap2label_list = colormap2label(palette)
+
+def clip_big_image(image_path, clip_save_dir, args, to_label=False):
+ """
+ Original image of GID dataset is very large, thus pre-processing
+ of them is adopted. Given fixed clip size and stride size to generate
+ clipped image, the intersection of width and height is determined.
+ For example, given one 6800 x 7200 original image, the clip size is
+ 256 and stride size is 256, thus it would generate 29 x 27 = 783 images
+ whose size are all 256 x 256.
+
+ """
+
+ image = mmcv.imread(image_path, channel_order='rgb')
+ # image = mmcv.bgr2gray(image)
+
+ h, w, c = image.shape
+ clip_size = args.clip_size
+ stride_size = args.stride_size
+
+ num_rows = math.ceil((h - clip_size) / stride_size) if math.ceil(
+ (h - clip_size) /
+ stride_size) * stride_size + clip_size >= h else math.ceil(
+ (h - clip_size) / stride_size) + 1
+ num_cols = math.ceil((w - clip_size) / stride_size) if math.ceil(
+ (w - clip_size) /
+ stride_size) * stride_size + clip_size >= w else math.ceil(
+ (w - clip_size) / stride_size) + 1
+
+ x, y = np.meshgrid(np.arange(num_cols + 1), np.arange(num_rows + 1))
+ xmin = x * clip_size
+ ymin = y * clip_size
+
+ xmin = xmin.ravel()
+ ymin = ymin.ravel()
+ xmin_offset = np.where(xmin + clip_size > w, w - xmin - clip_size,
+ np.zeros_like(xmin))
+ ymin_offset = np.where(ymin + clip_size > h, h - ymin - clip_size,
+ np.zeros_like(ymin))
+ boxes = np.stack([
+ xmin + xmin_offset, ymin + ymin_offset,
+ np.minimum(xmin + clip_size, w),
+ np.minimum(ymin + clip_size, h)
+ ], axis=1)
+
+ if to_label:
+ image = RGB2mask(image, colormap2label_list) #这里得改一下
+
+ for count, box in enumerate(boxes):
+ start_x, start_y, end_x, end_y = box
+ clipped_image = image[start_y:end_y,
+ start_x:end_x] if to_label else image[
+ start_y:end_y, start_x:end_x, :]
+ img_name = osp.basename(image_path).replace('.tif', '')
+ img_name = img_name.replace('_label', '')
+ if count % 3 == 0:
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(
+ clip_save_dir.replace('train', 'val'),
+ f'{img_name}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+ else:
+ mmcv.imwrite(
+ clipped_image.astype(np.uint8),
+ osp.join(
+ clip_save_dir,
+ f'{img_name}_{start_x}_{start_y}_{end_x}_{end_y}.png'))
+ count += 1
+
+def main():
+ args = parse_args()
+
+ """
+ According to this paper: https://ieeexplore.ieee.org/document/9343296/
+ select 15 images contained in GID, , which cover the whole six
+ categories, to generate train set and validation set.
+
+ According to Paper: https://ieeexplore.ieee.org/document/9343296/
+
+ """
+
+ if args.out_dir is None:
+ out_dir = osp.join('data', 'gid')
+ else:
+ out_dir = args.out_dir
+
+ print('Making directories...')
+ mkdir_or_exist(osp.join(out_dir, 'img_dir', 'train'))
+ mkdir_or_exist(osp.join(out_dir, 'img_dir', 'val'))
+ mkdir_or_exist(osp.join(out_dir, 'ann_dir', 'train'))
+ mkdir_or_exist(osp.join(out_dir, 'ann_dir', 'val'))
+
+ src_path_list = glob.glob(os.path.join(args.dataset_img_path, '*.tif'))
+ print(f'Find {len(src_path_list)} pictures')
+
+ prog_bar = ProgressBar(len(src_path_list))
+
+ dst_img_dir = osp.join(out_dir, 'img_dir', 'train')
+ dst_label_dir = osp.join(out_dir, 'ann_dir', 'train')
+
+ for i, img_path in enumerate(src_path_list):
+ label_path = osp.join(args.dataset_label_path, osp.basename(img_path.replace('.tif', '_label.tif')))
+
+ clip_big_image(img_path, dst_img_dir, args, to_label=False)
+ clip_big_image(label_path, dst_label_dir, args, to_label=True)
+ prog_bar.update()
+
+ print('Done!')
+
+if __name__ == '__main__':
+ main()
+```
+
+### 3.4 贡献`mmseg/datasets/gid.py`
+
+可参考[`projects/mapillary_dataset/mmseg/datasets/mapillary.py`](https://github.com/open-mmlab/mmsegmentation/blob/main/projects/mapillary_dataset/mmseg/datasets/mapillary.py)并在此基础上修改相应变量以适配您的数据集。
+
+```python
+# mmseg/datasets/gid.py
+# Copyright (c) OpenMMLab. All rights reserved.
+from mmseg.datasets.basesegdataset import BaseSegDataset
+from mmseg.registry import DATASETS
+
+# 注册数据集类
+@DATASETS.register_module()
+class GID_Dataset(BaseSegDataset):
+ """Gaofen Image Dataset (GID)
+
+ Dataset paper link:
+ https://www.sciencedirect.com/science/article/pii/S0034425719303414
+ https://x-ytong.github.io/project/GID.html
+
+ GID 6 classes: background(others), built-up, farmland, forest, meadow, water
+
+ In This example, select 10 images from GID dataset as training set,
+ and select 5 images as validation set.
+ The selected images are listed as follows:
+
+ GF2_PMS1__L1A0000647767-MSS1
+ GF2_PMS1__L1A0001064454-MSS1
+ GF2_PMS1__L1A0001348919-MSS1
+ GF2_PMS1__L1A0001680851-MSS1
+ GF2_PMS1__L1A0001680853-MSS1
+ GF2_PMS1__L1A0001680857-MSS1
+ GF2_PMS1__L1A0001757429-MSS1
+ GF2_PMS2__L1A0000607681-MSS2
+ GF2_PMS2__L1A0000635115-MSS2
+ GF2_PMS2__L1A0000658637-MSS2
+ GF2_PMS2__L1A0001206072-MSS2
+ GF2_PMS2__L1A0001471436-MSS2
+ GF2_PMS2__L1A0001642620-MSS2
+ GF2_PMS2__L1A0001787089-MSS2
+ GF2_PMS2__L1A0001838560-MSS2
+
+ The ``img_suffix`` is fixed to '.tif' and ``seg_map_suffix`` is
+ fixed to '.tif' for GID.
+ """
+ METAINFO = dict(
+ classes=('Others', 'Built-up', 'Farmland', 'Forest',
+ 'Meadow', 'Water'),
+
+ palette=[[0, 0, 0], [255, 0, 0], [0, 255, 0], [0, 255, 255],
+ [255, 255, 0], [0, 0, 255]])
+
+ def __init__(self,
+ img_suffix='.png',
+ seg_map_suffix='.png',
+ reduce_zero_label=None,
+ **kwargs) -> None:
+ super().__init__(
+ img_suffix=img_suffix,
+ seg_map_suffix=seg_map_suffix,
+ reduce_zero_label=reduce_zero_label,
+ **kwargs)
+```
+
+### 3.5 贡献使用 GID 的训练 config file
+
+```python
+_base_ = [
+ '../../../configs/_base_/models/deeplabv3plus_r50-d8.py',
+ './_base_/datasets/gid.py',
+ '../../../configs/_base_/default_runtime.py',
+ '../../../configs/_base_/schedules/schedule_240k.py'
+]
+custom_imports = dict(
+ imports=['projects.gid_dataset.mmseg.datasets.gid'])
+
+crop_size = (256, 256)
+data_preprocessor = dict(size=crop_size)
+model = dict(
+ data_preprocessor=data_preprocessor,
+ pretrained='open-mmlab://resnet101_v1c',
+ backbone=dict(depth=101),
+ decode_head=dict(num_classes=6),
+ auxiliary_head=dict(num_classes=6))
+
+```
+
+### 3.6 撰写`docs/zh_cn/user_guides/2_dataset_prepare.md`
+
+**Gaofen Image Dataset (GID)**
+
+- GID 数据集可在[此处](https://x-ytong.github.io/project/Five-Billion-Pixels.html)进行下载。
+- GID 数据集包含 150 张 6800x7200 的大尺寸图像,标签为 RGB 标签。
+- 此处选择 15 张图像生成训练集和验证集,该 15 张图像包含了所有六类信息。所选的图像名称如下:
+
+```None
+ GF2_PMS1__L1A0000647767-MSS1
+ GF2_PMS1__L1A0001064454-MSS1
+ GF2_PMS1__L1A0001348919-MSS1
+ GF2_PMS1__L1A0001680851-MSS1
+ GF2_PMS1__L1A0001680853-MSS1
+ GF2_PMS1__L1A0001680857-MSS1
+ GF2_PMS1__L1A0001757429-MSS1
+ GF2_PMS2__L1A0000607681-MSS2
+ GF2_PMS2__L1A0000635115-MSS2
+ GF2_PMS2__L1A0000658637-MSS2
+ GF2_PMS2__L1A0001206072-MSS2
+ GF2_PMS2__L1A0001471436-MSS2
+ GF2_PMS2__L1A0001642620-MSS2
+ GF2_PMS2__L1A0001787089-MSS2
+ GF2_PMS2__L1A0001838560-MSS2
+```
+
+执行以下命令进行裁切及标签的转换,需要修改为您所存储 15 张图像及标签的路径。
+
+```
+python projects/gid_dataset/tools/dataset_converters/gid.py [15 张图像的路径] [15 张标签的路径]
+```
+
+完成裁切后的 GID 数据结构如下:
+
+```none
+mmsegmentation
+├── mmseg
+├── tools
+├── configs
+├── data
+│ ├── gid
+│ │ ├── ann_dir
+| │ │ │ ├── train
+| │ │ │ ├── val
+│ │ ├── img_dir
+| │ │ │ ├── train
+| │ │ │ ├── val
+
+```
+
+### 3.7 贡献的代码及文档通过`pre-commit`检查
+
+使用命令
+
+```bash
+git add .
+git commit -m "添加描述"
+git push
+```
+
+### 3.8 在 GitHub 中向 mmsegmentation 提交 PR
+
+具体步骤可见[《OpenMMLab 贡献代码指南》](https://mmcv.readthedocs.io/zh_CN/latest/community/contributing.html)
diff --git a/docs/zh_cn/advanced_guides/data_flow.md b/docs/zh_cn/advanced_guides/data_flow.md
index 0716d36d1b4..20dbe07e75d 100644
--- a/docs/zh_cn/advanced_guides/data_flow.md
+++ b/docs/zh_cn/advanced_guides/data_flow.md
@@ -16,7 +16,7 @@ val_cfg = dict(type='ValLoop')
test_cfg = dict(type='TestLoop')
```
-在上图中,红色线表示 [train_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#train_step) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#train_step))*** ,在每次训练迭代中,数据加载器(dataloader)从存储中加载图像并传输到数据预处理器(data preprocessor),数据预处理器会将图像放到特定的设备上,并将数据堆叠到批处理中,之后模型接受批处理数据作为输入,最后将模型的输出发送给优化器(optimizer)。蓝色线表示 [val_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#val_step) 和 [test_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#test_step) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#test_step))*** 。这两个过程的数据流除了模型输出与 `train_step` 不同外,其余均和 `train_step` 类似。由于在评估时模型参数会被冻结,因此模型的输出将被传递给 [Evaluator](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/evaluation.md#ioumetric) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/evaluation.md#ioumetric))***
+在上图中,红色线表示 [train_step](./models.md#train_step),在每次训练迭代中,数据加载器(dataloader)从存储中加载图像并传输到数据预处理器(data preprocessor),数据预处理器会将图像放到特定的设备上,并将数据堆叠到批处理中,之后模型接受批处理数据作为输入,最后将模型的输出发送给优化器(optimizer)。蓝色线表示 [val_step](./models.md#val_step) 和 [test_step](./models.md#test_step)。这两个过程的数据流除了模型输出与 `train_step` 不同外,其余均和 `train_step` 类似。由于在评估时模型参数会被冻结,因此模型的输出将被传递给 [Evaluator](./evaluation.md#ioumetric)。
来计算指标。
## MMSegmentation 中的数据流约定
@@ -28,7 +28,7 @@ test_cfg = dict(type='TestLoop')
数据加载器(DataLoader)是 MMEngine 的训练和测试流程中的一个重要组件。
从概念上讲,它源于 [PyTorch](https://pytorch.org/) 并保持一致。DataLoader 从文件系统加载数据,原始数据通过数据准备流程后被发送给数据预处理器。
-MMSegmentation 在 [PackSegInputs](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/transforms/formatting.py#L12) 中定义了默认数据格式, 它是 `train_pipeline` 和 `test_pipeline` 的最后一个组件。有关数据转换 `pipeline` 的更多信息,请参阅[数据转换文档](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/transforms.html)。 ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/transforms.html))***
+MMSegmentation 在 [PackSegInputs](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/transforms/formatting.py#L12) 中定义了默认数据格式, 它是 `train_pipeline` 和 `test_pipeline` 的最后一个组件。有关数据转换 `pipeline` 的更多信息,请参阅[数据转换文档](./transforms.md)。
在没有任何修改的情况下,PackSegInputs 的返回值通常是一个包含 `inputs` 和 `data_samples` 的 `dict`。以下伪代码展示了 mmseg 中数据加载器输出的数据类型,它是从数据集中获取的一批数据样本,数据加载器将它们打包成一个字典列表。`inputs` 是输入进模型的张量列表,`data_samples` 包含了输入图像的 meta information 和相应的 ground truth。
@@ -39,11 +39,11 @@ dict(
)
```
-**注意:** [SegDataSample](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/structures/seg_data_sample.py) 是 MMSegmentation 的数据结构接口,用于连接不同组件。`SegDataSample` 实现了抽象数据元素 `mmengine.structures.BaseDataElement`,更多信息请在 [MMEngine](https://github.com/open-mmlab/mmengine) 中参阅 [SegDataSample 文档](https://mmsegmentation.readthedocs.io/zh_CN/1.x/advanced_guides/structures.html)和[数据元素文档](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/data_element.html)。
+**注意:** [SegDataSample](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/structures/seg_data_sample.py) 是 MMSegmentation 的数据结构接口,用于连接不同组件。`SegDataSample` 实现了抽象数据元素 `mmengine.structures.BaseDataElement`,更多信息请在 [MMEngine](https://github.com/open-mmlab/mmengine) 中参阅 [SegDataSample 文档](./structures.md)和[数据元素文档](https://mmengine.readthedocs.io/zh_CN/latest/advanced_tutorials/data_element.html)。
### 数据预处理器到模型
-虽然在[上面的图](##数据流概述)中分开绘制了数据预处理器和模型,但数据预处理器是模型的一部分,因此可以在[模型教程](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/models.html)中找到数据预处理器章节。 ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/models.html))***
+虽然在[上面的图](##数据流概述)中分开绘制了数据预处理器和模型,但数据预处理器是模型的一部分,因此可以在[模型教程](./models.md)中找到数据预处理器章节。
数据预处理器的返回值是一个包含 `inputs` 和 `data_samples` 的字典,其中 `inputs` 是批处理图像的 4D 张量,`data_samples` 中添加了一些用于数据预处理的额外元信息。当传递给网络时,字典将被解包为两个值。 以下伪代码展示了数据预处理器的返回值和模型的输入值。
@@ -61,22 +61,22 @@ class Network(BaseSegmentor):
pass
```
-**注意:** 模型的前向传播有 3 种模式,由输入参数 mode 控制,更多信息请参阅[模型教程](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md)。 ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md))***
+**注意:** 模型的前向传播有 3 种模式,由输入参数 mode 控制,更多信息请参阅[模型教程](./models.md)。
### 模型输出
-如[模型教程](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#forward) ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#forward))*** 所提到的 3 种前向传播具有 3 种输出。
+如[模型教程](./models.md#forward) ***([中文链接待更新](./models.md#forward))*** 所提到的 3 种前向传播具有 3 种输出。
`train_step` 和 `test_step`(或 `val_step`)分别对应于 `'loss'` 和 `'predict'`。
-在 `test_step` 或 `val_step` 中,推理结果会被传递给 `Evaluator` 。您可以参阅[评估文档](https://mmsegmentation.readthedocs.io/en/dev-1.x/advanced_guides/evaluation.html) ***([中文链接待更新](https://mmsegmentation.readthedocs.io/zh_CN/dev-1.x/advanced_guides/evaluation.html))*** 来获取更多关于 `Evaluator` 的信息。
+在 `test_step` 或 `val_step` 中,推理结果会被传递给 `Evaluator` 。您可以参阅[评估文档](./evaluation.md)来获取更多关于 `Evaluator` 的信息。
-在推理后,MMSegmentation 中的 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/segmentors/base.py#L15) 会对推理结果进行简单的后处理以打包推理结果。神经网络生成的分割 logits,经过 `argmax` 操作后的分割 mask 和 ground truth(如果存在)将被打包到类似 `SegDataSample` 的实例。 [postprocess_result](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/segmentors/base.py#L132) 的返回值是一个 **`SegDataSample`的`List`**。下图显示了这些 `SegDataSample` 实例的关键属性。
+在推理后,MMSegmentation 中的 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/segmentors/base.py#L15) 会对推理结果进行简单的后处理以打包推理结果。神经网络生成的分割 logits,经过 `argmax` 操作后的分割 mask 和 ground truth(如果存在)将被打包到类似 `SegDataSample` 的实例。 [postprocess_result](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/segmentors/base.py#L132) 的返回值是一个 **`SegDataSample`的`List`**。下图显示了这些 `SegDataSample` 实例的关键属性。

-与数据预处理器一致,损失函数也是模型的一部分,它是[解码头](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/decode_heads/decode_head.py#L142)的属性之一。
+与数据预处理器一致,损失函数也是模型的一部分,它是[解码头](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/decode_heads/decode_head.py#L142)的属性之一。
-在 MMSegmentation 中,`decode_head` 的 [loss_by_feat](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/models/decode_heads/decode_head.py#L291) 方法是用于计算损失的统一接口。
+在 MMSegmentation 中,`decode_head` 的 [loss_by_feat](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/models/decode_heads/decode_head.py#L291) 方法是用于计算损失的统一接口。
参数:
@@ -87,4 +87,4 @@ class Network(BaseSegmentor):
- dict\[str, Tensor\]:一个损失组件的字典
-**注意:** `train_step` 将损失传递进 OptimWrapper 以更新模型中的权重,更多信息请参阅 [train_step](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/advanced_guides/models.md#train_step)。 ***([中文链接待更新](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/models.md#train_step))***
+**注意:** `train_step` 将损失传递进 OptimWrapper 以更新模型中的权重,更多信息请参阅 [train_step](./models.md#train_step)。
diff --git a/docs/zh_cn/advanced_guides/datasets.md b/docs/zh_cn/advanced_guides/datasets.md
index 546e97f70d6..b45f2d22bb6 100644
--- a/docs/zh_cn/advanced_guides/datasets.md
+++ b/docs/zh_cn/advanced_guides/datasets.md
@@ -1,10 +1,10 @@
# 数据集
-在 MMSegmentation 算法库中, 所有 Dataset 类的功能有两个: 加载[预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/2_dataset_prepare.md) 之后的数据集的信息, 和将数据送入[数据集变换流水线](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/basesegdataset.py#L141) 中, 进行[数据变换操作](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/transforms.md). 加载的数据集信息包括两类: 元信息 (meta information), 数据集本身的信息, 例如数据集总共的类别, 和它们对应调色盘信息: 数据信息 (data information) 是指每组数据中图片和对应标签的路径. 下文中介绍了 MMSegmentation 1.x 中数据集的常用接口, 和 mmseg 数据集基类中数据信息加载与修改数据集类别的逻辑, 以及数据集与数据变换流水线 (pipeline) 的关系.
+在 MMSegmentation 算法库中, 所有 Dataset 类的功能有两个: 加载[预处理](../user_guides/2_dataset_prepare.md) 之后的数据集的信息, 和将数据送入[数据集变换流水线](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L141) 中, 进行[数据变换操作](./transforms.md). 加载的数据集信息包括两类: 元信息 (meta information), 数据集本身的信息, 例如数据集总共的类别, 和它们对应调色盘信息: 数据信息 (data information) 是指每组数据中图片和对应标签的路径. 下文中介绍了 MMSegmentation 1.x 中数据集的常用接口, 和 mmseg 数据集基类中数据信息加载与修改数据集类别的逻辑, 以及数据集与数据变换流水线 (pipeline) 的关系.
## 常用接口
-以 Cityscapes 为例, 介绍数据集常用接口. 如需运行以下示例, 请在当前工作目录下的 `data` 目录下载并[预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/2_dataset_prepare.md#cityscapes) Cityscapes 数据集.
+以 Cityscapes 为例, 介绍数据集常用接口. 如需运行以下示例, 请在当前工作目录下的 `data` 目录下载并[预处理](../user_guides/2_dataset_prepare.md#cityscapes) Cityscapes 数据集.
实例化 Cityscapes 训练数据集:
@@ -96,7 +96,7 @@ print(dataset.metainfo)
'reduce_zero_label': False}
```
-数据集 `__getitem__` 方法的返回值, 是经过数据增强的样本数据的输出, 同样也是一个字典, 包括两个字段, `'inputs'` 字段是当前样本经过数据增强操作的图像, 类型为 torch.Tensor, `'data_samples'` 字段存放的数据类型是 MMSegmentation 1.x 新添加的数据结构 [`Segdatasample`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/advanced_guides/structures.md), 其中`gt_sem_seg` 字段是经过数据增强的标签数据.
+数据集 `__getitem__` 方法的返回值, 是经过数据增强的样本数据的输出, 同样也是一个字典, 包括两个字段, `'inputs'` 字段是当前样本经过数据增强操作的图像, 类型为 torch.Tensor, `'data_samples'` 字段存放的数据类型是 MMSegmentation 1.x 新添加的数据结构 [`Segdatasample`](./structures.md), 其中`gt_sem_seg` 字段是经过数据增强的标签数据.
```python
print(dataset[0])
@@ -166,13 +166,13 @@ print(dataset[0])
## BaseSegDataset
-由于 MMSegmentation 中的所有数据集的基本功能均包括(1) 加载[数据集预处理](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/zh_cn/user_guides/2_dataset_prepare.md) 之后的数据信息和 (2) 将数据送入数据变换流水线中进行数据变换, 因此在 MMSegmentation 中将其中的共同接口抽象成 [`BaseSegDataset`](https://mmsegmentation.readthedocs.io/en/dev-1.x/api.html?highlight=BaseSegDataset#mmseg.datasets.BaseSegDataset),它继承自 [MMEngine 的 `BaseDataset`](https://github.com/open-mmlab/mmengine/blob/main/docs/en/advanced_tutorials/basedataset.md), 遵循 OpenMMLab 数据集初始化统一流程, 支持高效的内部数据存储格式, 支持数据集拼接、数据集重复采样等功能.
+由于 MMSegmentation 中的所有数据集的基本功能均包括(1) 加载[数据集预处理](../user_guides/2_dataset_prepare.md) 之后的数据信息和 (2) 将数据送入数据变换流水线中进行数据变换, 因此在 MMSegmentation 中将其中的共同接口抽象成 [`BaseSegDataset`](https://mmsegmentation.readthedocs.io/zh_CN/latest/api.html?highlight=BaseSegDataset#mmseg.datasets.BaseSegDataset),它继承自 [MMEngine 的 `BaseDataset`](https://github.com/open-mmlab/mmengine/blob/main/docs/en/advanced_tutorials/basedataset.md), 遵循 OpenMMLab 数据集初始化统一流程, 支持高效的内部数据存储格式, 支持数据集拼接、数据集重复采样等功能.
在 MMSegmentation BaseSegDataset 中重新定义了**数据信息加载方法**(`load_data_list`)和并新增了 `get_label_map` 方法用来**修改数据集的类别信息**.
### 数据信息加载
-数据信息加载的内容是样本数据的图片路径和标签路径, 具体实现在 MMSegmentation 的 BaseSegDataset 的 [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/163277bfe0fa8fefb63ee5137917fafada1b301c/mmseg/datasets/basesegdataset.py#L231) 中.
-主要有两种获取图片和标签的路径方法, 如果当数据集目录按以下目录结构组织, [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/163277bfe0fa8fefb63ee5137917fafada1b301c/mmseg/datasets/basesegdataset.py#L231)) 会根据数据路径和后缀来解析.
+数据信息加载的内容是样本数据的图片路径和标签路径, 具体实现在 MMSegmentation 的 BaseSegDataset 的 [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L231) 中.
+主要有两种获取图片和标签的路径方法, 如果当数据集目录按以下目录结构组织, [`load_data_list`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L231)) 会根据数据路径和后缀来解析.
```
├── data
@@ -322,7 +322,7 @@ print(dataset.metainfo)
'reduce_zero_label': False}
```
-可以看到, 数据集元信息的类别和默认 Cityscapes 不同. 并且, 定义了标签重映射的字段 `label_map` 用来修改每个分割掩膜上的像素的类别索引, 分割标签类别会根据 `label_map`, 将类别重映射, [具体实现](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/datasets/basesegdataset.py#L151):
+可以看到, 数据集元信息的类别和默认 Cityscapes 不同. 并且, 定义了标签重映射的字段 `label_map` 用来修改每个分割掩膜上的像素的类别索引, 分割标签类别会根据 `label_map`, 将类别重映射, [具体实现](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/datasets/basesegdataset.py#L151):
```python
gt_semantic_seg_copy = gt_semantic_seg.copy()
diff --git a/docs/zh_cn/advanced_guides/engine.md b/docs/zh_cn/advanced_guides/engine.md
index a5746fcec89..79b4c8d2296 100644
--- a/docs/zh_cn/advanced_guides/engine.md
+++ b/docs/zh_cn/advanced_guides/engine.md
@@ -61,21 +61,21 @@ OpenMMLab 将模型训练和测试过程抽象为 `Runner`, 插入钩子可以
- 默认钩子 (default hooks)
-它们实现了训练时所必需的功能, 在配置文件中用 `default_hooks` 定义传给 `Runner`, `Runner` 通过 [`register_default_hooks`](https://github.com/open-mmlab/mmengine/blob/090104df21acd05a8aadae5a0d743a7da3314f6f/mmengine/runner/runner.py#L1780) 方法注册.
+它们实现了训练时所必需的功能, 在配置文件中用 `default_hooks` 定义传给 `Runner`, `Runner` 通过 [`register_default_hooks`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/runner.py#L1780) 方法注册.
钩子有对应的优先级, 优先级越高, 越早被执行器调用. 如果优先级一样, 被调用的顺序和钩子注册的顺序一致.
不建议用户修改默认钩子的优先级, 可以参考 [mmengine hooks 文档](https://github.com/open-mmlab/mmengine/blob/main/docs/zh_cn/tutorials/hook.md) 了解钩子优先级的定义.
下面是 MMSegmentation 中所用到的默认钩子:
-| 钩子 | 功能 | 优先级 |
-| :-----------------------------------------------------------------------------------------------------------------------: | :---------------------------------------------------------------------------------------------: | :---------------: |
-| [IterTimerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/iter_timer_hook.py) | 记录 iteration 花费的时间. | NORMAL (50) |
-| [LoggerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/logger_hook.py) | 从 `Runner` 里不同的组件中收集日志记录, 并将其输出到终端, JSON 文件, tensorboard, wandb 等下游. | BELOW_NORMAL (60) |
-| [ParamSchedulerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/param_scheduler_hook.py) | 更新优化器里面的一些超参数, 例如学习率的动量. | LOW (70) |
-| [CheckpointHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py) | 规律性地保存 checkpoint 文件. | VERY_LOW (90) |
-| [DistSamplerSeedHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/sampler_seed_hook.py) | 确保分布式采样器 shuffle 是打开的. | NORMAL (50) |
-| [SegVisualizationHook](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/visualization/local_visualizer.py) | 可视化验证和测试过程里的预测结果. | NORMAL (50) |
+| 钩子 | 功能 | 优先级 |
+| :--------------------------------------------------------------------------------------------------------------------: | :---------------------------------------------------------------------------------------------: | :---------------: |
+| [IterTimerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/iter_timer_hook.py) | 记录 iteration 花费的时间. | NORMAL (50) |
+| [LoggerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/logger_hook.py) | 从 `Runner` 里不同的组件中收集日志记录, 并将其输出到终端, JSON 文件, tensorboard, wandb 等下游. | BELOW_NORMAL (60) |
+| [ParamSchedulerHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/param_scheduler_hook.py) | 更新优化器里面的一些超参数, 例如学习率的动量. | LOW (70) |
+| [CheckpointHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py) | 规律性地保存 checkpoint 文件. | VERY_LOW (90) |
+| [DistSamplerSeedHook](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/sampler_seed_hook.py) | 确保分布式采样器 shuffle 是打开的. | NORMAL (50) |
+| [SegVisualizationHook](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/visualization/local_visualizer.py) | 可视化验证和测试过程里的预测结果. | NORMAL (50) |
-MMSegmentation 会在 [`defualt_hooks`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/_base_/schedules/schedule_160k.py#L19-L25) 里面注册一些训练所必需功能的钩子::
+MMSegmentation 会在 [`defualt_hooks`](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/_base_/schedules/schedule_160k.py#L19-L25) 里面注册一些训练所必需功能的钩子::
```python
default_hooks = dict(
@@ -94,6 +94,7 @@ default_hooks = dict(
以 `default_hooks` 里面的 `logger` 和 `checkpoint` 为例, 我们来介绍如何修改 `default_hooks` 中默认的钩子.
(1) 模型保存配置
+
`default_hooks` 使用 `checkpoint` 字段来初始化[模型保存钩子 (CheckpointHook)](https://github.com/open-mmlab/mmengine/blob/main/mmengine/hooks/checkpoint_hook.py#L19).
```python
@@ -104,6 +105,7 @@ checkpoint = dict(type='CheckpointHook', interval=1)
更多相关参数的细节可以参考[这里](https://mmengine.readthedocs.io/zh_CN/latest/api/generated/mmengine.hooks.CheckpointHook.html#checkpointhook).
(2) 日志配置
+
`日志钩子 (LoggerHook)` 被用来收集 `执行器 (Runner)` 里面不同组件的日志信息然后写入终端, JSON 文件, tensorboard 和 wandb 等地方.
```python
@@ -126,7 +128,7 @@ visualizer = dict(
- 自定义钩子 (custom hooks)
-自定义钩子在配置通过 `custom_hooks` 定义, `Runner` 通过 [`register_custom_hooks`](https://github.com/open-mmlab/mmengine/blob/090104df21acd05a8aadae5a0d743a7da3314f6f/mmengine/runner/runner.py#L1852) 方法注册.
+自定义钩子在配置通过 `custom_hooks` 定义, `Runner` 通过 [`register_custom_hooks`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/runner/runner.py#L1820) 方法注册.
自定义钩子优先级需要在配置文件里设置, 如果没有设置, 则会被默认设置为 `NORMAL`. 下面是部分 MMEngine 中实现的自定义钩子:
| 钩子 | 用法 |
@@ -145,7 +147,7 @@ custom_hooks = [
### SegVisualizationHook
-MMSegmentation 实现了 [`SegVisualizationHook`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/mmseg/engine/hooks/visualization_hook.py#L17), 用来在验证和测试时可视化预测结果.
+MMSegmentation 实现了 [`SegVisualizationHook`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/engine/hooks/visualization_hook.py#L17), 用来在验证和测试时可视化预测结果.
`SegVisualizationHook` 重写了基类 `Hook` 中的 `_after_iter` 方法, 在验证或测试时, 根据指定的迭代次数间隔调用 `visualizer` 的 `add_datasample` 方法绘制语义分割结果, 具体实现如下:
```python
@@ -181,7 +183,7 @@ class SegVisualizationHook(Hook):
```
-关于可视化更多的细节可以查看[这里](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/docs/en/user_guides/visualization.md).
+关于可视化更多的细节可以查看[这里](../user_guides/visualization.md).
## 优化器
@@ -234,7 +236,7 @@ optim_wrapper = dict(type='AmpOptimWrapper', optimizer=optimizer)
在模型训练中, 如果想在优化器里为不同参数分别设置优化策略, 例如设置不同的学习率、权重衰减等超参数, 可以通过设置配置文件里 `optim_wrapper` 中的 `paramwise_cfg` 来实现.
-下面的配置文件以 [ViT `optim_wrapper`](https://github.com/open-mmlab/mmsegmentation/blob/dev-1.x/configs/vit/vit_vit-b16-ln_mln_upernet_8xb2-160k_ade20k-512x512.py#L15-L27) 为例介绍 `paramwise_cfg` 参数使用.
+下面的配置文件以 [ViT `optim_wrapper`](https://github.com/open-mmlab/mmsegmentation/blob/main/configs/vit/vit_vit-b16-ln_mln_upernet_8xb2-160k_ade20k-512x512.py#L15-L27) 为例介绍 `paramwise_cfg` 参数使用.
训练时将 `pos_embed`, `mask_token`, `norm` 模块的 weight decay 参数的系数设置成 0.
即: 在训练时, 这些模块的 weight decay 将被变为 `weight_decay * decay_mult`=0.
@@ -257,9 +259,9 @@ optim_wrapper = dict(
### 优化器封装构造器
-默认的优化器封装构造器 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/376251961da47ea8254ab808ae5c51e1430f18dc/mmengine/optim/optimizer/default_constructor.py#L19) 根据输入的 `optim_wrapper` 和 `optim_wrapper` 中定义的 `paramwise_cfg` 来构建训练中使用的优化器. 当 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/376251961da47ea8254ab808ae5c51e1430f18dc/mmengine/optim/optimizer/default_constructor.py#L19) 功能不能满足需求时, 可以自定义优化器封装构造器来实现超参数的配置.
+默认的优化器封装构造器 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/optim/optimizer/default_constructor.py#L19) 根据输入的 `optim_wrapper` 和 `optim_wrapper` 中定义的 `paramwise_cfg` 来构建训练中使用的优化器. 当 [`DefaultOptimWrapperConstructor`](https://github.com/open-mmlab/mmengine/blob/main/mmengine/optim/optimizer/default_constructor.py#L19) 功能不能满足需求时, 可以自定义优化器封装构造器来实现超参数的配置.
-MMSegmentation 中的实现了 [`LearningRateDecayOptimizerConstructor`](https://github.com/open-mmlab/mmsegmentation/blob/b21df463d47447f33c28d9a4f46136ad64d34a40/mmseg/engine/optimizers/layer_decay_optimizer_constructor.py#L104), 可以对以 ConvNeXt, BEiT 和 MAE 为骨干网络的模型训练时, 骨干网络的模型参数的学习率按照定义的衰减比例(`decay_rate`)逐层递减, 在配置文件中的配置如下:
+MMSegmentation 中的实现了 [`LearningRateDecayOptimizerConstructor`](https://github.com/open-mmlab/mmsegmentation/blob/main/mmseg/engine/optimizers/layer_decay_optimizer_constructor.py#L104), 可以对以 ConvNeXt, BEiT 和 MAE 为骨干网络的模型训练时, 骨干网络的模型参数的学习率按照定义的衰减比例(`decay_rate`)逐层递减, 在配置文件中的配置如下:
```python
optim_wrapper = dict(
diff --git a/docs/zh_cn/advanced_guides/models.md b/docs/zh_cn/advanced_guides/models.md
index 408a57863c4..6eb22517a46 100644
--- a/docs/zh_cn/advanced_guides/models.md
+++ b/docs/zh_cn/advanced_guides/models.md
@@ -30,17 +30,17 @@
## 基本接口
-MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmenter](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/models/segmentors/base.py#L15) 类,主要提供 `forward`、`train_step`、`val_step` 和 `test_step` 接口。接下来将详细介绍这些接口。
+MMSegmentation 封装 `BaseModel` 并实现了 [BaseSegmentor](https://github.com/open-mmlab/mmsegmentation/blob/1.x/mmseg/models/segmentors/base.py#L15) 类,主要提供 `forward`、`train_step`、`val_step` 和 `test_step` 接口。接下来将详细介绍这些接口。
### forward
 +
+ 


 +
+ 
 +
+ 